Paper List
-
Ill-Conditioning in Dictionary-Based Dynamic-Equation Learning: A Systems Biology Case Study
This paper addresses the critical challenge of numerical ill-conditioning and multicollinearity in library-based sparse regression methods (e.g., SIND...
-
Hybrid eTFCE–GRF: Exact Cluster-Size Retrieval with Analytical pp-Values for Voxel-Based Morphometry
This paper addresses the computational bottleneck in voxel-based neuroimaging analysis by providing a method that delivers exact cluster-size retrieva...
-
abx_amr_simulator: A simulation environment for antibiotic prescribing policy optimization under antimicrobial resistance
This paper addresses the critical challenge of quantitatively evaluating antibiotic prescribing policies under realistic uncertainty and partial obser...
-
PesTwin: a biology-informed Digital Twin for enabling precision farming
This paper addresses the critical bottleneck in precision agriculture: the inability to accurately forecast pest outbreaks in real-time, leading to su...
-
Equivariant Asynchronous Diffusion: An Adaptive Denoising Schedule for Accelerated Molecular Conformation Generation
This paper addresses the core challenge of generating physically plausible 3D molecular structures by bridging the gap between autoregressive methods ...
-
Omics Data Discovery Agents
This paper addresses the core challenge of making published omics data computationally reusable by automating the extraction, quantification, and inte...
-
Single-cell directional sensing at ultra-low chemoattractant concentrations from extreme first-passage events
