Paper List
-
Ill-Conditioning in Dictionary-Based Dynamic-Equation Learning: A Systems Biology Case Study
This paper addresses the critical challenge of numerical ill-conditioning and multicollinearity in library-based sparse regression methods (e.g., SIND...
-
Hybrid eTFCE–GRF: Exact Cluster-Size Retrieval with Analytical pp-Values for Voxel-Based Morphometry
This paper addresses the computational bottleneck in voxel-based neuroimaging analysis by providing a method that delivers exact cluster-size retrieva...
-
abx_amr_simulator: A simulation environment for antibiotic prescribing policy optimization under antimicrobial resistance
This paper addresses the critical challenge of quantitatively evaluating antibiotic prescribing policies under realistic uncertainty and partial obser...
-
PesTwin: a biology-informed Digital Twin for enabling precision farming
This paper addresses the critical bottleneck in precision agriculture: the inability to accurately forecast pest outbreaks in real-time, leading to su...
-
Equivariant Asynchronous Diffusion: An Adaptive Denoising Schedule for Accelerated Molecular Conformation Generation
This paper addresses the core challenge of generating physically plausible 3D molecular structures by bridging the gap between autoregressive methods ...
-
Omics Data Discovery Agents
This paper addresses the core challenge of making published omics data computationally reusable by automating the extraction, quantification, and inte...
-
Single-cell directional sensing at ultra-low chemoattractant concentrations from extreme first-passage events
This work addresses the core challenge of how a cell can rapidly and accurately determine the direction of a chemoattractant source when the signal is...
