Paper List
-
SpikGPT: A High-Accuracy and Interpretable Spiking Attention Framework for Single-Cell Annotation
This paper addresses the core challenge of robust single-cell annotation across heterogeneous datasets with batch effects and the critical need to ide...
-
Unlocking hidden biomolecular conformational landscapes in diffusion models at inference time
This paper addresses the core challenge of efficiently and accurately sampling the conformational landscape of biomolecules from diffusion-based struc...
-
Personalized optimization of pediatric HD-tDCS for dose consistency and target engagement
This paper addresses the critical limitation of one-size-fits-all HD-tDCS protocols in pediatric populations by developing a personalized optimization...
-
Realistic Transition Paths for Large Biomolecular Systems: A Langevin Bridge Approach
This paper addresses the core challenge of generating physically realistic and computationally efficient transition paths between distinct protein con...
-
Consistent Synthetic Sequences Unlock Structural Diversity in Fully Atomistic De Novo Protein Design
This paper addresses the core pain point of low sequence-structure alignment in existing synthetic datasets (e.g., AFDB), which severely limits the pe...
-
MoRSAIK: Sequence Motif Reactor Simulation, Analysis and Inference Kit in Python
This work addresses the computational bottleneck in simulating prebiotic RNA reactor dynamics by developing a Python package that tracks sequence moti...
-
On the Approximation of Phylogenetic Distance Functions by Artificial Neural Networks
This paper addresses the core challenge of developing computationally efficient and scalable neural network architectures that can learn accurate phyl...
-
EcoCast: A Spatio-Temporal Model for Continual Biodiversity and Climate Risk Forecasting
This paper addresses the critical bottleneck in conservation: the lack of timely, high-resolution, near-term forecasts of species distribution shifts ...
Approximate Bayesian Inference on Mechanisms of Network Growth and Evolution
Harvard T.H. Chan School of Public Health
30秒速读
IN SHORT: This paper addresses the core challenge of inferring the relative contributions of multiple, simultaneous generative mechanisms in network formation when the true likelihood is intractable.
核心创新
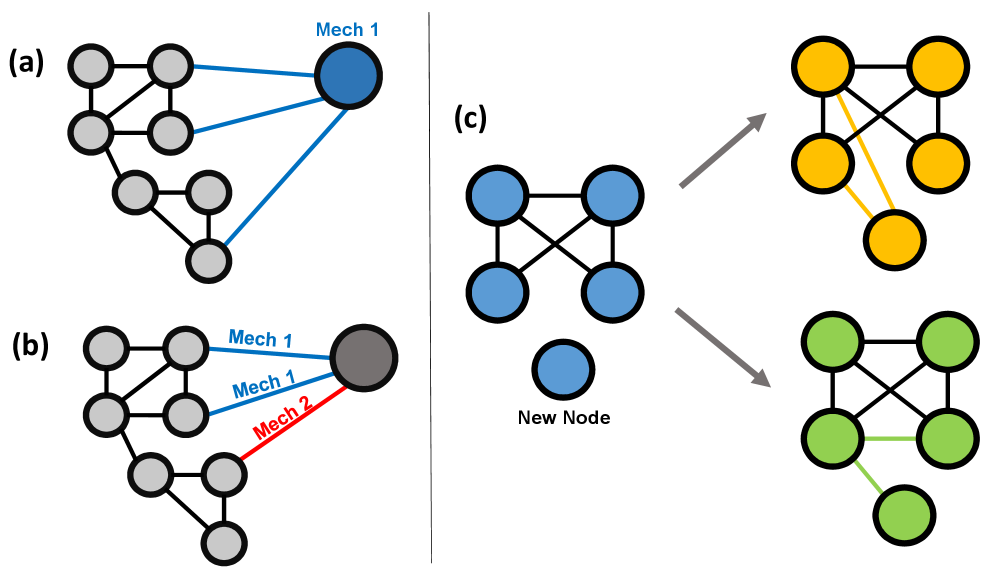
- Methodology Proposes an event-wise mixture-of-mechanisms model that assigns generative rules (e.g., Preferential Attachment, Random Attachment) to each edge formation event, rather than to nodes, increasing model flexibility and realism.