This work addresses the core challenge of how a cell can rapidly and accurately determine the direction of a chemoattractant source when the signal is...
-
SDSR: A Spectral Divide-and-Conquer Approach for Species Tree Reconstruction
This paper addresses the computational bottleneck in reconstructing species trees from thousands of species and multiple genes by introducing a scalab...
Fast and Accurate Node-Age Estimation Under Fossil Calibration Uncertainty Using the Adjusted Pairwise Likelihood
Department of Statistics, University of Georgia, Athens, 30601, USA
30秒速读
IN SHORT: This paper addresses the dual challenge of computational inefficiency and sensitivity to fossil calibration errors in Bayesian divergence time estimation for large phylogenomic datasets.
核心创新
- Methodology Introduces two Adjusted Pairwise Likelihood (APW) formulations (APW1 and APW2) that use asymptotic moment-matching weights to correct composite likelihoods within a Bayesian MCMC framework.
- Methodology Demonstrates that APW methods reduce computational cost by more than an order of magnitude compared to full-likelihood methods while maintaining comparable accuracy in node-age estimation.
- Methodology Shows that APW methods exhibit greater robustness to fossil misplacement and prior misspecification due to the reduced sensitivity of composite likelihoods to local calibration errors.
主要结论
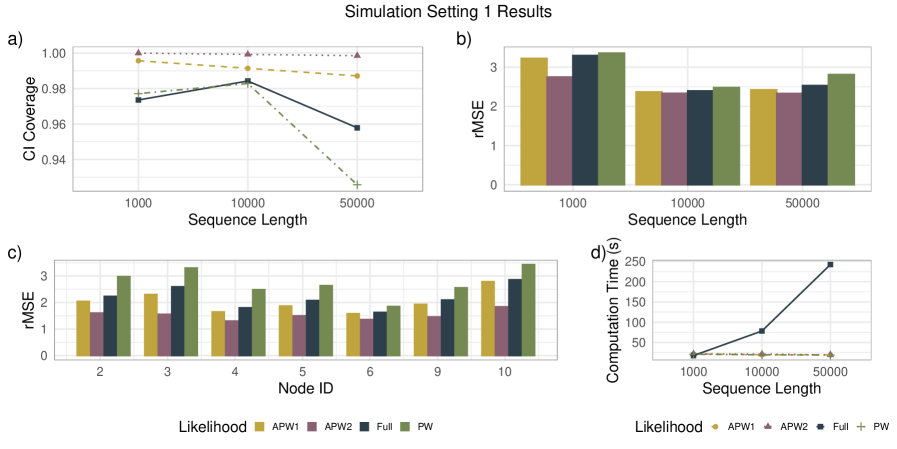
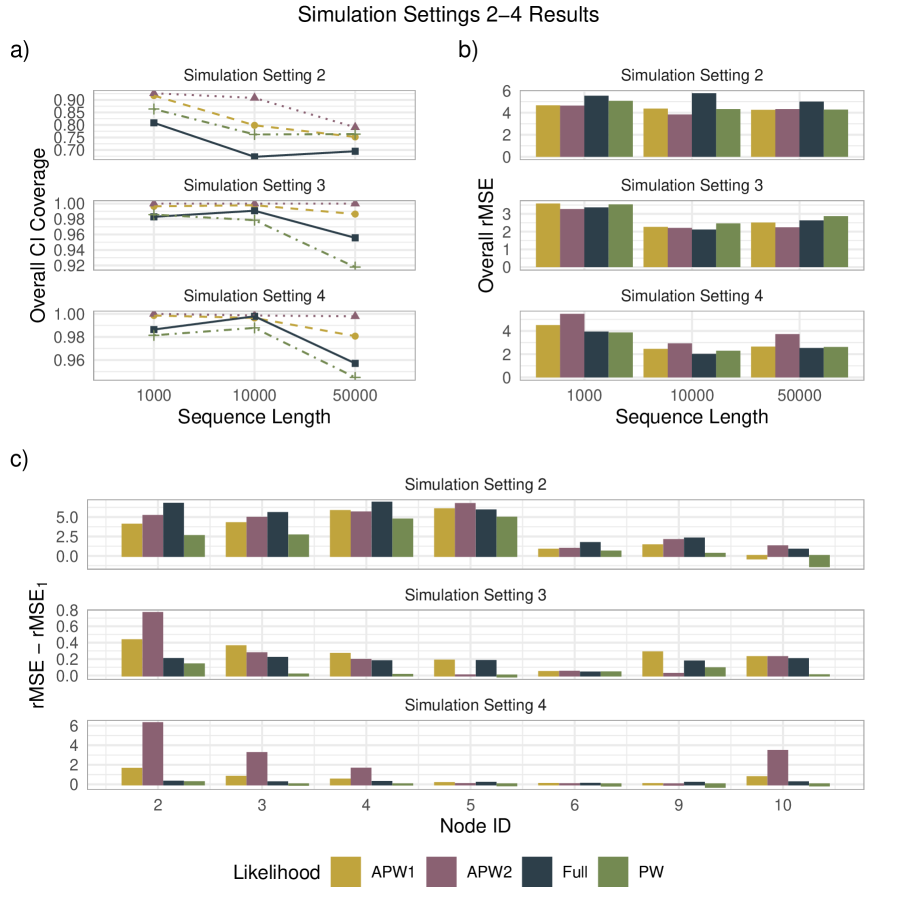
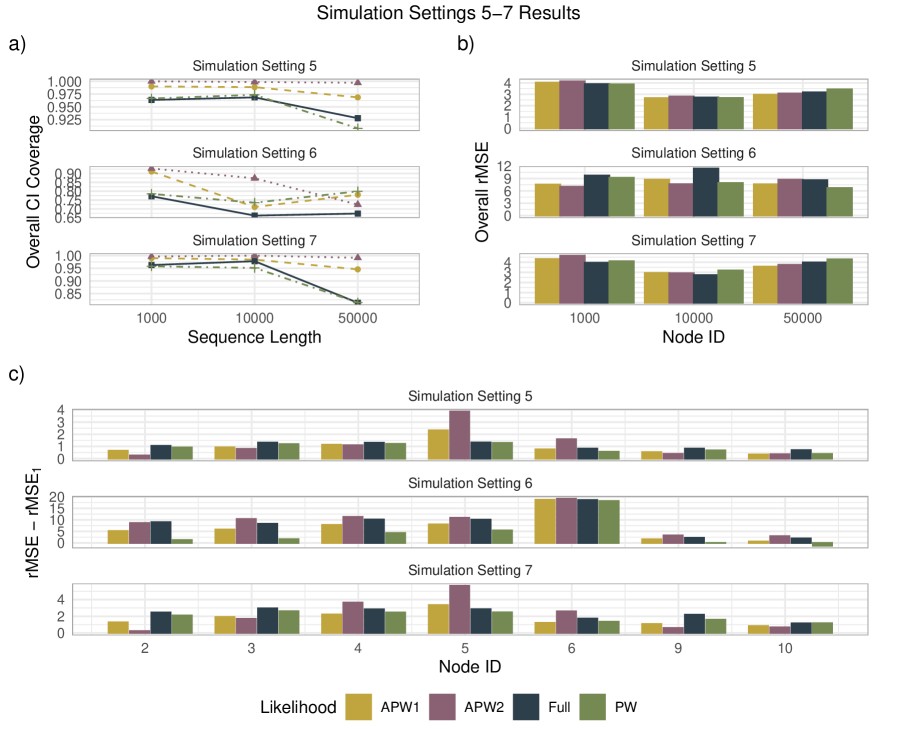
- APW methods produce node-age estimates statistically comparable to full-likelihood methods across diverse simulation scenarios, with reduced sensitivity to local calibration errors.
