Paper List
-
A Unified Variational Principle for Branching Transport Networks: Wave Impedance, Viscous Flow, and Tissue Metabolism
This paper solves the core problem of predicting the empirically observed branching exponent (α≈2.7) in mammalian arterial trees, which neither Murray...
-
Household Bubbling Strategies for Epidemic Control and Social Connectivity
This paper addresses the core challenge of designing household merging (social bubble) strategies that effectively control epidemic risk while maximiz...
-
Empowering Chemical Structures with Biological Insights for Scalable Phenotypic Virtual Screening
This paper addresses the core challenge of bridging the gap between scalable chemical structure screening and biologically informative but resource-in...
-
A mechanical bifurcation constrains the evolution of cell sheet folding in the family Volvocaceae
This paper addresses the core problem of why there is an evolutionary gap in species with intermediate cell numbers (e.g., 256 cells) in Volvocaceae, ...
-
Bayesian Inference in Epidemic Modelling: A Beginner’s Guide Illustrated with the SIR Model
This guide addresses the core challenge of estimating uncertain epidemiological parameters (like transmission and recovery rates) from noisy, real-wor...
-
Geometric framework for biological evolution
This paper addresses the fundamental challenge of developing a coordinate-independent, geometric description of evolutionary dynamics that bridges gen...
-
A multiscale discrete-to-continuum framework for structured population models
This paper addresses the core challenge of systematically deriving uniformly valid continuum approximations from discrete structured population models...
-
Whole slide and microscopy image analysis with QuPath and OMERO
使QuPath能够直接分析存储在OMERO服务器中的图像而无需下载整个数据集,克服了大规模研究的本地存储限制。
SpikGPT: A High-Accuracy and Interpretable Spiking Attention Framework for Single-Cell Annotation
Department of Biomedical Informatics, Emory University | Department of Surgery, Duke University
30秒速读
IN SHORT: This paper addresses the core challenge of robust single-cell annotation across heterogeneous datasets with batch effects and the critical need to identify previously unseen cell populations.
核心创新
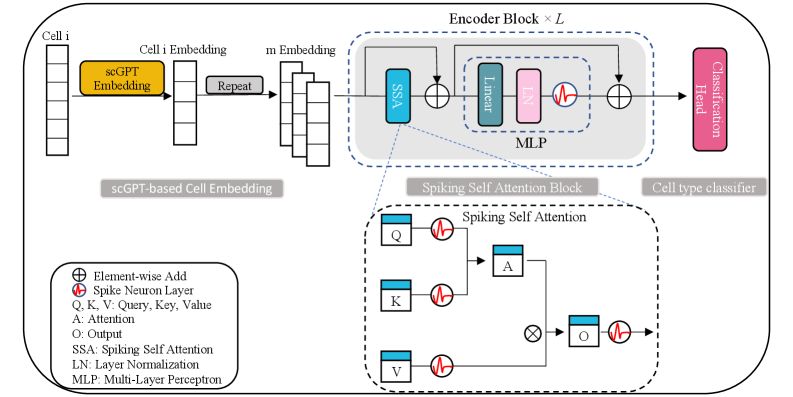
- Methodology First integration of spiking neural networks with transformer architecture for single-cell analysis, using Leaky Integrate-and-Fire (LIF) neurons in a multi-head Spiking Self-Attention mechanism for energy-efficient computation.
- Methodology Novel two-step embedding expansion strategy: repeating cell embeddings along feature channels (default m=300) and temporal dimensions (default T=4) to enhance representation richness and training stability.
