Paper List
-
Autonomous Agents Coordinating Distributed Discovery Through Emergent Artifact Exchange
This paper addresses the fundamental limitation of current AI-assisted scientific research by enabling truly autonomous, decentralized investigation w...
-
D-MEM: Dopamine-Gated Agentic Memory via Reward Prediction Error Routing
This paper addresses the fundamental scalability bottleneck in LLM agentic memory systems: the O(N²) computational complexity and unbounded API token ...
-
Countershading coloration in blue shark skin emerges from hierarchically organized and spatially tuned photonic architectures inside skin denticles
This paper solves the core problem of how blue sharks achieve their striking dorsoventral countershading camouflage, revealing that coloration origina...
-
Human-like Object Grouping in Self-supervised Vision Transformers
This paper addresses the core challenge of quantifying how well self-supervised vision models capture human-like object grouping in natural scenes, br...
-
Hierarchical pp-Adic Framework for Gene Regulatory Networks: Theory and Stability Analysis
This paper addresses the core challenge of mathematically capturing the inherent hierarchical organization and multi-scale stability of gene regulator...
-
Towards unified brain-to-text decoding across speech production and perception
This paper addresses the core challenge of developing a unified brain-to-text decoding framework that works across both speech production and percepti...
-
Dual-Laws Model for a theory of artificial consciousness
This paper addresses the core challenge of developing a comprehensive, testable theory of consciousness that bridges biological and artificial systems...
-
Pulse desynchronization of neural populations by targeting the centroid of the limit cycle in phase space
This work addresses the core challenge of determining optimal pulse timing and intensity for desynchronizing pathological neural oscillations when the...
DeepFRI Demystified: Interpretability vs. Accuracy in AI Protein Function Prediction
Yale University | Microsoft
30秒速读
IN SHORT: This study addresses the critical gap between high predictive accuracy and biological interpretability in DeepFRI, revealing that the model often prioritizes structural motifs over functional residues, complicating reliable identification of drug targets.
核心创新
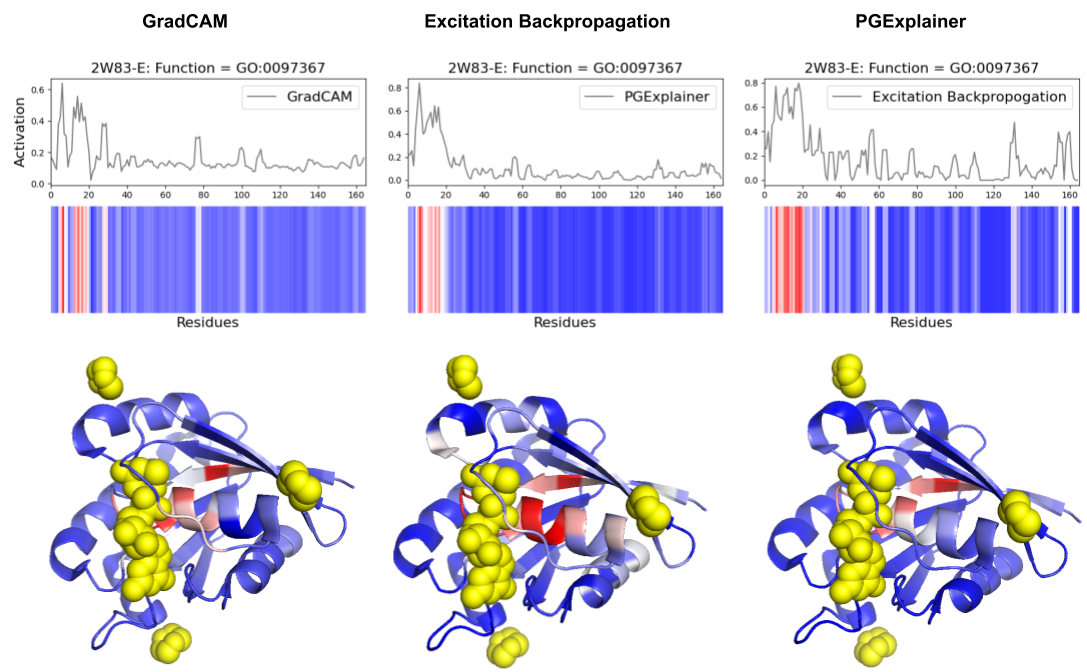
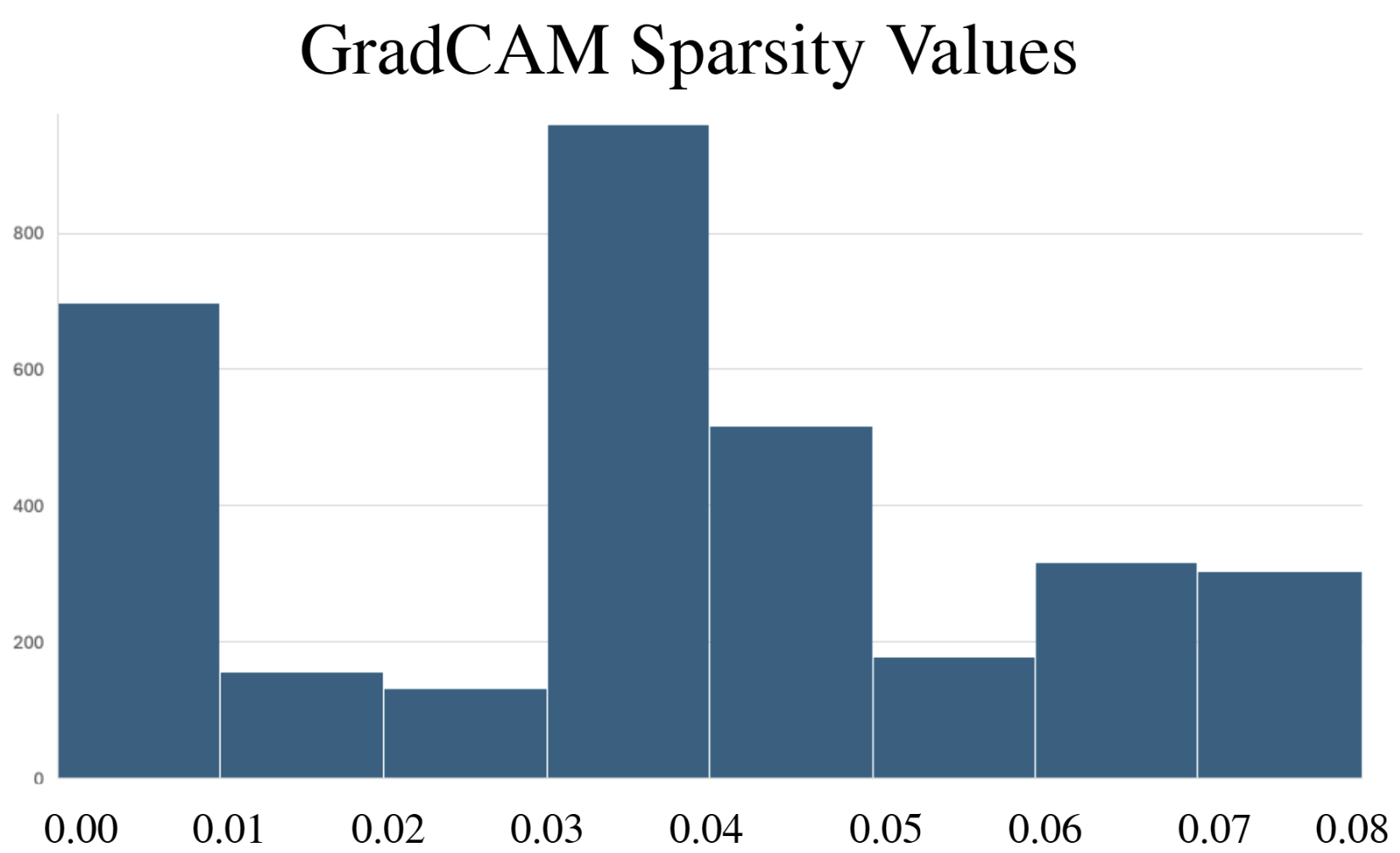
- Methodology Comprehensive benchmarking of three post-hoc explainability methods (GradCAM, Excitation Backpropagation, PGExplainer) on DeepFRI with quantitative sparsity analysis.
