Paper List
-
A Unified Variational Principle for Branching Transport Networks: Wave Impedance, Viscous Flow, and Tissue Metabolism
This paper solves the core problem of predicting the empirically observed branching exponent (α≈2.7) in mammalian arterial trees, which neither Murray...
-
Household Bubbling Strategies for Epidemic Control and Social Connectivity
This paper addresses the core challenge of designing household merging (social bubble) strategies that effectively control epidemic risk while maximiz...
-
Empowering Chemical Structures with Biological Insights for Scalable Phenotypic Virtual Screening
This paper addresses the core challenge of bridging the gap between scalable chemical structure screening and biologically informative but resource-in...
-
A mechanical bifurcation constrains the evolution of cell sheet folding in the family Volvocaceae
This paper addresses the core problem of why there is an evolutionary gap in species with intermediate cell numbers (e.g., 256 cells) in Volvocaceae, ...
-
Bayesian Inference in Epidemic Modelling: A Beginner’s Guide Illustrated with the SIR Model
This guide addresses the core challenge of estimating uncertain epidemiological parameters (like transmission and recovery rates) from noisy, real-wor...
-
Geometric framework for biological evolution
This paper addresses the fundamental challenge of developing a coordinate-independent, geometric description of evolutionary dynamics that bridges gen...
-
A multiscale discrete-to-continuum framework for structured population models
This paper addresses the core challenge of systematically deriving uniformly valid continuum approximations from discrete structured population models...
-
Whole slide and microscopy image analysis with QuPath and OMERO
使QuPath能够直接分析存储在OMERO服务器中的图像而无需下载整个数据集,克服了大规模研究的本地存储限制。
DeepFRI Demystified: Interpretability vs. Accuracy in AI Protein Function Prediction
Yale University | Microsoft
30秒速读
IN SHORT: This study addresses the critical gap between high predictive accuracy and biological interpretability in DeepFRI, revealing that the model often prioritizes structural motifs over functional residues, complicating reliable identification of drug targets.
核心创新
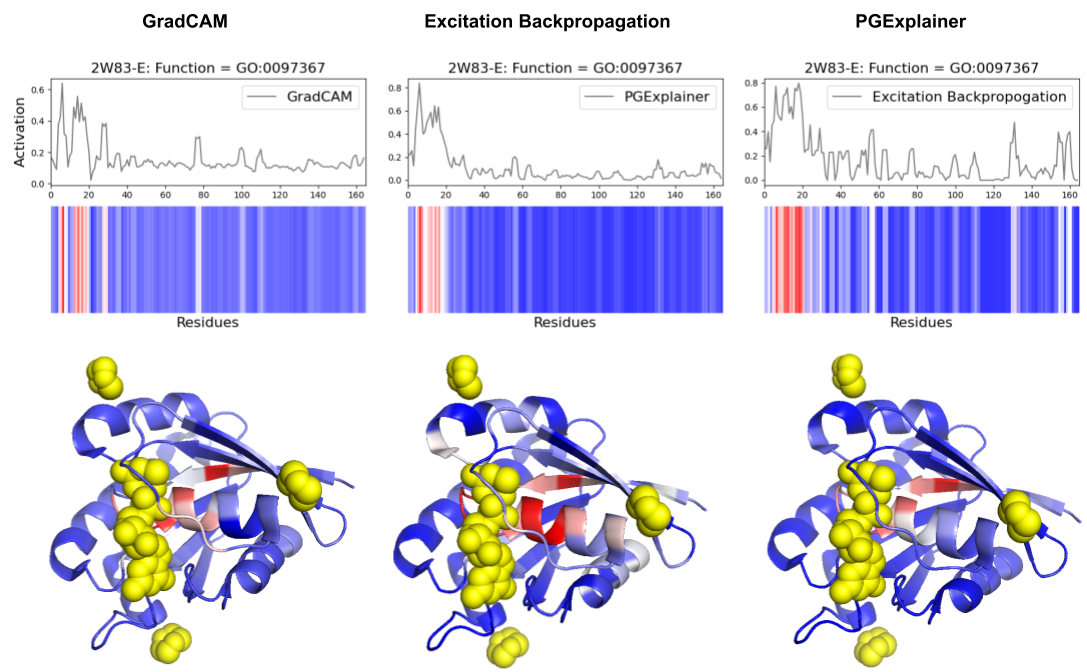
- Methodology Comprehensive benchmarking of three post-hoc explainability methods (GradCAM, Excitation Backpropagation, PGExplainer) on DeepFRI with quantitative sparsity analysis.