-
SDSR: A Spectral Divide-and-Conquer Approach for Species Tree Reconstruction
This paper addresses the computational bottleneck in reconstructing species trees from thousands of species and multiple genes by introducing a scalab...
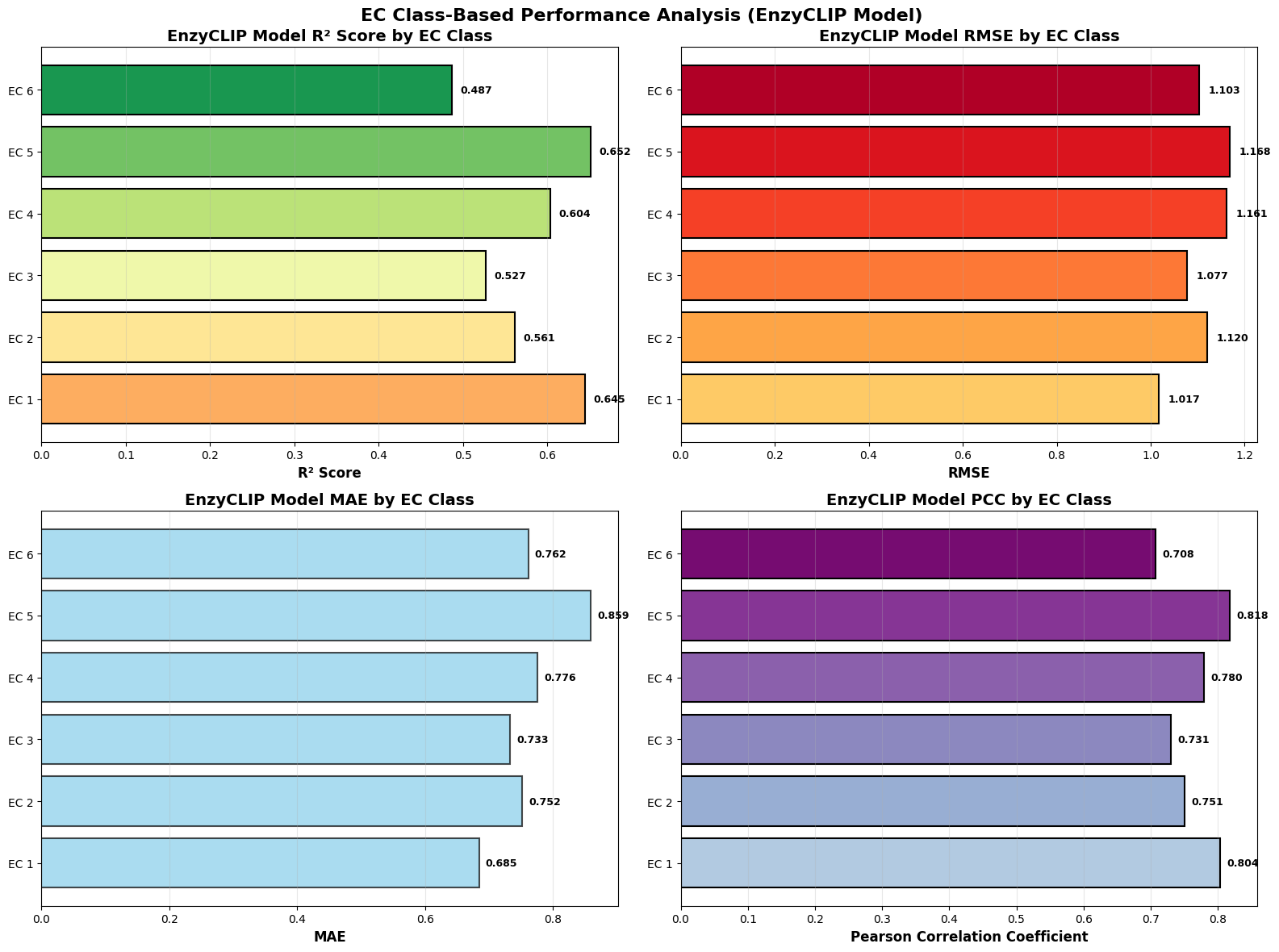
EnzyCLIP: A Cross-Attention Dual Encoder Framework with Contrastive Learning for Predicting Enzyme Kinetic Constants
Vellore Institute of Technology | BIT (Department of Computer Science) | BIT (Department of Bioengineering and Biotechnology)
30秒速读
IN SHORT: This paper addresses the core challenge of jointly predicting enzyme kinetic parameters (Kcat and Km) by modeling dynamic enzyme-substrate interactions through a multimodal contrastive learning framework.
核心创新
