Paper List
-
A Unified Variational Principle for Branching Transport Networks: Wave Impedance, Viscous Flow, and Tissue Metabolism
This paper solves the core problem of predicting the empirically observed branching exponent (α≈2.7) in mammalian arterial trees, which neither Murray...
-
Household Bubbling Strategies for Epidemic Control and Social Connectivity
This paper addresses the core challenge of designing household merging (social bubble) strategies that effectively control epidemic risk while maximiz...
-
Empowering Chemical Structures with Biological Insights for Scalable Phenotypic Virtual Screening
This paper addresses the core challenge of bridging the gap between scalable chemical structure screening and biologically informative but resource-in...
-
A mechanical bifurcation constrains the evolution of cell sheet folding in the family Volvocaceae
This paper addresses the core problem of why there is an evolutionary gap in species with intermediate cell numbers (e.g., 256 cells) in Volvocaceae, ...
-
Bayesian Inference in Epidemic Modelling: A Beginner’s Guide Illustrated with the SIR Model
This guide addresses the core challenge of estimating uncertain epidemiological parameters (like transmission and recovery rates) from noisy, real-wor...
-
Geometric framework for biological evolution
This paper addresses the fundamental challenge of developing a coordinate-independent, geometric description of evolutionary dynamics that bridges gen...
-
A multiscale discrete-to-continuum framework for structured population models
This paper addresses the core challenge of systematically deriving uniformly valid continuum approximations from discrete structured population models...
-
Whole slide and microscopy image analysis with QuPath and OMERO
使QuPath能够直接分析存储在OMERO服务器中的图像而无需下载整个数据集,克服了大规模研究的本地存储限制。
STAR-GO: Improving Protein Function Prediction by Learning to Hierarchically Integrate Ontology-Informed Semantic Embeddings
Department of Computer Engineering, Bogazici University, Istanbul, Turkiye
30秒速读
IN SHORT: This paper addresses the core challenge of generalizing protein function prediction to unseen or newly introduced Gene Ontology (GO) terms by overcoming the limitations of existing models that either prioritize graph structure at the expense of semantic meaning or vice versa.
核心创新
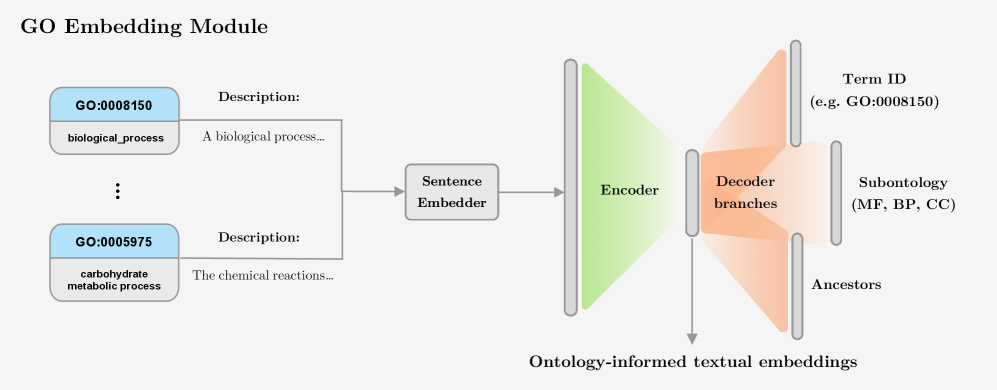
- Methodology Introduces a novel GO embedding module that integrates textual definitions (via SBERT-BioBERT) with ontology graph structure through a multi-task autoencoder, learning unified representations that preserve both semantic similarity and hierarchical dependencies.
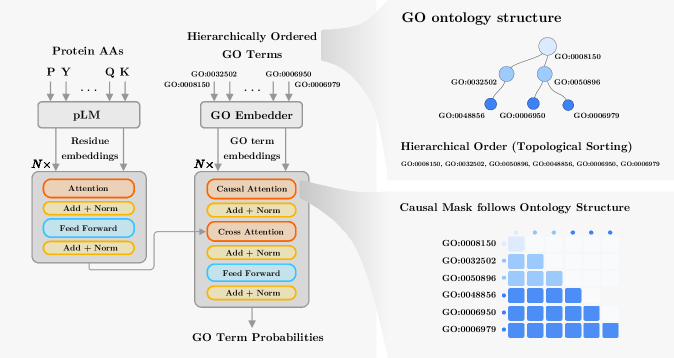
- Methodology Proposes a hierarchical Transformer decoder that processes GO terms in topological order (ancestors to descendants) using causal self-attention, enabling information propagation across ontology levels and capturing functional dependencies.
- Biology Demonstrates superior zero-shot generalization to unseen GO terms, particularly for Molecular Function and Biological Process terms, by effectively leveraging semantic information from textual definitions, which transfers better to novel ontology concepts than purely structural embeddings.
主要结论
- STAR-GO achieves state-of-the-art or competitive performance across all three GO subontologies (BP, CC, MF), with the highest AUC scores (e.g., 0.989 for BP, 0.988 for CC, 0.995 for MF), indicating strong term-level discriminability.
- In zero-shot evaluation on 16 held-out GO terms, STAR-GO variants achieve the highest AUCs in 13 cases, significantly outperforming baselines like DeepGOZero and DeepGO-SE, demonstrating superior generalization to unseen functions.
- Ablation studies reveal that semantic embeddings (STAR_T) achieve the best zero-shot results for most MF and BP terms (e.g., AUC of 0.949 for GO:0001228), while structural embeddings (STAR_S) perform best for a few terms but poorly for MF, highlighting the critical role of semantic information for generalization.
摘要: Motivation: Accurate prediction of protein function is essential for elucidating molecular mechanisms and advancing biological and therapeutic discovery. Yet experimental annotation lags far behind the rapid growth of protein sequence data. Computational approaches address this gap by associating proteins with Gene Ontology (GO) terms, which encode functional knowledge through hierarchical relations and textual definitions. However, existing models often emphasize one modality over the other, limiting their ability to generalize, particularly to unseen or newly introduced GO terms that frequently arise as the ontology evolves, and making the previously trained models outdated. Results: We present STAR-GO, a Transformer-based framework that jointly models the semantic and structural characteristics of GO terms to enhance zero-shot protein function prediction. STAR-GO integrates textual definitions with ontology graph structure to learn unified GO representations, which are processed in hierarchical order to propagate information from general to specific terms. These representations are then aligned with protein sequence embeddings to capture sequence–function relationships. STAR-GO achieves state-of-the-art performance and superior zero-shot generalization, demonstrating the utility of integrating semantics and structure for robust and adaptable protein function prediction. Availability: Code and pre-trained models are available at https://github.com/boun-tabi-lifelu/stargo.