Paper List
-
STAR-GO: Improving Protein Function Prediction by Learning to Hierarchically Integrate Ontology-Informed Semantic Embeddings
This paper addresses the core challenge of generalizing protein function prediction to unseen or newly introduced Gene Ontology (GO) terms by overcomi...
-
Incorporating indel channels into average-case analysis of seed-chain-extend
This paper addresses the core pain point of bridging the theoretical gap for the widely used seed-chain-extend heuristic by providing the first rigoro...
-
Competition, stability, and functionality in excitatory-inhibitory neural circuits
This paper addresses the core challenge of extending interpretable energy-based frameworks to biologically realistic asymmetric neural networks, where...
-
Enhancing Clinical Note Generation with ICD-10, Clinical Ontology Knowledge Graphs, and Chain-of-Thought Prompting Using GPT-4
This paper addresses the core challenge of generating accurate and clinically relevant patient notes from sparse inputs (ICD codes and basic demograph...
-
Learning From Limited Data and Feedback for Cell Culture Process Monitoring: A Comparative Study
This paper addresses the core challenge of developing accurate real-time bioprocess monitoring soft sensors under severe data constraints: limited his...
-
Cell-cell communication inference and analysis: biological mechanisms, computational approaches, and future opportunities
This review addresses the critical need for a systematic framework to navigate the rapidly expanding landscape of computational methods for inferring ...
-
Generating a Contact Matrix for Aged Care Settings in Australia: an agent-based model study
This study addresses the critical gap in understanding heterogeneous contact patterns within aged care facilities, where existing population-level con...
-