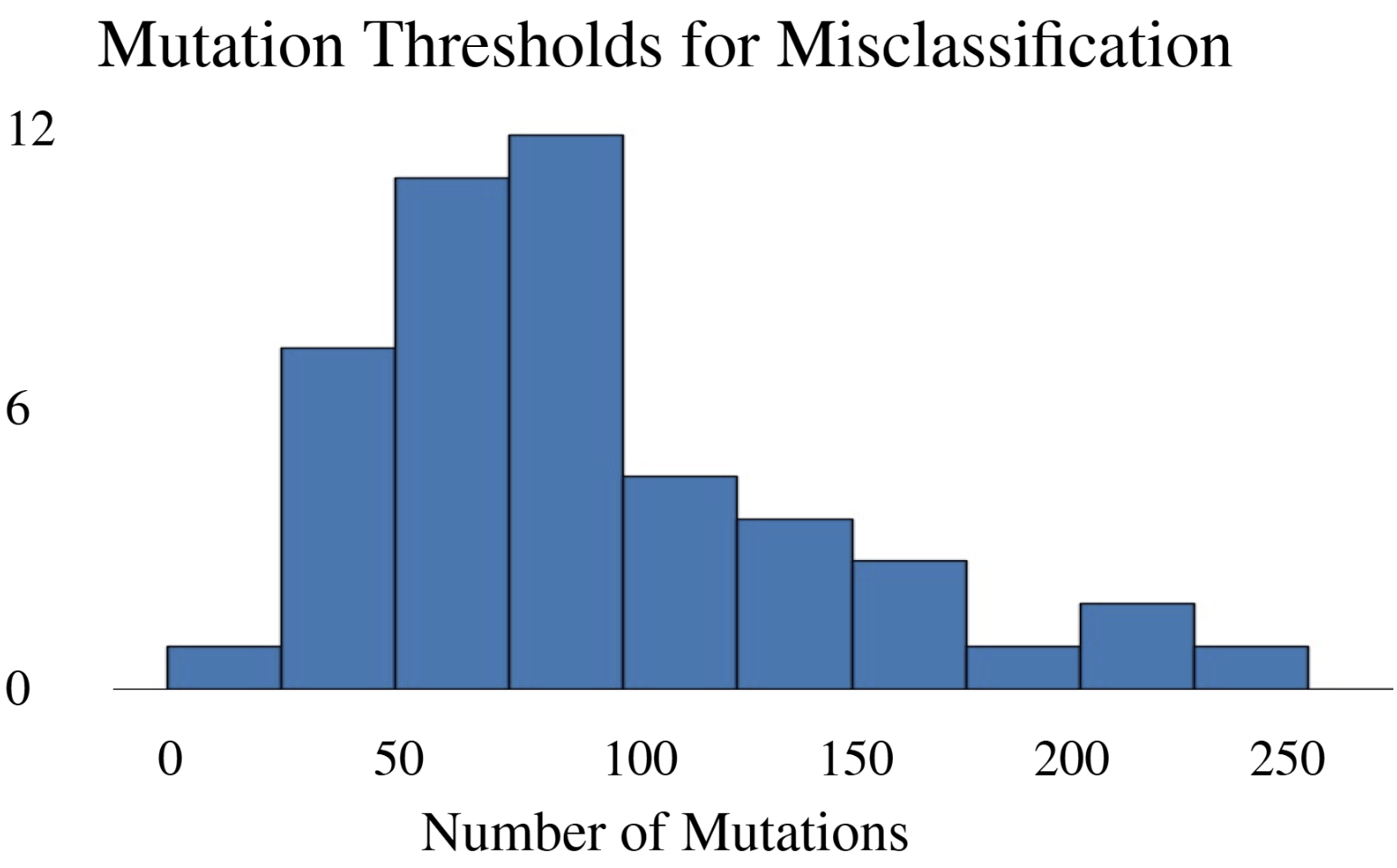
- Methodology Development of a modified DeepFool adversarial testing framework for protein sequences, measuring mutation thresholds required for misclassification.
- Biology Revealed that DeepFRI prioritizes amino acids controlling protein structure over function in >50% of tested proteins, highlighting a fundamental accuracy-interpretability trade-off.
