Paper List
-
STAR-GO: Improving Protein Function Prediction by Learning to Hierarchically Integrate Ontology-Informed Semantic Embeddings
This paper addresses the core challenge of generalizing protein function prediction to unseen or newly introduced Gene Ontology (GO) terms by overcomi...
-
Incorporating indel channels into average-case analysis of seed-chain-extend
This paper addresses the core pain point of bridging the theoretical gap for the widely used seed-chain-extend heuristic by providing the first rigoro...
-
Competition, stability, and functionality in excitatory-inhibitory neural circuits
This paper addresses the core challenge of extending interpretable energy-based frameworks to biologically realistic asymmetric neural networks, where...
-
Enhancing Clinical Note Generation with ICD-10, Clinical Ontology Knowledge Graphs, and Chain-of-Thought Prompting Using GPT-4
This paper addresses the core challenge of generating accurate and clinically relevant patient notes from sparse inputs (ICD codes and basic demograph...
-
Learning From Limited Data and Feedback for Cell Culture Process Monitoring: A Comparative Study
This paper addresses the core challenge of developing accurate real-time bioprocess monitoring soft sensors under severe data constraints: limited his...
-
Cell-cell communication inference and analysis: biological mechanisms, computational approaches, and future opportunities
This review addresses the critical need for a systematic framework to navigate the rapidly expanding landscape of computational methods for inferring ...
-
Generating a Contact Matrix for Aged Care Settings in Australia: an agent-based model study
This study addresses the critical gap in understanding heterogeneous contact patterns within aged care facilities, where existing population-level con...
-
Emergent Spatiotemporal Dynamics in Large-Scale Brain Networks with Next Generation Neural Mass Models
This work addresses the core challenge of understanding how complex, brain-wide spatiotemporal patterns emerge from the interaction of biophysically d...
Exactly Solvable Population Model with Square-Root Growth Noise and Cell-Size Regulation
Institute for Theoretical Physics, Department of Physics, Utrecht University, Utrecht, Netherlands | Centre for Complex Systems Studies, Utrecht University, Utrecht, Netherlands
30秒速读
IN SHORT: This paper addresses the fundamental gap in understanding how microscopic growth fluctuations, specifically those with size-dependent (square-root) noise, shape population-level fitness and statistics in cell populations, providing an exactly solvable model that contrasts sharply with existing size-independent noise models.
核心创新
- Theory Demonstrates that the asymptotic population growth rate Λ is exactly equal to the mean single-cell growth rate k, independent of noise strength σ and division mechanisms, establishing square-root growth noise as neutral for long-term fitness.
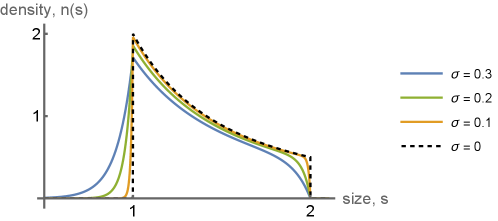
- Methodology Derives exact, closed-form expressions for the steady-state snapshot cell-size distribution, showing it results from a universal one-sided exponential convolution of the deterministic inverse-square-law solution, with kernel width σ².
- Theory Proves that the mean-rescaled population size Nt/⟨Nt⟩ converges to a stationary compound Poisson–exponential distribution determined solely by the growth noise parameter σ, independent of division or partitioning noise.
主要结论
- Population growth rate Λ = k exactly, demonstrating fitness neutrality of square-root noise (contrasting with models where Λ increases with variance of size-independent noise).
- Steady-state population mean cell size shifts by -σ² (e.g., ⟨s⟩pop = 2ln2 - σ² + O(e^{-1/σ²})), while variance is modified only at order σ⁴, showing a hierarchy of decoupling.
- The coefficient of variation of total cell number saturates to √(2σ²), and the full distribution of the mean-rescaled population size is a compound Poisson–exponential, providing concrete, testable signatures.
摘要: We analyze a size-structured branching process in which individual cells grow exponentially according to a Feller square-root process and divide under general size-control mechanisms. We obtain exact expressions for the asymptotic population growth rate, the steady-state snapshot distribution of cell sizes, and the fluctuations of the total cell number. Our first result is that the population growth rate is exactly equal to the mean single-cell growth rate, for all noise strengths and for all division and size-regulation schemes that maintain size homeostasis. Thus square-root growth noise is neutral with respect to long-term fitness, in sharp contrast to models with size-independent stochastic growth rates. Second, we show that the steady-state population cell-size distribution is obtained from the deterministic inverse-square-law solution by a one-sided exponential convolution with kernel width set by the strength of growth fluctuations. Third, the mean-rescaled population size Nt/⟨Nt⟩ converges to a stationary compound Poisson–exponential distribution that depends only on growth noise. This distribution, and hence the long-time shape of population-size fluctuations, is unchanged by division-size noise or asymmetric partitioning. These results identify Feller-type exponential growth with square-root noise as an exactly solvable benchmark for stochastic growth in size-controlled populations and provide concrete signatures that distinguish it from models with size-independent growth-rate noise.