Paper List
-
A Unified Variational Principle for Branching Transport Networks: Wave Impedance, Viscous Flow, and Tissue Metabolism
This paper solves the core problem of predicting the empirically observed branching exponent (α≈2.7) in mammalian arterial trees, which neither Murray...
-
Household Bubbling Strategies for Epidemic Control and Social Connectivity
This paper addresses the core challenge of designing household merging (social bubble) strategies that effectively control epidemic risk while maximiz...
-
Empowering Chemical Structures with Biological Insights for Scalable Phenotypic Virtual Screening
This paper addresses the core challenge of bridging the gap between scalable chemical structure screening and biologically informative but resource-in...
-
A mechanical bifurcation constrains the evolution of cell sheet folding in the family Volvocaceae
This paper addresses the core problem of why there is an evolutionary gap in species with intermediate cell numbers (e.g., 256 cells) in Volvocaceae, ...
-
Bayesian Inference in Epidemic Modelling: A Beginner’s Guide Illustrated with the SIR Model
This guide addresses the core challenge of estimating uncertain epidemiological parameters (like transmission and recovery rates) from noisy, real-wor...
-
Geometric framework for biological evolution
This paper addresses the fundamental challenge of developing a coordinate-independent, geometric description of evolutionary dynamics that bridges gen...
-
A multiscale discrete-to-continuum framework for structured population models
This paper addresses the core challenge of systematically deriving uniformly valid continuum approximations from discrete structured population models...
-
Whole slide and microscopy image analysis with QuPath and OMERO
使QuPath能够直接分析存储在OMERO服务器中的图像而无需下载整个数据集,克服了大规模研究的本地存储限制。
Approximate Bayesian Inference on Mechanisms of Network Growth and Evolution
Harvard T.H. Chan School of Public Health
30秒速读
IN SHORT: This paper addresses the core challenge of inferring the relative contributions of multiple, simultaneous generative mechanisms in network formation when the true likelihood is intractable.
核心创新
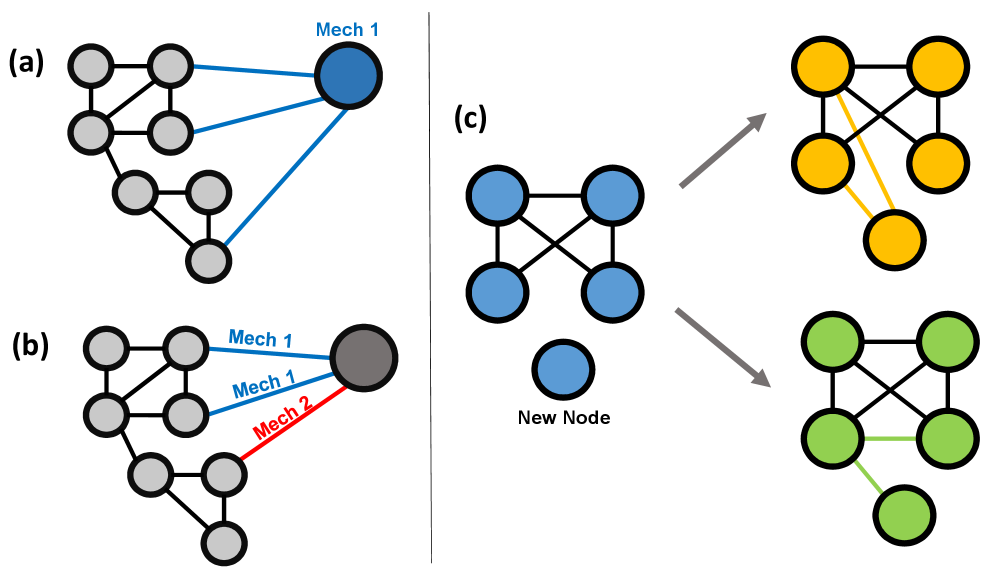
- Methodology Proposes an event-wise mixture-of-mechanisms model that assigns generative rules (e.g., Preferential Attachment, Random Attachment) to each edge formation event, rather than to nodes, increasing model flexibility and realism.
