Paper List
-
A Unified Variational Principle for Branching Transport Networks: Wave Impedance, Viscous Flow, and Tissue Metabolism
This paper solves the core problem of predicting the empirically observed branching exponent (α≈2.7) in mammalian arterial trees, which neither Murray...
-
Household Bubbling Strategies for Epidemic Control and Social Connectivity
This paper addresses the core challenge of designing household merging (social bubble) strategies that effectively control epidemic risk while maximiz...
-
Empowering Chemical Structures with Biological Insights for Scalable Phenotypic Virtual Screening
This paper addresses the core challenge of bridging the gap between scalable chemical structure screening and biologically informative but resource-in...
-
A mechanical bifurcation constrains the evolution of cell sheet folding in the family Volvocaceae
This paper addresses the core problem of why there is an evolutionary gap in species with intermediate cell numbers (e.g., 256 cells) in Volvocaceae, ...
-
Bayesian Inference in Epidemic Modelling: A Beginner’s Guide Illustrated with the SIR Model
This guide addresses the core challenge of estimating uncertain epidemiological parameters (like transmission and recovery rates) from noisy, real-wor...
-
Geometric framework for biological evolution
This paper addresses the fundamental challenge of developing a coordinate-independent, geometric description of evolutionary dynamics that bridges gen...
-
A multiscale discrete-to-continuum framework for structured population models
This paper addresses the core challenge of systematically deriving uniformly valid continuum approximations from discrete structured population models...
-
Whole slide and microscopy image analysis with QuPath and OMERO
使QuPath能够直接分析存储在OMERO服务器中的图像而无需下载整个数据集,克服了大规模研究的本地存储限制。
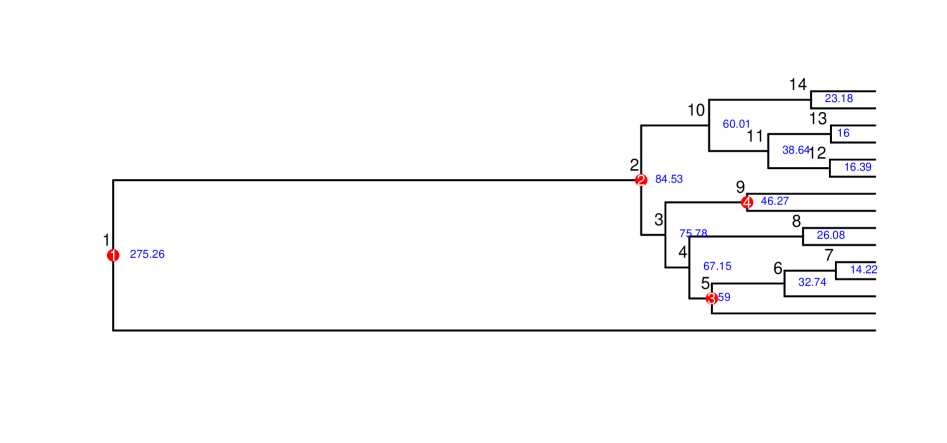
Fast and Accurate Node-Age Estimation Under Fossil Calibration Uncertainty Using the Adjusted Pairwise Likelihood
Department of Statistics, University of Georgia, Athens, 30601, USA
30秒速读
IN SHORT: This paper addresses the dual challenge of computational inefficiency and sensitivity to fossil calibration errors in Bayesian divergence time estimation for large phylogenomic datasets.
核心创新
- Methodology Introduces two Adjusted Pairwise Likelihood (APW) formulations (APW1 and APW2) that use asymptotic moment-matching weights to correct composite likelihoods within a Bayesian MCMC framework.
- Methodology Demonstrates that APW methods reduce computational cost by more than an order of magnitude compared to full-likelihood methods while maintaining comparable accuracy in node-age estimation.
- Methodology Shows that APW methods exhibit greater robustness to fossil misplacement and prior misspecification due to the reduced sensitivity of composite likelihoods to local calibration errors.
