Paper List
-
Autonomous Agents Coordinating Distributed Discovery Through Emergent Artifact Exchange
This paper addresses the fundamental limitation of current AI-assisted scientific research by enabling truly autonomous, decentralized investigation w...
-
D-MEM: Dopamine-Gated Agentic Memory via Reward Prediction Error Routing
This paper addresses the fundamental scalability bottleneck in LLM agentic memory systems: the O(N²) computational complexity and unbounded API token ...
-
Countershading coloration in blue shark skin emerges from hierarchically organized and spatially tuned photonic architectures inside skin denticles
This paper solves the core problem of how blue sharks achieve their striking dorsoventral countershading camouflage, revealing that coloration origina...
-
Human-like Object Grouping in Self-supervised Vision Transformers
This paper addresses the core challenge of quantifying how well self-supervised vision models capture human-like object grouping in natural scenes, br...
-
Hierarchical pp-Adic Framework for Gene Regulatory Networks: Theory and Stability Analysis
This paper addresses the core challenge of mathematically capturing the inherent hierarchical organization and multi-scale stability of gene regulator...
-
Towards unified brain-to-text decoding across speech production and perception
This paper addresses the core challenge of developing a unified brain-to-text decoding framework that works across both speech production and percepti...
-
Dual-Laws Model for a theory of artificial consciousness
This paper addresses the core challenge of developing a comprehensive, testable theory of consciousness that bridges biological and artificial systems...
-
Pulse desynchronization of neural populations by targeting the centroid of the limit cycle in phase space
This work addresses the core challenge of determining optimal pulse timing and intensity for desynchronizing pathological neural oscillations when the...
Hierarchical Molecular Language Models (HMLMs)
Department of Chemical Engineering, University of Arkansas, Fayetteville, AR 72701, USA
30秒速读
IN SHORT: This paper addresses the core challenge of accurately modeling context-dependent signaling, pathway cross-talk, and temporal dynamics across multiple biological scales in cellular signaling networks.
核心创新
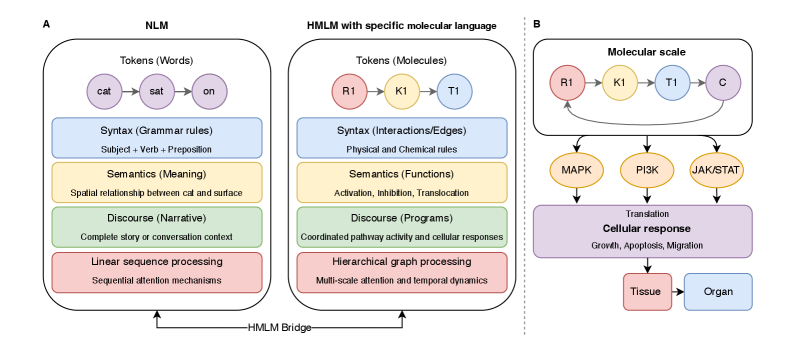
- Methodology Introduces cellular signaling as a molecular language with unique grammar and semantics, establishing a theoretical foundation for molecular artificial intelligence (MAI).
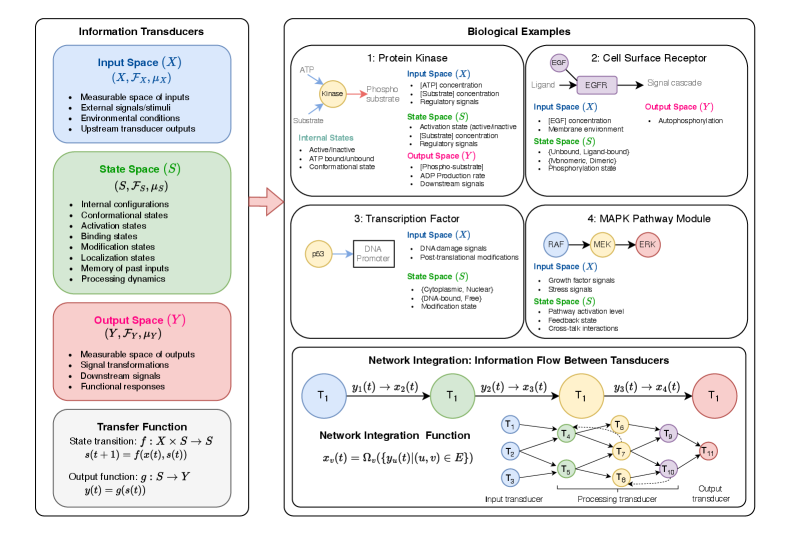
- Methodology Develops HMLMs as a novel computational architecture adapting transformer architecture to model signaling networks as information-processing systems across molecular, pathway, and cellular scales.