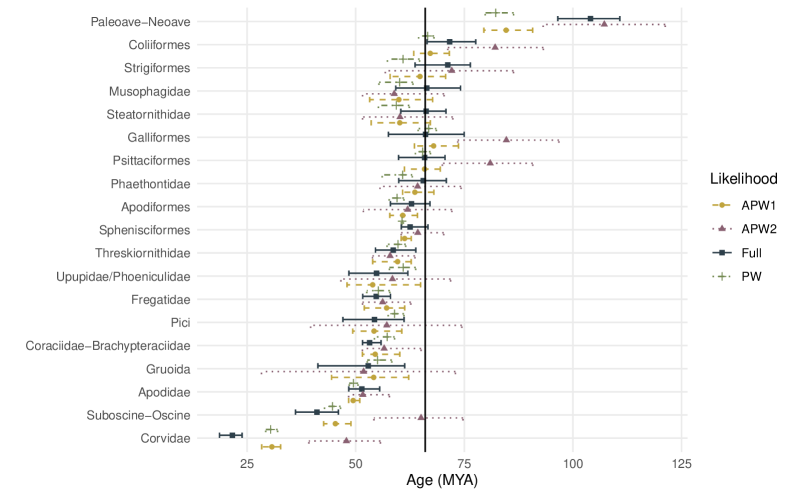
- Applied to a genome-scale avian dataset, APW recovered divergence time patterns consistent with recent studies while achieving a >10x reduction in computational cost.
- The robustness of APW to fossil misplacement stems from the composite likelihood's inherent property of being less sensitive to errors in individual calibration points, as demonstrated in simulations modeling various prior misspecifications.
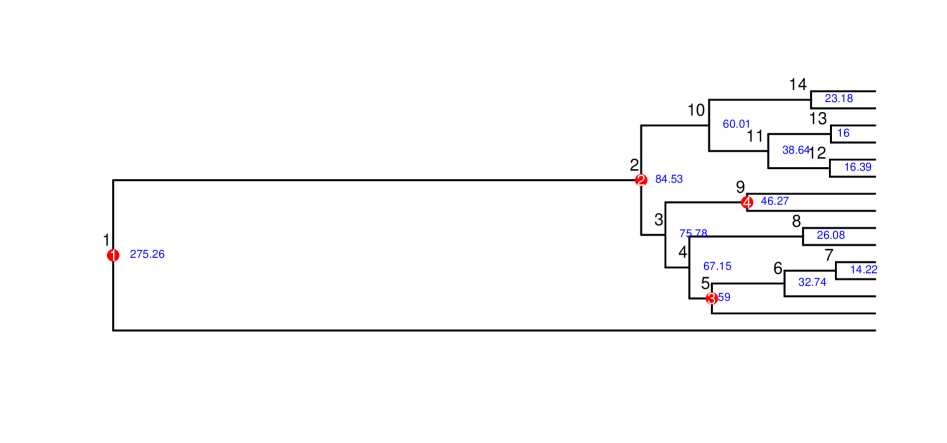
摘要: Estimating divergence times from molecular sequence data is central to reconstructing the evolutionary history of lineages. Although Bayesian relaxed-clock methods provide a principled framework for incorporating fossil information, their dependence on repeated evaluations of the full phylogenetic likelihood makes them computationally demanding for large genomic datasets. Furthermore, because disagreements in divergence-time estimates often arise from uncertainty or error in fossil placement and prior specification, there is a need for methods that are both computationally efficient and robust to fossil-calibration uncertainty. In this study, we introduce fast and accurate alternatives based on the phylogenetic pairwise composite likelihood, presenting two adjusted pairwise likelihood (APW) formulations that employ asymptotic moment-matching weights to better approximate the behavior of the full likelihood within a Bayesian MCMC framework. Extensive simulations across diverse fossil-calibration scenarios show that APW methods produce node-age estimates comparable to those obtained from the full likelihood while offering greater robustness to fossil misplacement and prior misspecification, due to the reduced sensitivity of composite likelihoods to local calibration errors. Applied to a genome-scale dataset of modern birds, APW methods recover divergence time patterns consistent with recent studies, while reducing computational cost by more than an order of magnitude. Overall, our results demonstrate that adjusted pairwise likelihoods provide a calibration-robust and computationally efficient framework for Bayesian node dating, especially suited for large phylogenomic datasets and analyses in which fossil priors may be uncertain or imperfectly placed.