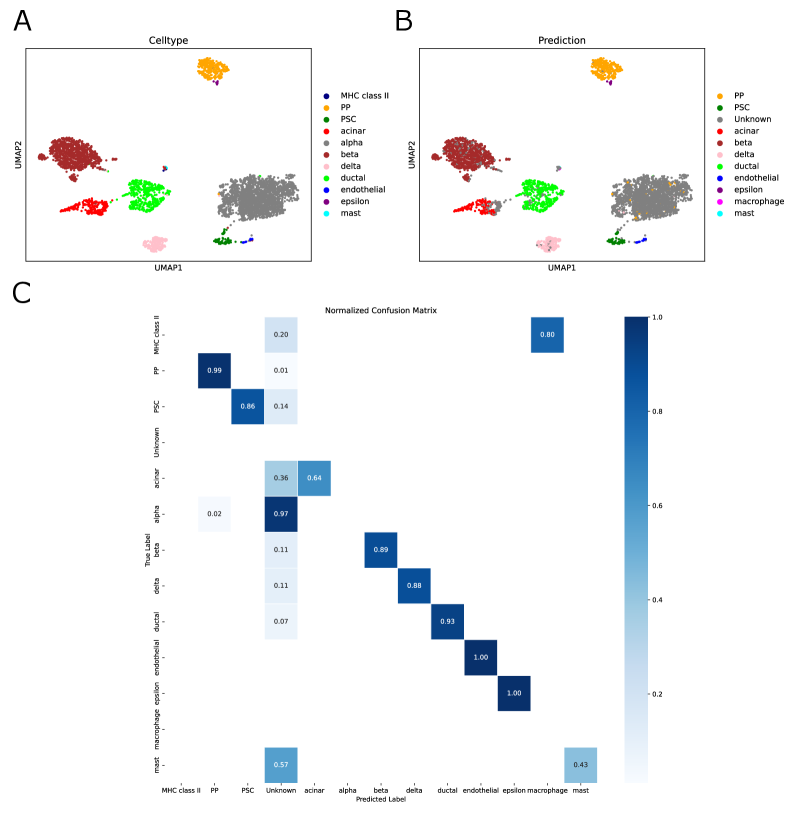
- Biology Confidence-based rejection mechanism that successfully identifies 97% of unseen 'alpha cells' as 'Unknown' in pancreas datasets, enabling robust detection of novel cell types absent from training data.
主要结论
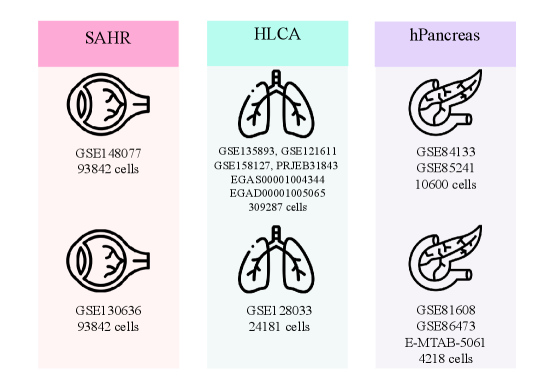
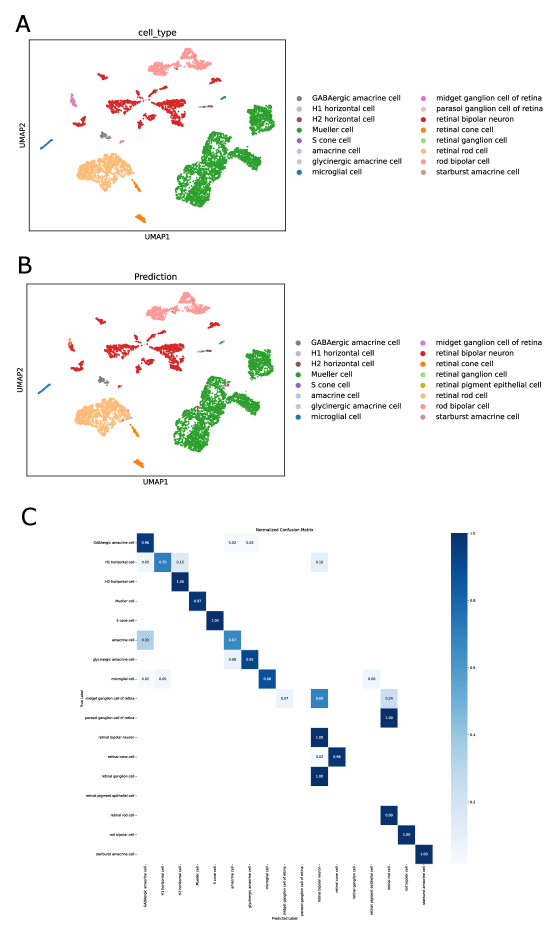
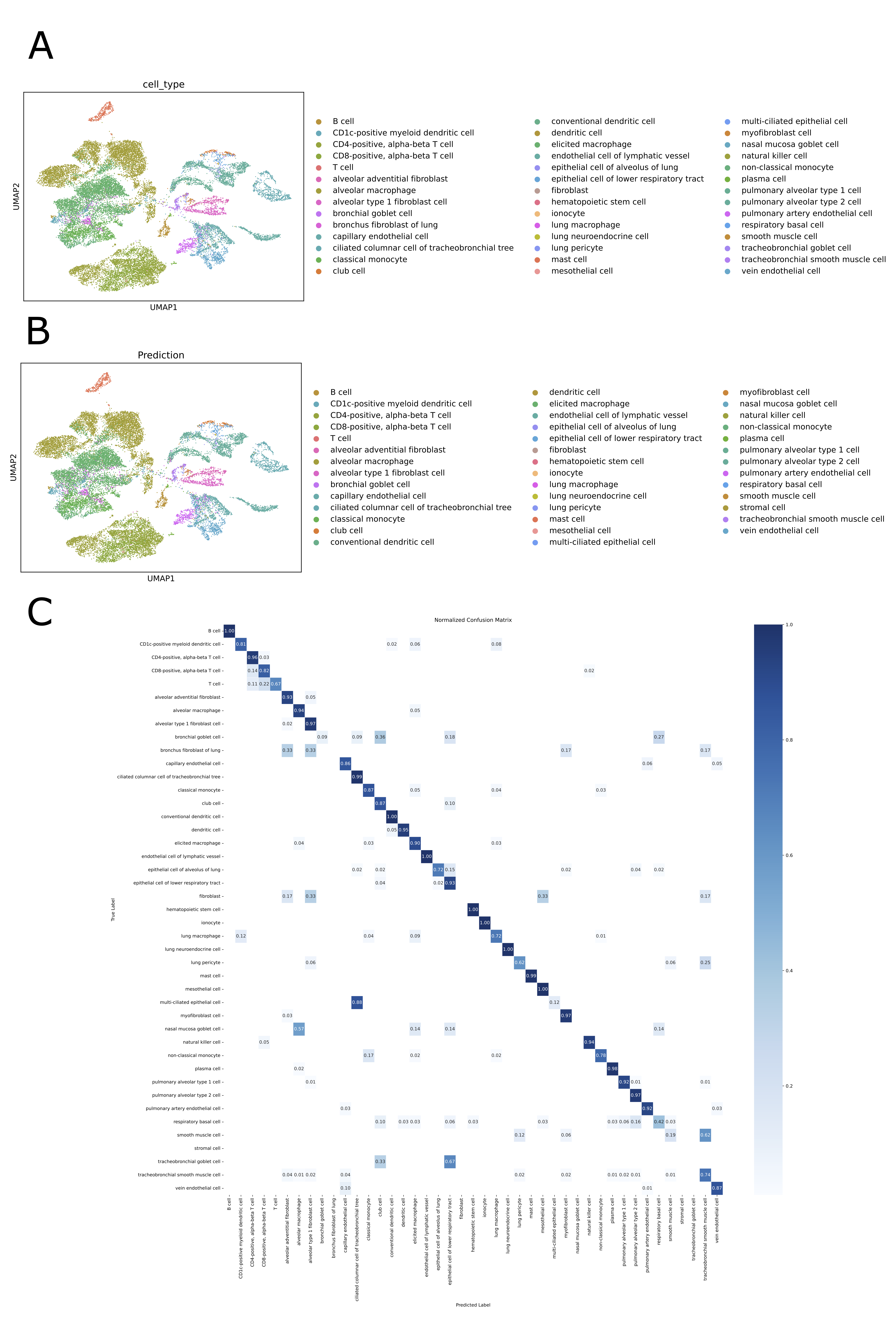
- SpikGPT achieves accuracy of 0.991 on SAHR dataset and 0.920 on HLCA dataset, outperforming or matching 8 benchmark methods including scGPT, CCA, and scPred.
- The model demonstrates superior robustness to batch effects, maintaining macro F1-score of 0.711 on heterogeneous HLCA data where traditional methods like SingleR drop to 0.207 F1-score.
- SpikGPT successfully identifies 97% of unseen 'alpha cells' as 'Unknown' using confidence thresholding (p<0.05), enabling reliable detection of novel cell populations.
摘要: Accurate and scalable cell type annotation remains a challenge in single-cell transcriptomics, especially when datasets exhibit strong batch effects or contain previously unseen cell populations. Here we introduce SpikGPT, a hybrid deep learning framework that integrates scGPT-derived cell embeddings with a spiking Transformer architecture to achieve efficient and robust annotation. scGPT provides biologically informed dense representations of each cell, which are further processed by a multi-head Spiking Self-Attention mechanism, energy-efficient feature extraction. Across multiple benchmark datasets, SpikGPT consistently matches or exceeds the performance of leading annotation tools. Notably, SpikGPT uniquely identifies unseen cell types by assigning low-confidence predictions to an 'Unknown' category, allowing accurate rejection of cell states absent from the training reference. Together, these results demonstrate that SpikGPT is a versatile and reliable annotation tool capable of generalizing across datasets, resolving complex cellular heterogeneity, and facilitating discovery of novel or disease-associated cell populations.