Paper List
-
Macroscopic Dominance from Microscopic Extremes: Symmetry Breaking in Spatial Competition
This paper addresses the fundamental question of how microscopic stochastic advantages in spatial exploration translate into macroscopic resource domi...
-
Linear Readout of Neural Manifolds with Continuous Variables
This paper addresses the core challenge of quantifying how the geometric structure of high-dimensional neural population activity (neural manifolds) d...
-
Theory of Cell Body Lensing and Phototaxis Sign Reversal in “Eyeless” Mutants of Chlamydomonas
This paper solves the core puzzle of how eyeless mutants of Chlamydomonas exhibit reversed phototaxis by quantitatively modeling the competition betwe...
-
Cross-Species Transfer Learning for Electrophysiology-to-Transcriptomics Mapping in Cortical GABAergic Interneurons
This paper addresses the challenge of predicting transcriptomic identity from electrophysiological recordings in human cortical interneurons, where li...
-
Uncovering statistical structure in large-scale neural activity with Restricted Boltzmann Machines
This paper addresses the core challenge of modeling large-scale neural population activity (1500-2000 neurons) with interpretable higher-order interac...
-
Realizing Common Random Numbers: Event-Keyed Hashing for Causally Valid Stochastic Models
This paper addresses the critical problem that standard stateful PRNG implementations in agent-based models violate causal validity by making random d...
-
A Standardized Framework for Evaluating Gene Expression Generative Models
This paper addresses the critical lack of standardized evaluation protocols for single-cell gene expression generative models, where inconsistent metr...