Emergent Spatiotemporal Dynamics in Large-Scale Brain Networks with Next Generation Neural Mass Models
This work addresses the core challenge of understanding how complex, brain-wide spatiotemporal patterns emerge from the interaction of biophysically d...
SSDLabeler: Realistic semi-synthetic data generation for multi-label artifact classification in EEG
Sony Computer Science Laboratories, Inc., Tokyo, Japan
30秒速读
IN SHORT: This paper addresses the core challenge of training robust multi-label EEG artifact classifiers by overcoming the scarcity and limited diversity of manually labeled training data through a novel semi-synthetic data generation framework.
核心创新
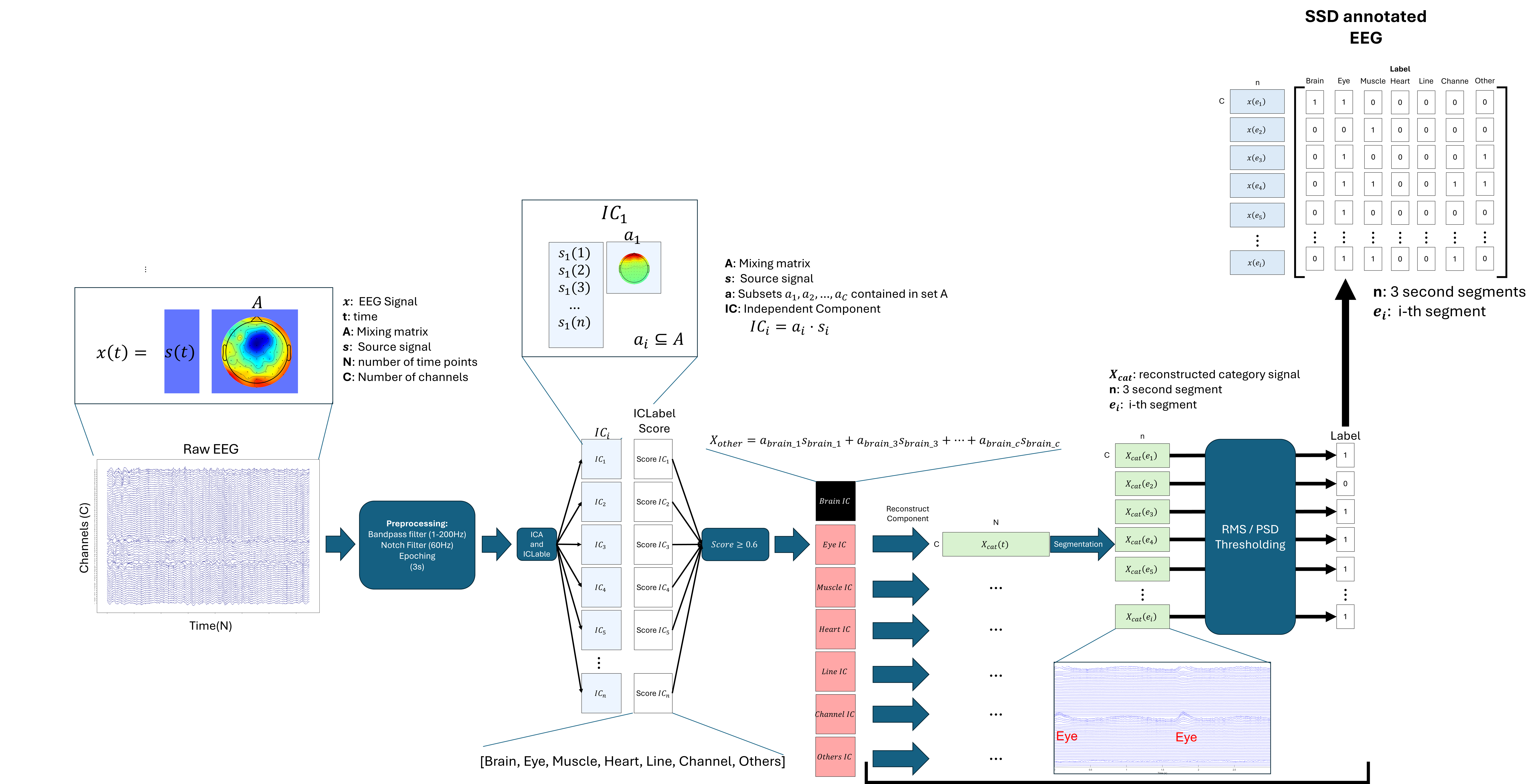
- Methodology Introduces SSDLabeler, a framework that generates realistic semi-synthetic EEG data by simultaneously reinjecting multiple ICA-isolated artifact types into clean data, preserving the co-occurrence structure of real-world contamination.