主要结论
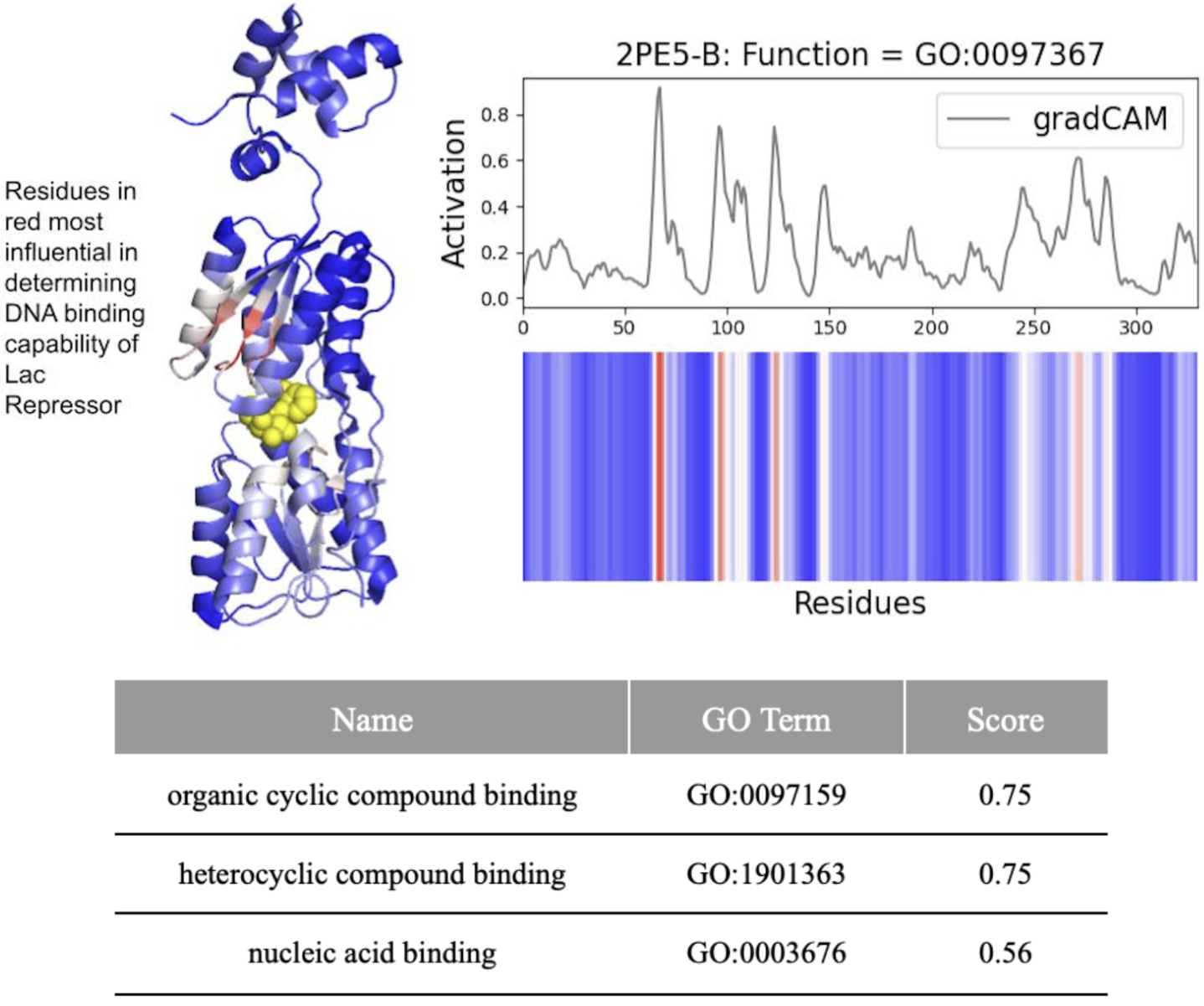
- DeepFRI required 206 mutations (62.4% of 330 residues) in the lac repressor for misclassification, demonstrating extreme robustness but potentially missing subtle functional alterations.