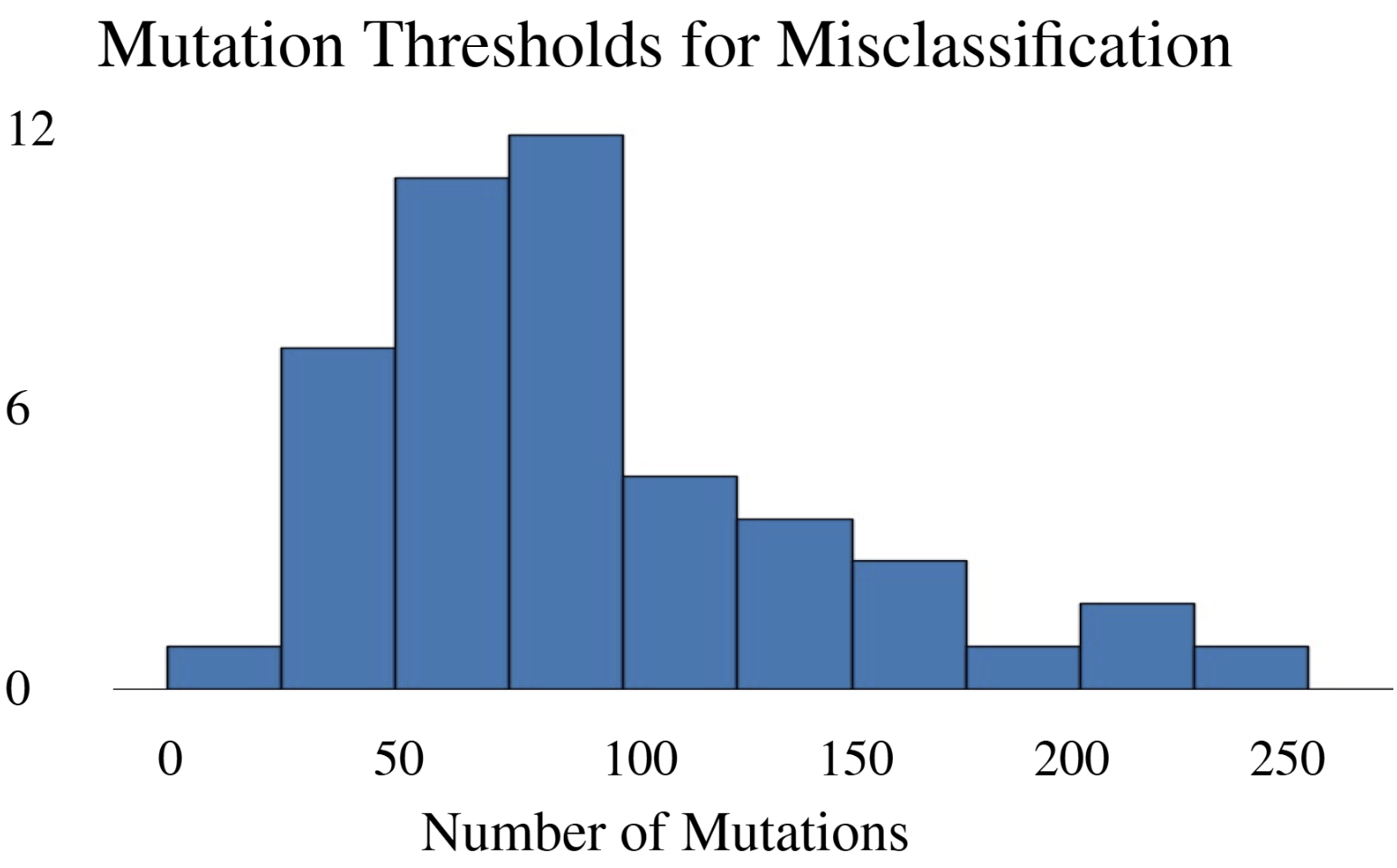
- Methodology Development of a modified DeepFool adversarial testing framework for protein sequences, measuring mutation thresholds required for misclassification.
- Biology Revealed that DeepFRI prioritizes amino acids controlling protein structure over function in >50% of tested proteins, highlighting a fundamental accuracy-interpretability trade-off.