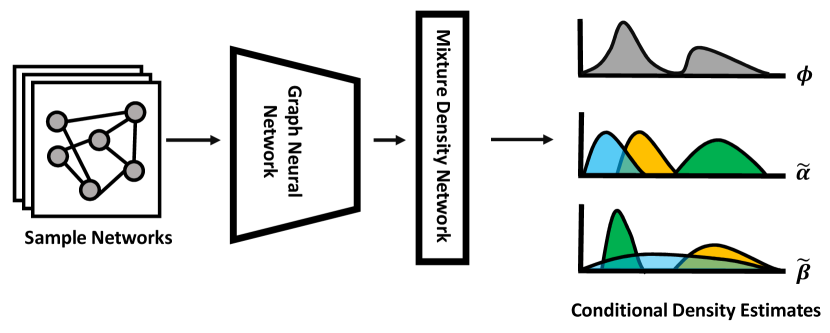
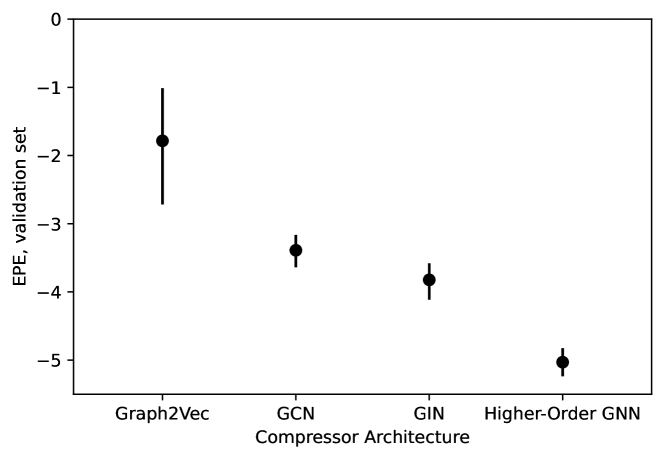
- Methodology Introduces a novel GNN-MDN (Graph Neural Network - Mixture Density Network) architecture that automatically learns informative, low-dimensional network embeddings for conditional density estimation, bypassing the need for manually specified summary statistics.
- Theory Formalizes a unified framework that incorporates both growth mechanisms (adding nodes/edges) and evolution mechanisms (modifying existing edges), allowing the model to capture a wider range of network dynamics like triangle formation.
主要结论
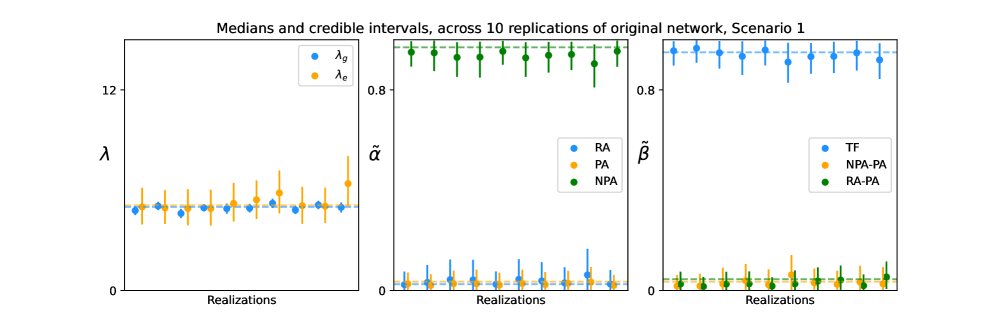
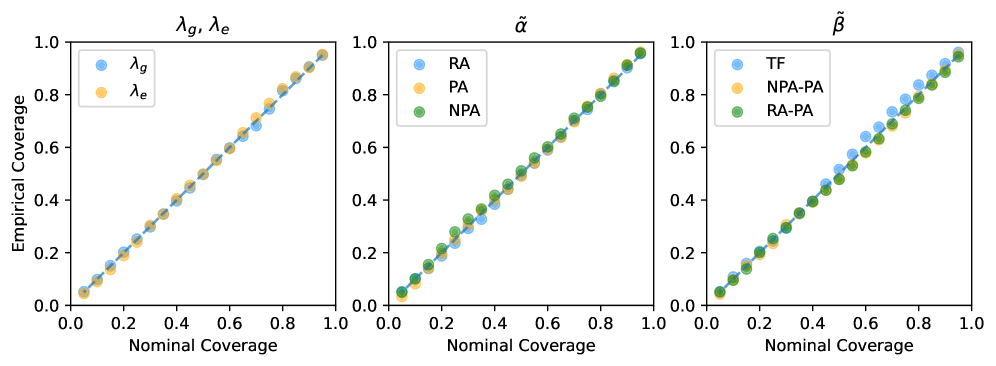
- The proposed GNN-MDN method provides valid approximate Bayesian inference, demonstrated via simulation studies showing that the 95% credible intervals achieve nominal coverage (e.g., containing the true parameter values).
- The event-wise model successfully infers dominant mechanisms in simulated scenarios; for instance, it accurately recovers a weight vector of (0.95, 0.025, 0.025) for a scenario where Preferential Attachment is the primary growth mechanism.
- The method is applicable to real-world networks, providing interpretable decompositions of their formation processes into quantifiable contributions from mechanisms like Random Attachment, Preferential Attachment, and Triangle Formation.
摘要: Mechanistic models can provide an intuitive and interpretable explanation of network growth by specifying a set of generative rules. These rules can be defined by domain knowledge about real-world mechanisms governing network growth or may be designed to facilitate the appearance of certain network motifs. In the formation of real-world networks, multiple mechanisms may be simultaneously involved; it is then important to understand the relative contribution of each of these mechanisms. In this paper, we propose the use of a conditional density estimator, augmented with a graph neural network, to perform inference on a flexible mixture of network-forming mechanisms. This event-wise mixture-of-mechanisms model assigns mechanisms to each edge formation event rather than stipulating node-level mechanisms, thus allowing for an explanation of the network generation process, as well as the dynamic evolution of the network over time. We demonstrate that our approximate Bayesian approach yields valid inferences for the relative weights of the mechanisms in our model, and we utilize this method to investigate the mechanisms behind the formation of a variety of real-world networks.