- Methodology Proposes a CLIP-inspired dual-encoder architecture with bidirectional cross-attention that dynamically models enzyme-substrate interactions, overcoming the limitation of separate processing in existing methods.
- Methodology Integrates contrastive learning (InfoNCE loss) with multi-task regression (Huber loss) to learn aligned multimodal representations while jointly predicting both Kcat and Km parameters.
- Biology Addresses the critical gap in existing literature that typically focuses on single parameter prediction (mainly Kcat) by providing a unified framework for joint prediction of both fundamental kinetic constants.
主要结论
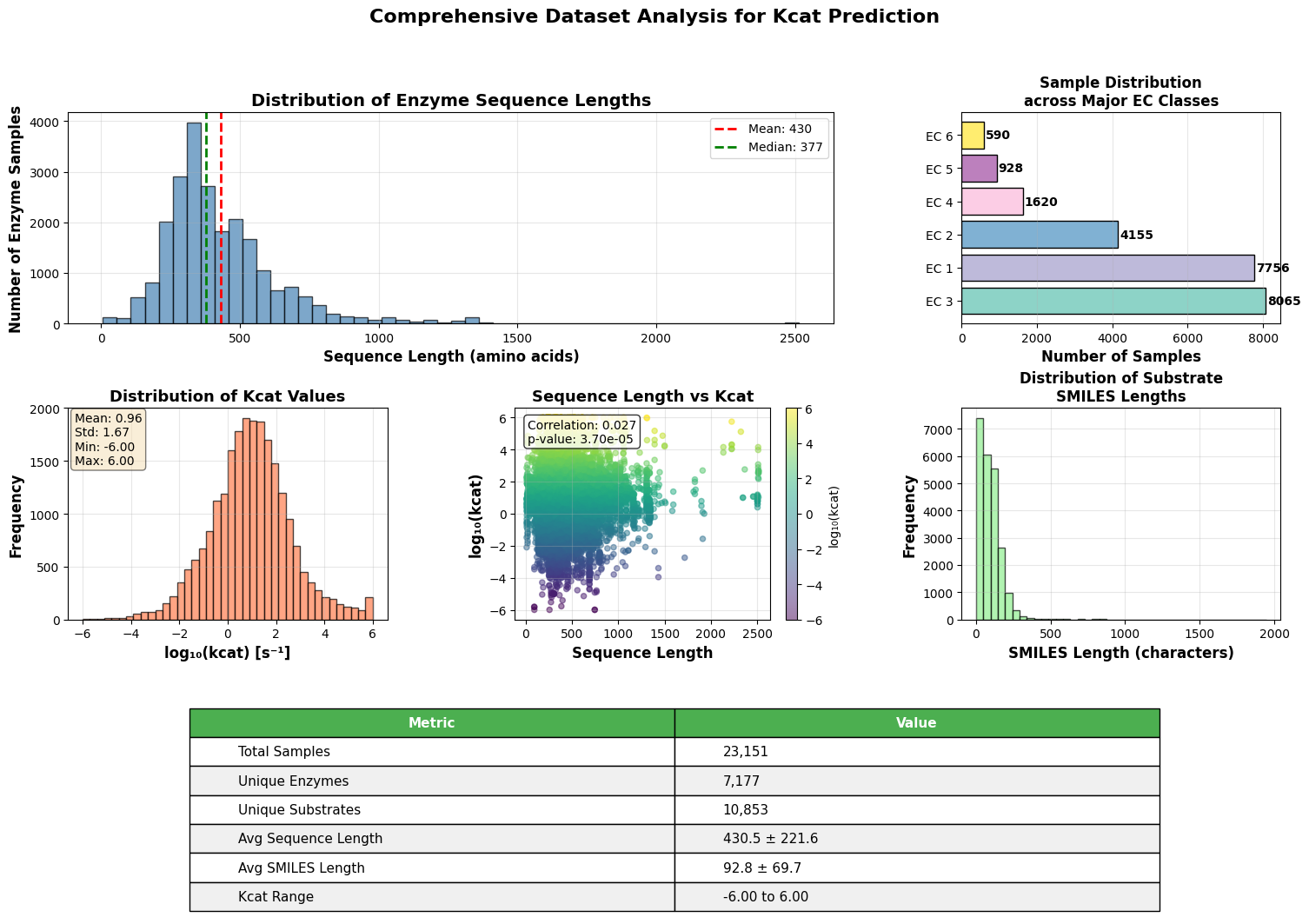
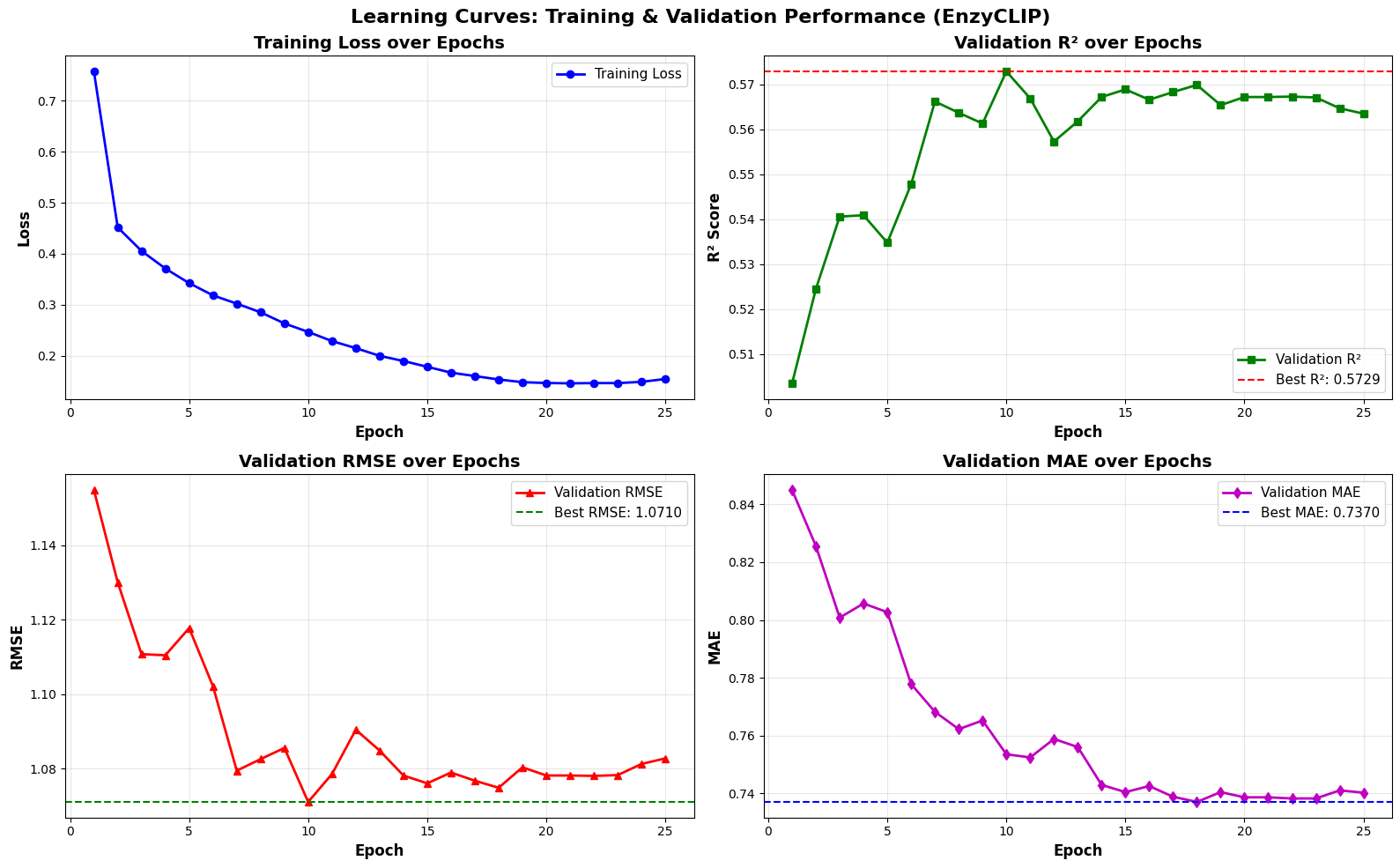
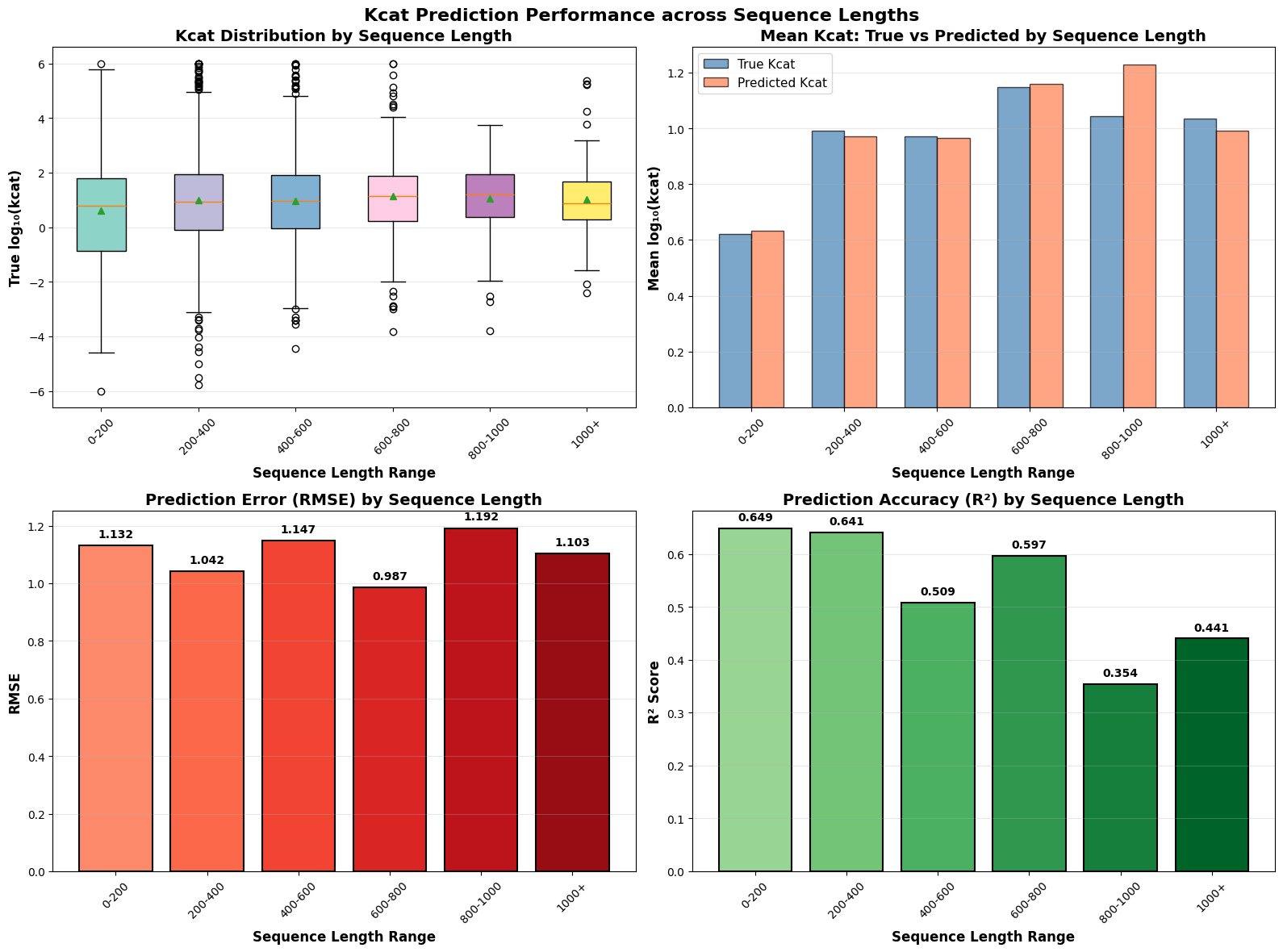
- EnzyCLIP achieves competitive baseline performance with R² scores of 0.593 for Kcat and 0.607 for Km prediction on the CatPred-DB dataset containing 23,151 Kcat and 41,174 Km measurements.