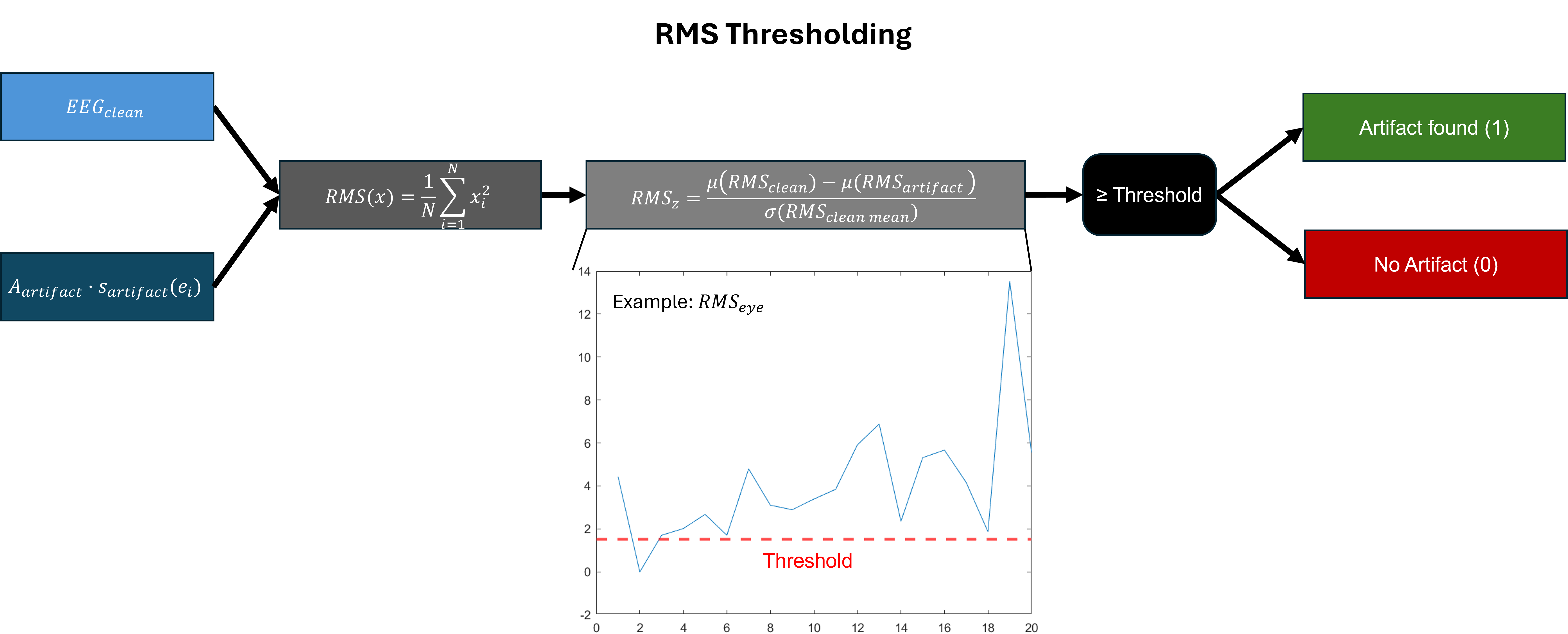
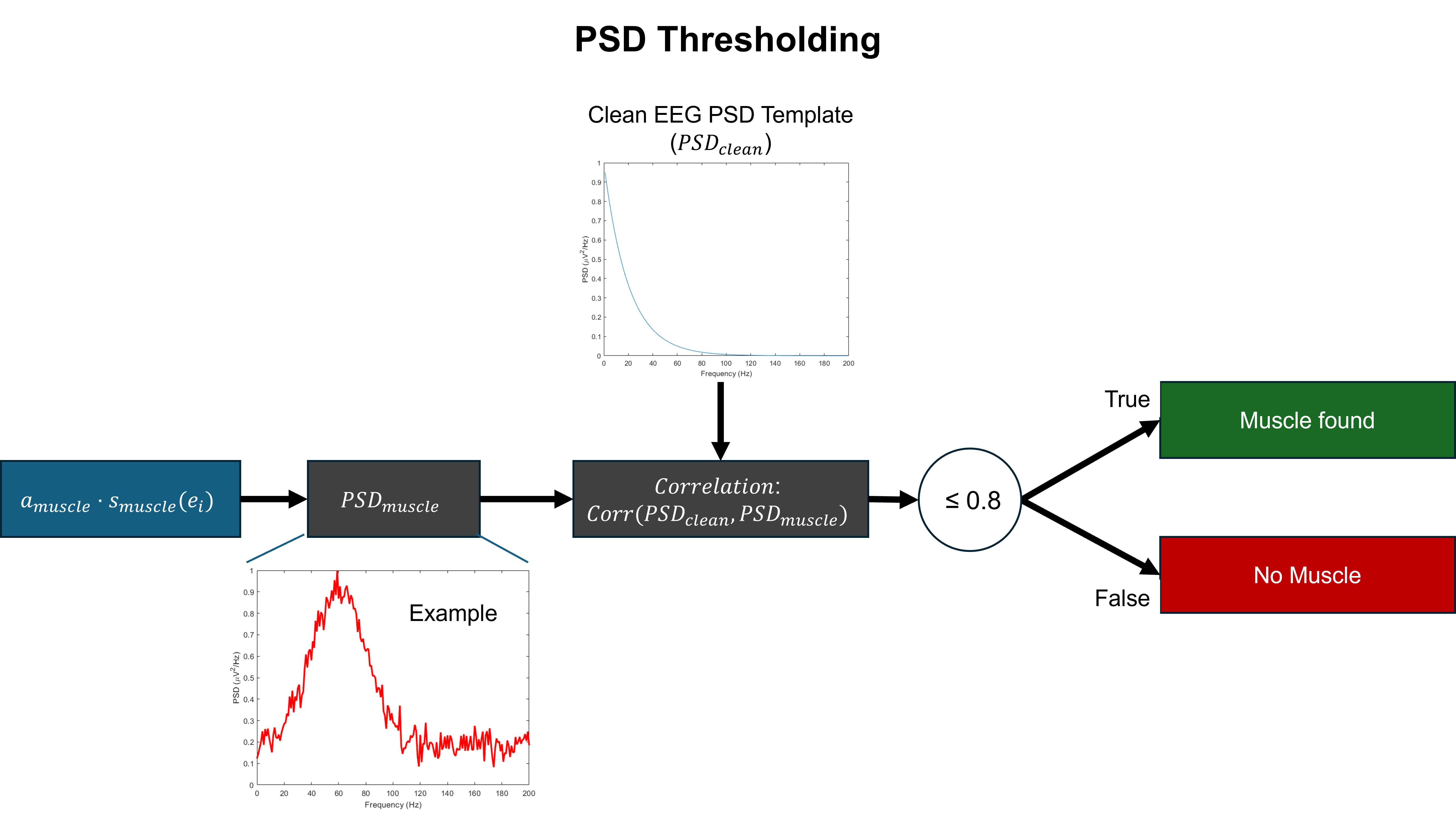
- Methodology Develops a novel artifact verification step using RMS and PSD thresholding criteria at the epoch level to ensure the physiological plausibility of generated contaminations, moving beyond simple ICA component injection.
- Biology Proposes a multi-label artifact classification paradigm that identifies multiple co-occurring artifact types (eye, muscle, heart, line, channel, other) within single EEG epochs, providing transparent contamination information for flexible preprocessing decisions.
主要结论
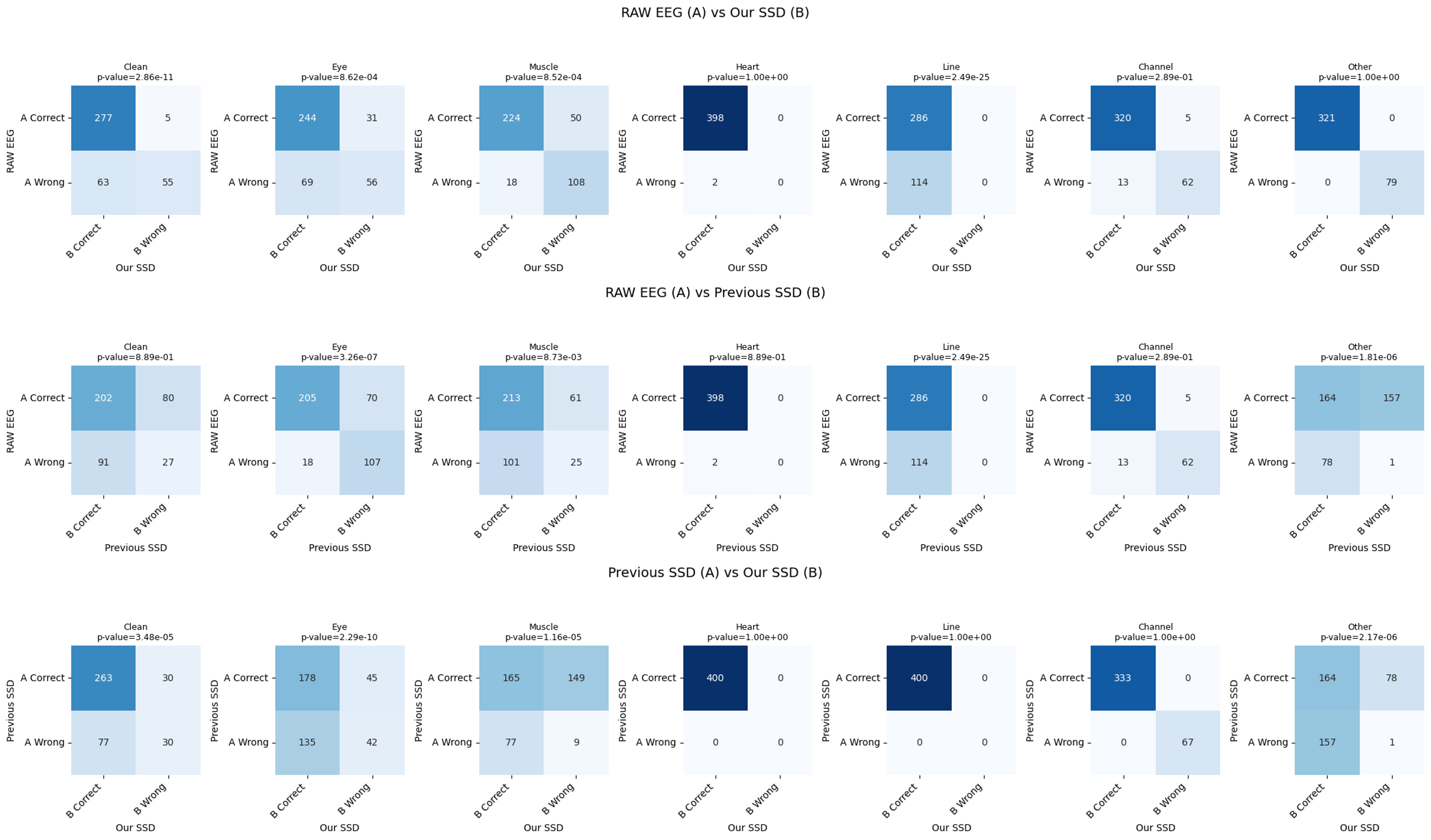
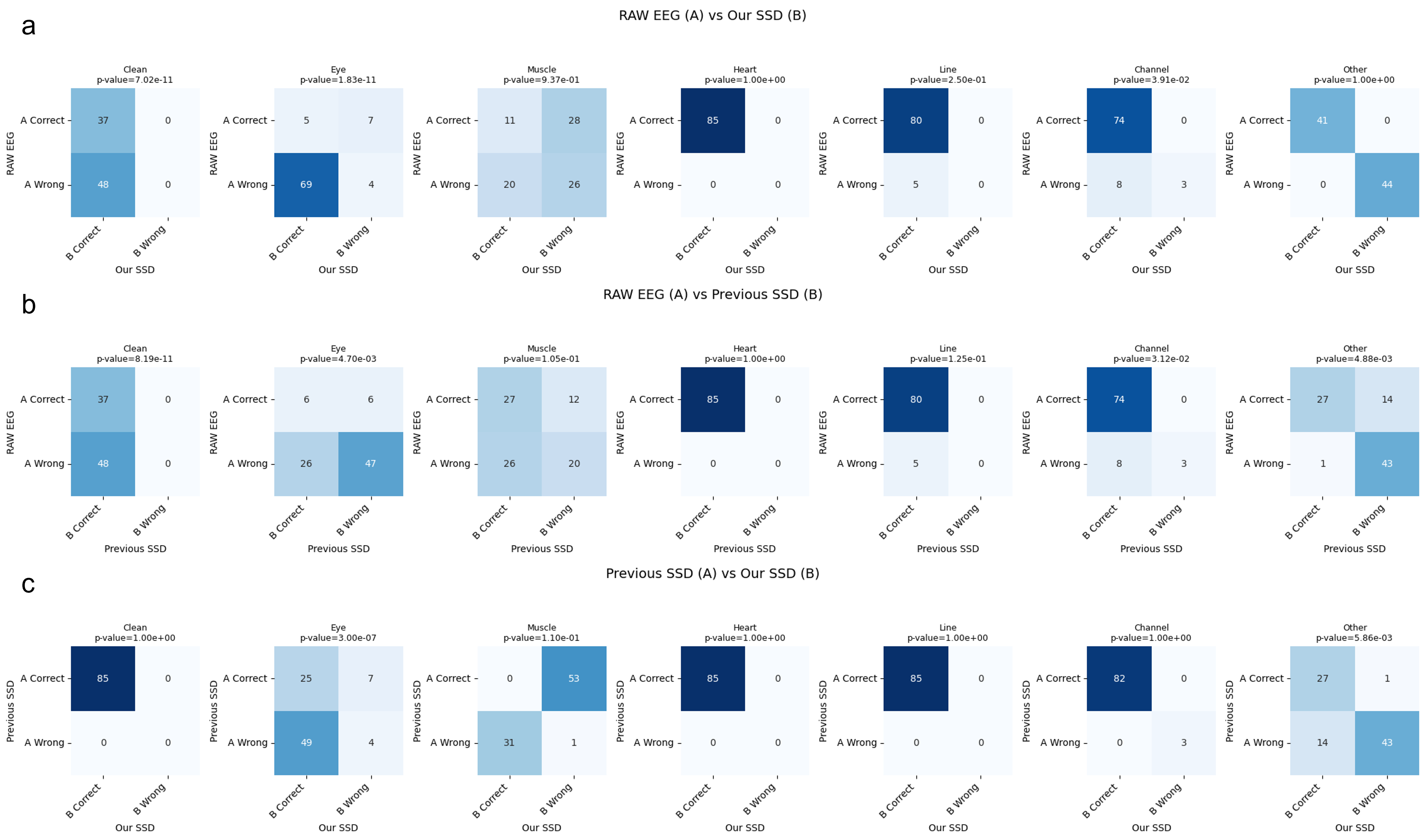
- SSDLabeler-trained classifiers achieved the highest overall accuracy (0.839) on motor execution test data, significantly outperforming raw EEG training (0.772, p<0.05 for Clean, Eye, and Line categories) and prior SSD methods (0.788).