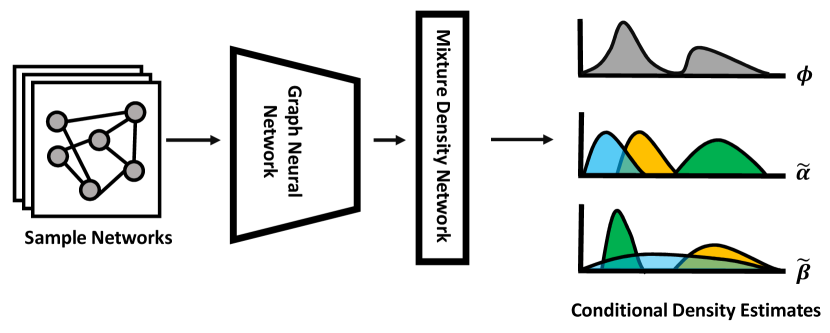
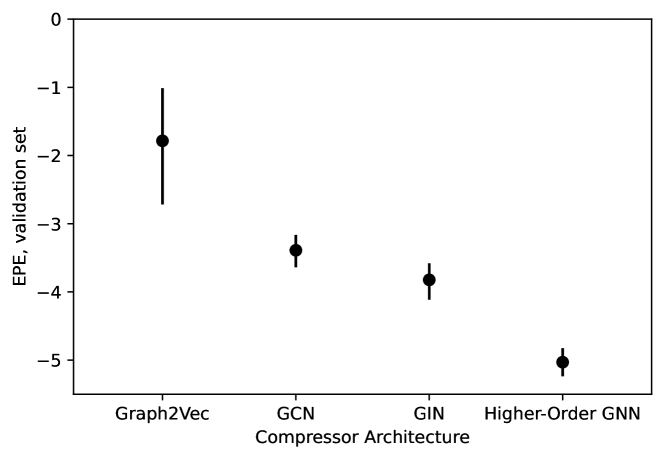
- Methodology Introduces a novel GNN-MDN (Graph Neural Network - Mixture Density Network) architecture that automatically learns informative, low-dimensional network embeddings for conditional density estimation, bypassing the need for manually specified summary statistics.
- Theory Formalizes a unified framework that incorporates both growth mechanisms (adding nodes/edges) and evolution mechanisms (modifying existing edges), allowing the model to capture a wider range of network dynamics like triangle formation.
主要结论
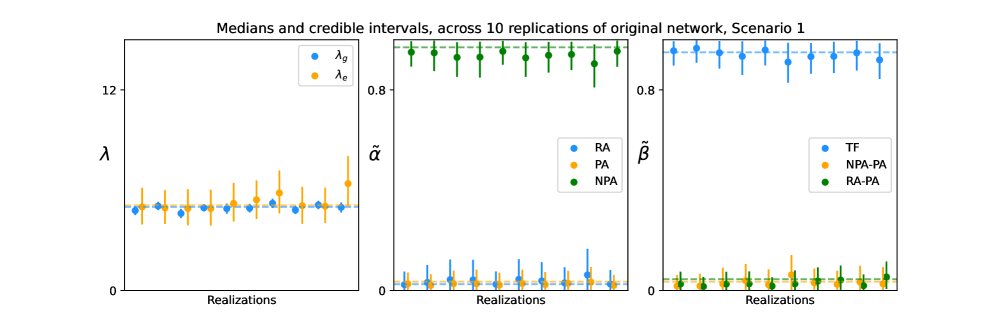
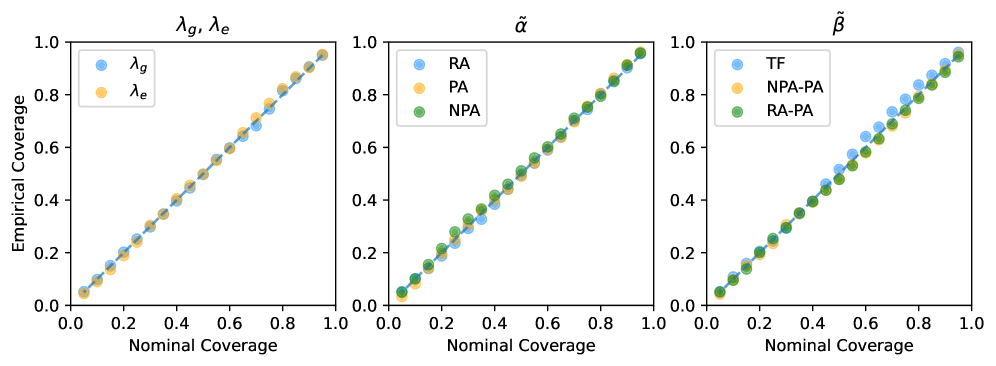
- The proposed GNN-MDN method provides valid approximate Bayesian inference, demonstrated via simulation studies showing that the 95% credible intervals achieve nominal coverage (e.g., containing the true parameter values).
- The event-wise model successfully infers dominant mechanisms in simulated scenarios; for instance, it accurately recovers a weight vector of (0.95, 0.025, 0.025) for a scenario where Preferential Attachment is the primary growth mechanism.
- The method is applicable to real-world networks, providing interpretable decompositions of their formation processes into quantifiable contributions from mechanisms like Random Attachment, Preferential Attachment, and Triangle Formation.
摘要: Mechanistic models can provide an intuitive and interpretable explanation of network growth by specifying a set of generative rules. These rules can be defined by domain knowledge about real-world mechanisms governing network growth or may be designed to facilitate the appearance of certain network motifs. In the formation of real-world networks, multiple mechanisms may be simultaneously involved; it is then important to understand the relative contribution of each of these mechanisms. In this paper, we propose the use of a conditional density estimator, augmented with a graph neural network, to perform inference on a flexible mixture of network-forming mechanisms. This event-wise mixture-of-mechanisms model assigns mechanisms to each edge formation event rather than stipulating node-level mechanisms, thus allowing for an explanation of the network generation process, as well as the dynamic evolution of the network over time. We demonstrate that our approximate Bayesian approach yields valid inferences for the relative weights of the mechanisms in our model, and we utilize this method to investigate the mechanisms behind the formation of a variety of real-world networks.