主要结论
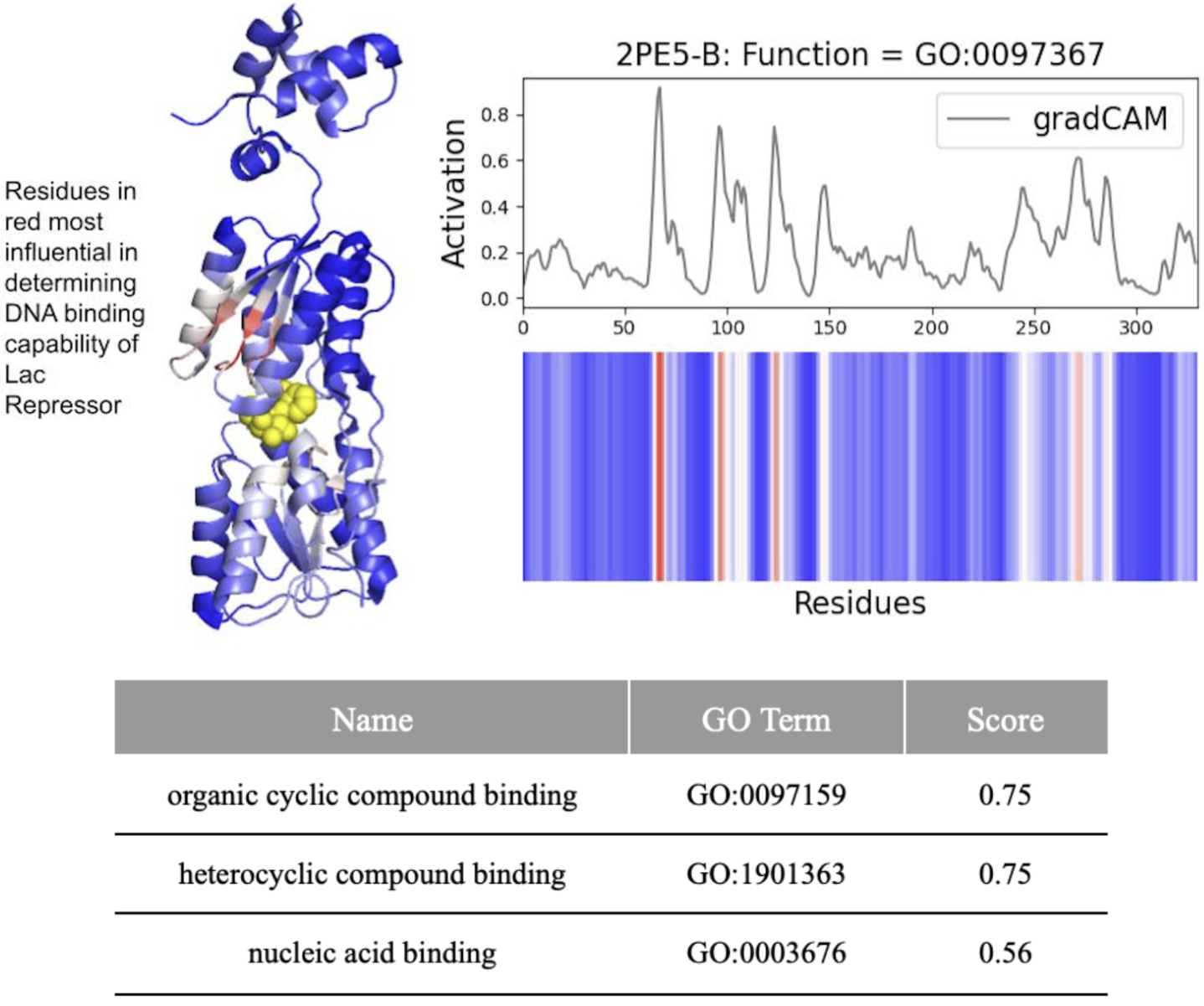
- DeepFRI required 206 mutations (62.4% of 330 residues) in the lac repressor for misclassification, demonstrating extreme robustness but potentially missing subtle functional alterations.
- Explainability methods showed significant granularity differences: PGExplainer was 3× sparser than GradCAM and 17× sparser than Excitation Backpropagation across 124 binding proteins.
- All three methods converged on biochemically critical P-loop residues (0-20) in ARF6 GTPase, validating DeepFRI's focus on conserved functional motifs in straightforward domains.
摘要: Machine learning technologies for protein function prediction are black box models. Despite their potential to identify key drug targets with high accuracy and accelerate therapy development, the adoption of these methods depends on verifying their findings. This study evaluates DeepFRI, a leading Graph Convolutional Network (GCN)-based tool, using advanced explainability techniques—GradCAM, Excitation Backpropagation, and PGExplainer—and adversarial robustness tests. Our findings reveal that the model’s predictions often prioritize conserved motifs over truly deterministic residues, complicating the identification of functional sites. Quantitative analyses show that explainability methods differ significantly in granularity, with GradCAM providing broad relevance and PGExplainer pinpointing specific active sites. These results highlight trade-offs between accuracy and interpretability, suggesting areas for improvement in DeepFRI’s architecture to enhance its trustworthiness in drug discovery and regulatory settings.