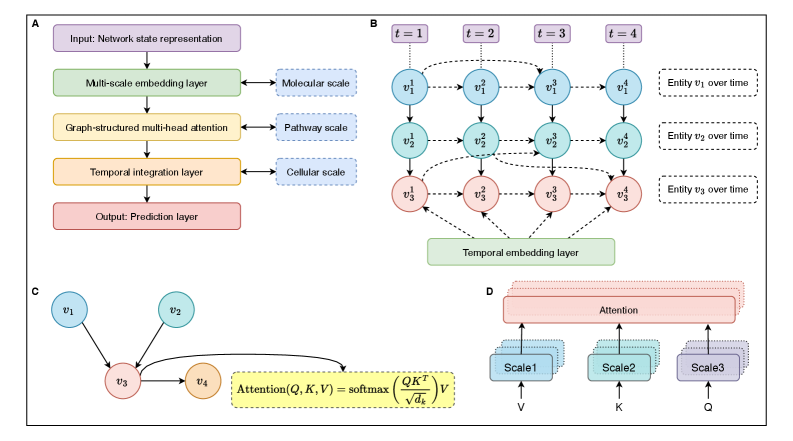
- Methodology Implements graph-structured attention mechanisms and hierarchical scale-bridging operators (aggregation, decomposition, translation) to accommodate signaling network topology and multi-scale organization.
主要结论
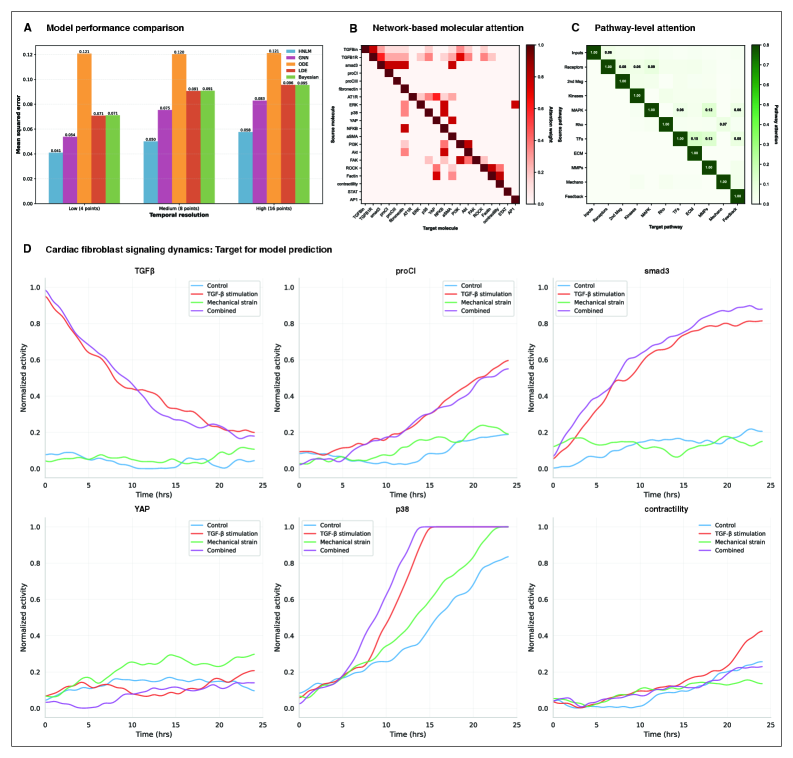
- HMLM achieved MSE of 0.058 for temporal signaling predictions, representing 30% improvement over GNNs (0.083) and 52% improvement over ODE models (0.121).
- Under sparse temporal sampling with only 4 timepoints, HMLM maintained superior performance with MSE = 0.041, demonstrating robustness to limited temporal data.
- Attention mechanisms identified biologically plausible pathway interactions including mechanotransduction-MAPK coupling and TGFβ to ERK signaling, validating the model's ability to capture meaningful biological relationships.
摘要: Cellular signaling networks represent complex information processing systems that have been modeled via traditional mathematical or statistical approaches. However, these methods often struggle to capture context-dependent signaling, pathway cross-talk, and temporal dynamics across multiple biological scales. Here, we introduce hierarchical molecular language models (HMLMs), a novel architecture that proposes a molecular network-specifiac large language model (LLM) to use in intracellular communication as a specialized molecular language, which includes molecules as tokens, protein interactions, post-translational modifications, and regulatory events modeled as semantic relationships within an adapted transformer architecture. HMLMs employ graph-structured attention mechanisms to accommodate signaling network topology while integrating information across the molecular, pathway, and cellular scales through hierarchical attention patterns. We demonstrate HMLM superiority using a cardiac fibroblast signaling network comprising over 100 molecular species across functional modules connected by regulatory edges. HMLM achieved a mean squared error (MSE) of 0.058 for temporal signaling predictions, representing 30% improvement over graph neural networks (GNNs: 0.083) and 52% improvement over ordinary differential equation models (ODEs: 0.121), with particular advantages under sparse temporal sampling conditions where HMLM maintained MSE = 0.041 with only 4 timepoints. The attention-based computational analysis identified key inter-pathway cross-talk patterns through learned attention mechanisms, including mechanotransduction-MAPK coupling and TGFβ to ERK signaling, demonstrating the model's capability to capture biologically plausible pathway interactions from network topology and temporal dynamics and convergent regulatory mechanisms controlling fibrosis markers in simulated cardiac fibroblast networks. The HMLMs offer a foundation for AI-driven biology and medicine with predictable scaling characteristics suitable for interactive applications. By bridging molecular mechanisms with cellular phenotypes through AI-driven molecular language representation, HMLMs provide a powerful paradigm for systems biology that advances precision medicine applications and therapeutic discovery in the era of AI.