主要结论
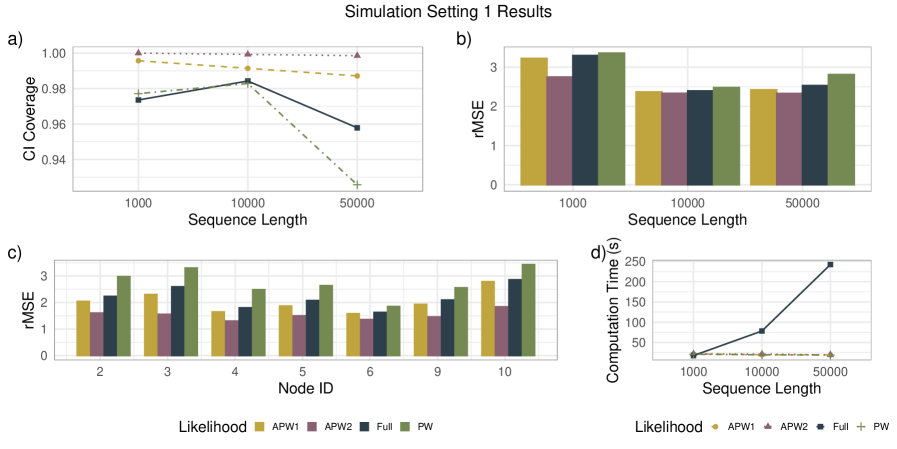
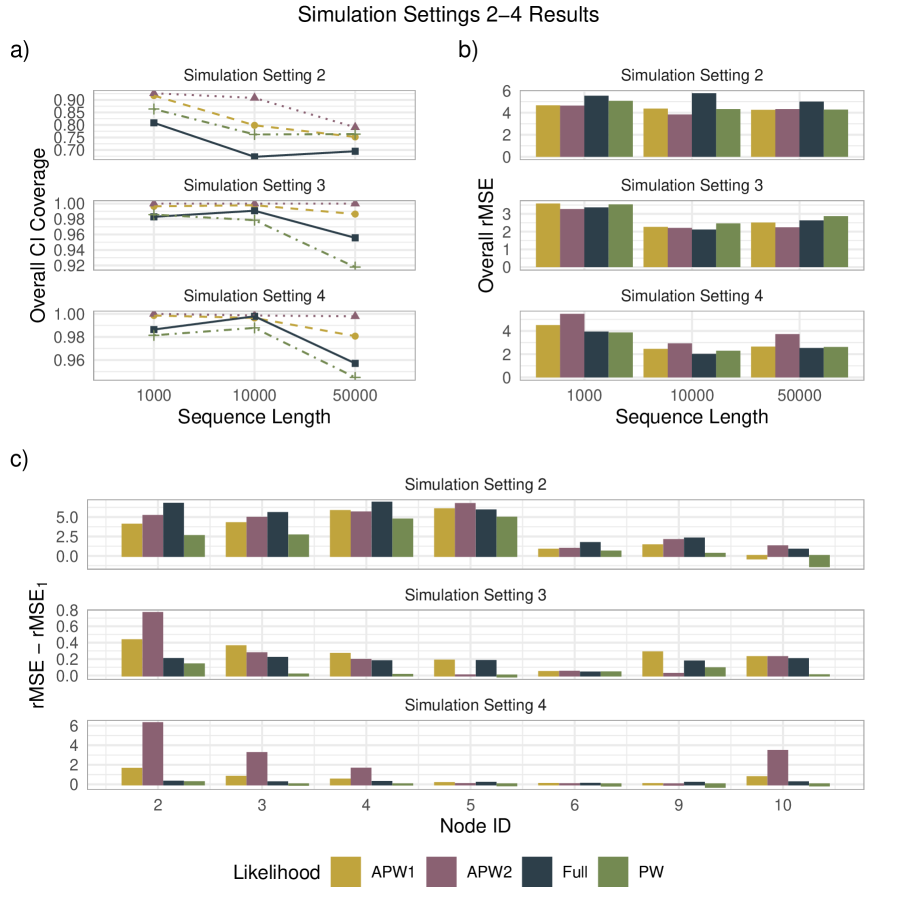
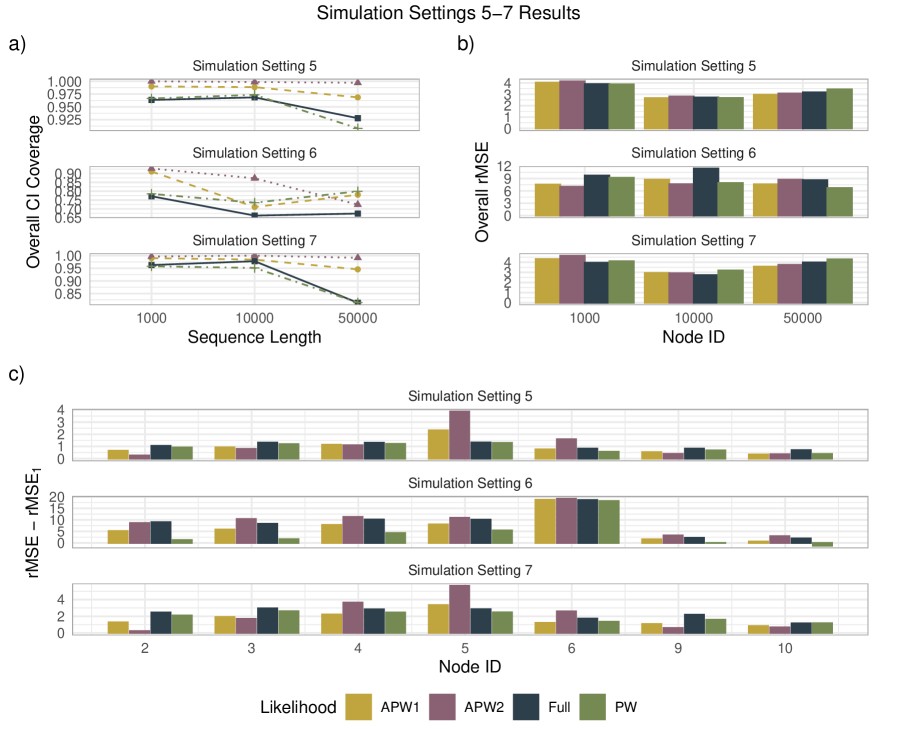
- APW methods produce node-age estimates statistically comparable to full-likelihood methods across diverse simulation scenarios, with reduced sensitivity to local calibration errors.
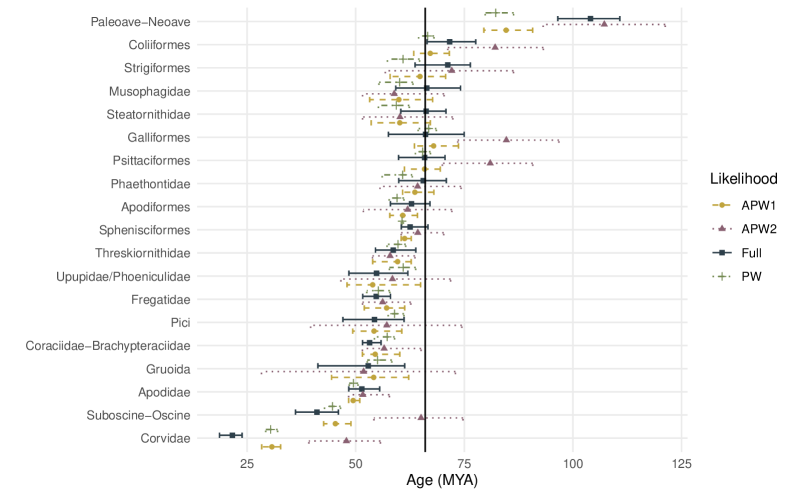
- Applied to a genome-scale avian dataset, APW recovered divergence time patterns consistent with recent studies while achieving a >10x reduction in computational cost.
- The robustness of APW to fossil misplacement stems from the composite likelihood's inherent property of being less sensitive to errors in individual calibration points, as demonstrated in simulations modeling various prior misspecifications.
摘要: Estimating divergence times from molecular sequence data is central to reconstructing the evolutionary history of lineages. Although Bayesian relaxed-clock methods provide a principled framework for incorporating fossil information, their dependence on repeated evaluations of the full phylogenetic likelihood makes them computationally demanding for large genomic datasets. Furthermore, because disagreements in divergence-time estimates often arise from uncertainty or error in fossil placement and prior specification, there is a need for methods that are both computationally efficient and robust to fossil-calibration uncertainty. In this study, we introduce fast and accurate alternatives based on the phylogenetic pairwise composite likelihood, presenting two adjusted pairwise likelihood (APW) formulations that employ asymptotic moment-matching weights to better approximate the behavior of the full likelihood within a Bayesian MCMC framework. Extensive simulations across diverse fossil-calibration scenarios show that APW methods produce node-age estimates comparable to those obtained from the full likelihood while offering greater robustness to fossil misplacement and prior misspecification, due to the reduced sensitivity of composite likelihoods to local calibration errors. Applied to a genome-scale dataset of modern birds, APW methods recover divergence time patterns consistent with recent studies, while reducing computational cost by more than an order of magnitude. Overall, our results demonstrate that adjusted pairwise likelihoods provide a calibration-robust and computationally efficient framework for Bayesian node dating, especially suited for large phylogenomic datasets and analyses in which fossil priors may be uncertain or imperfectly placed.