- On instructed-noise session data, the proposed method achieved 0.812 accuracy, demonstrating strong generalization with significant improvements over raw EEG (0.618, p<0.05 for Clean, Eye, and Channel categories) and prior SSD (0.756).
- The framework successfully captures artifact co-occurrence, with the classifier showing balanced performance across most artifact types, though muscle artifact detection remained challenging (accuracy 0.605 vs. 0.785 for prior SSD).
摘要: EEG recordings are inherently contaminated by artifacts such as ocular, muscular, and environmental noise, which obscure neural activity and complicate preprocessing. Artifact classification offers advantages in stability and transparency, providing a viable alternative to ICA-based methods that enable flexible use alongside human inspections and across various applications. However, artifact classification is limited by its training data as it requires extensive manual labeling, which cannot fully cover the diversity of real-world EEG. Semi-synthetic data (SSD) methods have been proposed to address this limitation, but prior approaches typically injected single artifact types using ICA components or required separately recorded artifact signals, reducing both the realism of the generated data and the applicability of the method. To overcome these issues, we introduce SSDLabeler, a framework that generates realistic, annotated SSDs by decomposing real EEG with ICA, epoch-level artifact verification using RMS and PSD criteria, and reinjecting multiple artifact types into clean data. When applied to train a multi-label artifact classifier, it improved accuracy on raw EEG across diverse conditions compared to prior SSD and raw EEG training, establishing a scalable foundation for artifact handling that captures the co-occurrence and complexity of real EEG.