- The integration of contrastive learning with cross-attention mechanisms enables the model to capture biochemical relationships and substrate preferences even for unseen enzyme-substrate pairs.
- XGBoost ensemble methods applied to learned embeddings further improved Km prediction performance to R² = 0.61 while maintaining robust Kcat prediction capabilities.
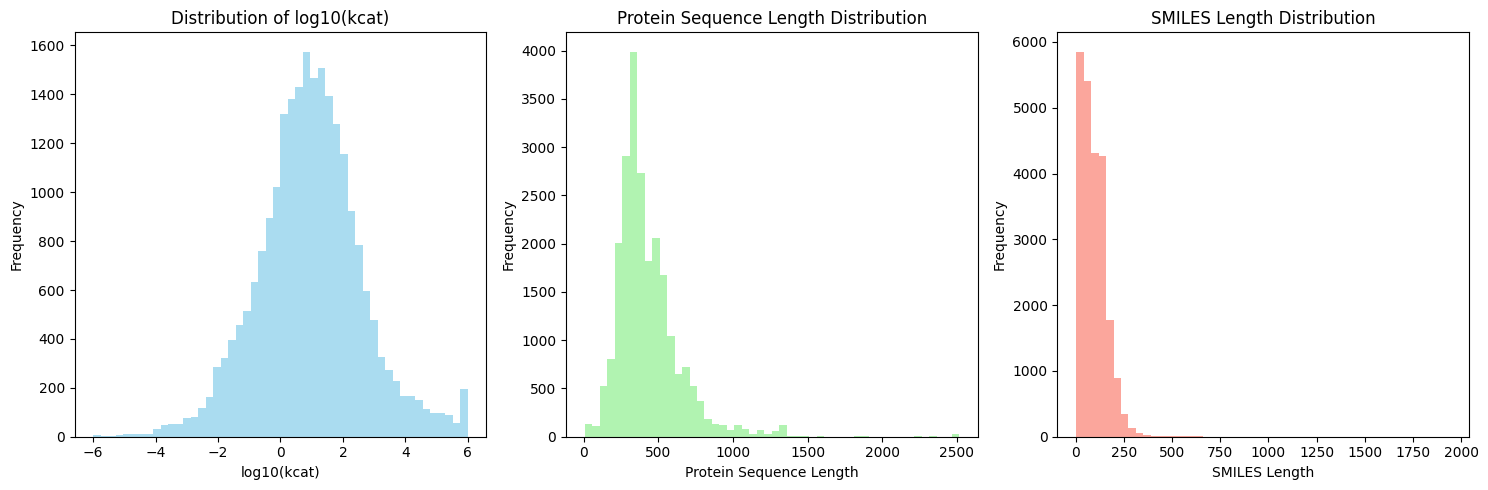
摘要: Accurate prediction of enzyme kinetic parameters is crucial for drug discovery, metabolic engineering, and synthetic biology applications. Current computational approaches face limitations in capturing complex enzyme–substrate interactions and often focus on single parameters while neglecting the joint prediction of catalytic turnover numbers (Kcat) and Michaelis–Menten constants (Km). We present EnzyCLIP, a novel dual-encoder framework that leverages contrastive learning and cross-attention mechanisms to predict enzyme kinetic parameters from protein sequences and substrate molecular structures. Our approach integrates ESM-2 protein language model embeddings with ChemBERTa chemical representations through a CLIP-inspired architecture enhanced with bidirectional cross-attention for dynamic enzyme–substrate interaction modeling. EnzyCLIP combines InfoNCE contrastive loss with Huber regression loss to learn aligned multimodal representations while predicting log10-transformed kinetic parameters. EnzyCLIP is trained on the CatPred-DB database containing 23,151 Kcat and 41,174 Km experimentally validated measurements, and achieved competitive baseline performance with R2 scores of 0.593 for Kcat and 0.607 for Km prediction. XGBoost ensemble methods on learned embeddings further improved Km prediction (R2 = 0.61) while maintaining robust Kcat performance.