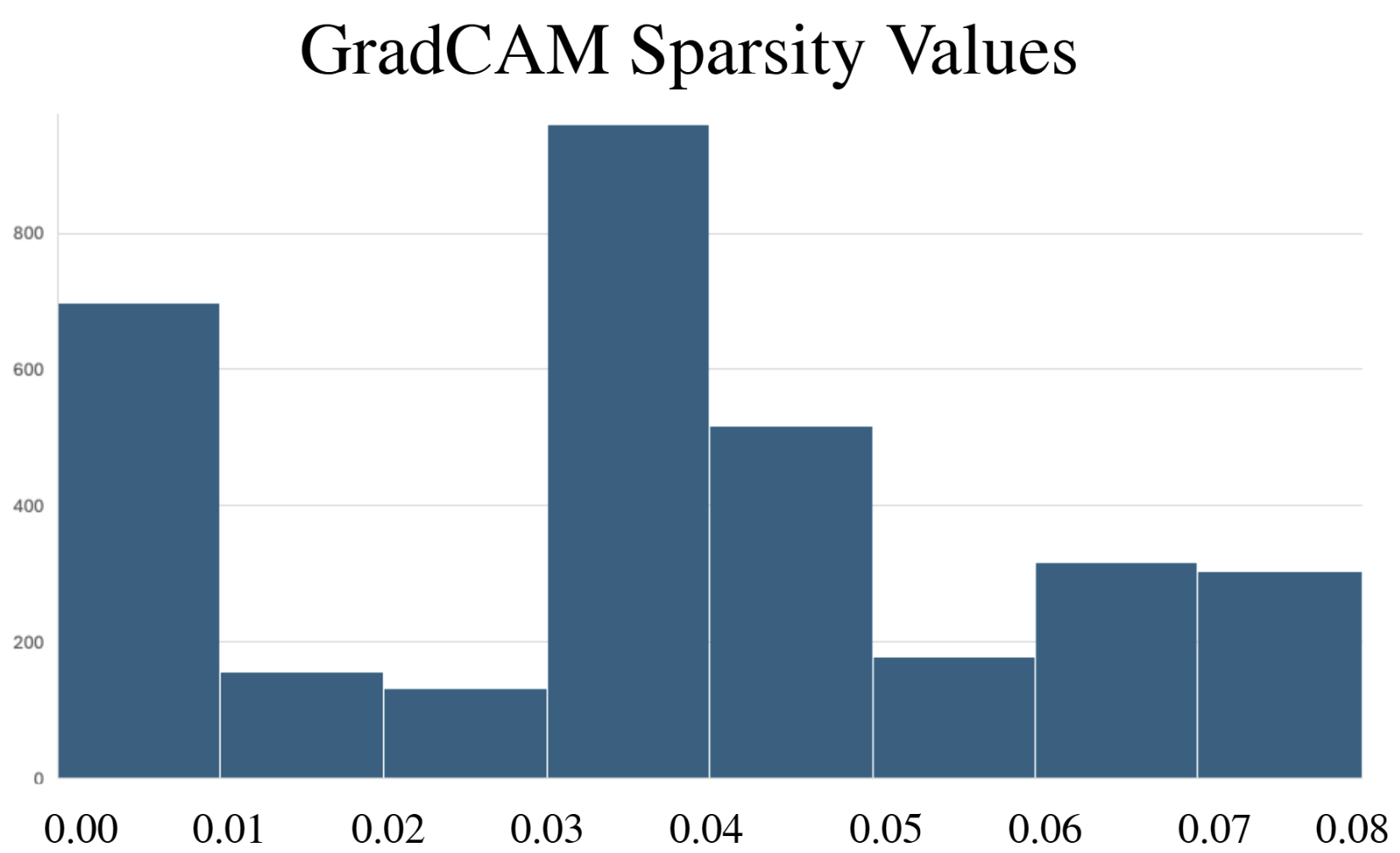
- Explainability methods showed significant granularity differences: PGExplainer was 3× sparser than GradCAM and 17× sparser than Excitation Backpropagation across 124 binding proteins.
- All three methods converged on biochemically critical P-loop residues (0-20) in ARF6 GTPase, validating DeepFRI's focus on conserved functional motifs in straightforward domains.
摘要: Machine learning technologies for protein function prediction are black box models. Despite their potential to identify key drug targets with high accuracy and accelerate therapy development, the adoption of these methods depends on verifying their findings. This study evaluates DeepFRI, a leading Graph Convolutional Network (GCN)-based tool, using advanced explainability techniques—GradCAM, Excitation Backpropagation, and PGExplainer—and adversarial robustness tests. Our findings reveal that the model’s predictions often prioritize conserved motifs over truly deterministic residues, complicating the identification of functional sites. Quantitative analyses show that explainability methods differ significantly in granularity, with GradCAM providing broad relevance and PGExplainer pinpointing specific active sites. These results highlight trade-offs between accuracy and interpretability, suggesting areas for improvement in DeepFRI’s architecture to enhance its trustworthiness in drug discovery and regulatory settings.