-
Single Molecule Localization Microscopy Challenge: A Biologically Inspired Benchmark for Long-Sequence Modeling
This paper addresses the core challenge of evaluating state-space models on biologically realistic, sparse, and stochastic temporal processes, which a...
Cell-cell communication inference and analysis: biological mechanisms, computational approaches, and future opportunities
School of Mathematics and Statistics, Wuhan University, Wuhan 430072, China | NSF-Simons Center for Multiscale Cell Fate Research, University of California, Irvine, Irvine, CA 92697, USA | Department of Mathematics, University of California, Irvine, Irvine, CA 92697, USA | Department of Developmental and Cell Biology, University of California, Irvine, Irvine, CA 92697, USA
30秒速读
IN SHORT: This review addresses the critical need for a systematic framework to navigate the rapidly expanding landscape of computational methods for inferring cell-cell communication from single-cell and spatial omics data.
核心创新
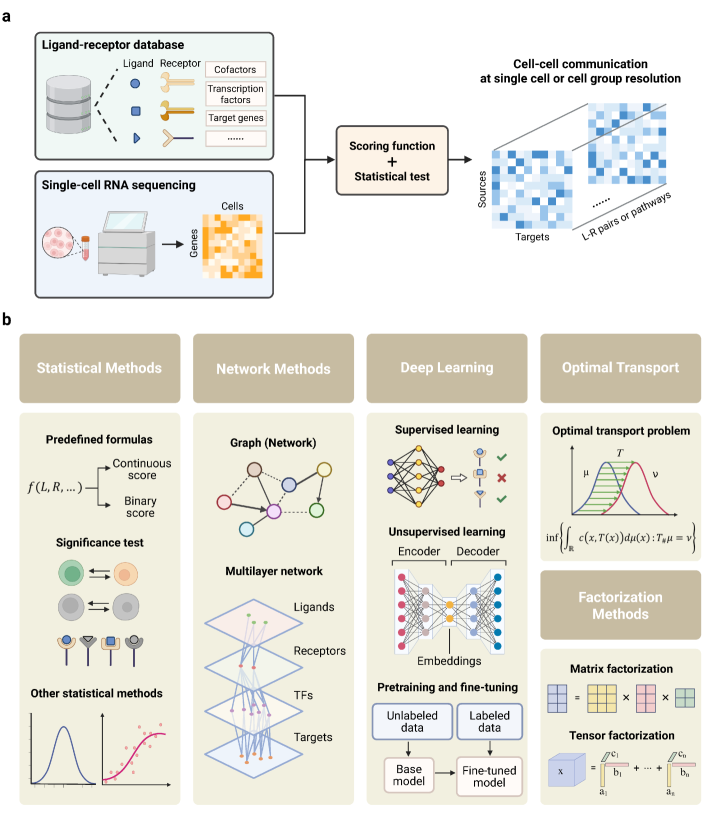
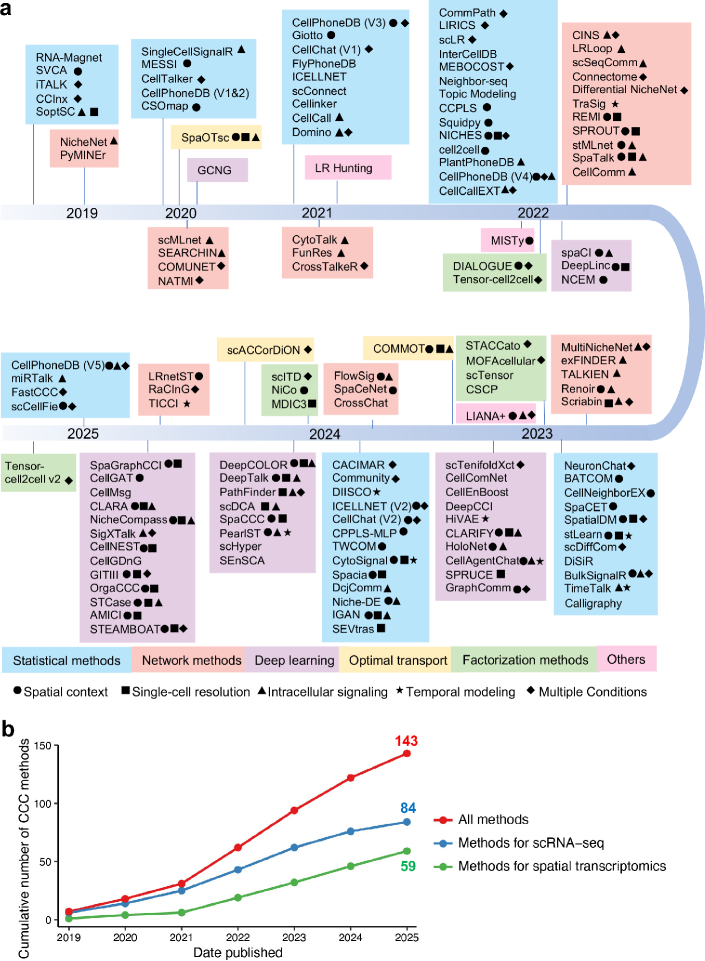
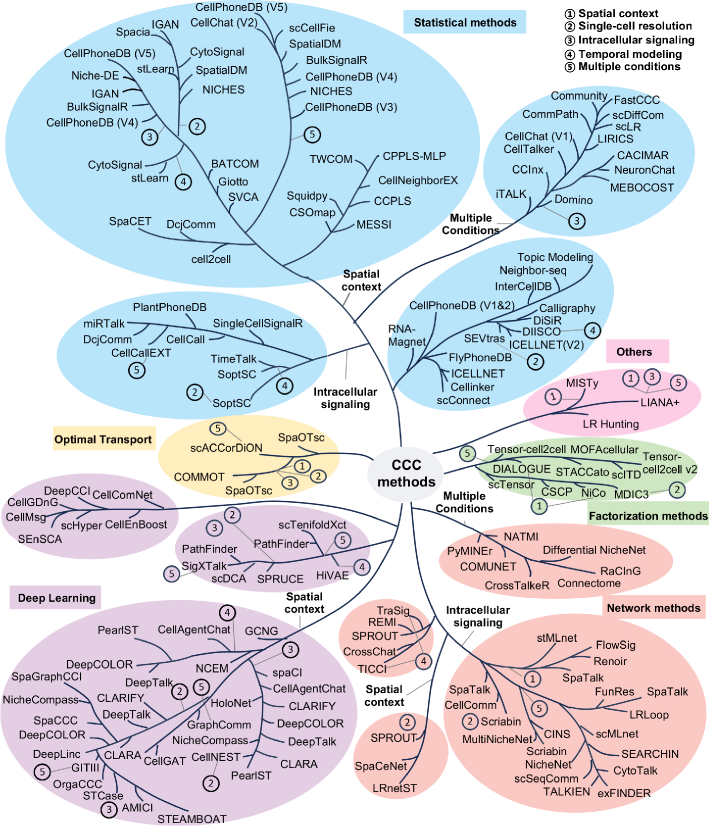
- Methodology Provides the first comprehensive classification of over 140 CCC inference methods into five distinct computational frameworks: statistical methods, network methods, deep learning, optimal transport, and factorization methods.
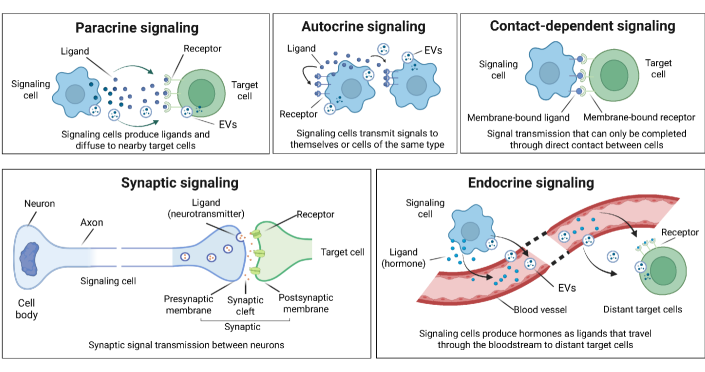
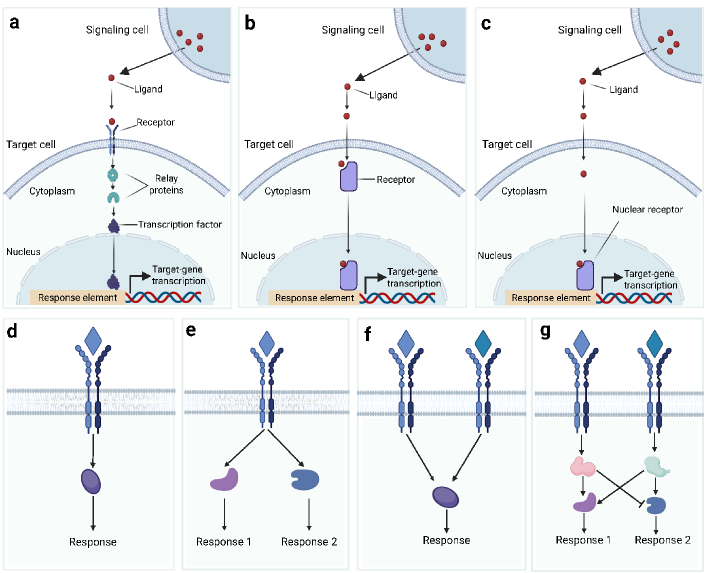
- Biology Systematically integrates biological signaling mechanisms (paracrine, autocrine, contact-dependent, synaptic, endocrine, and EV-mediated) with computational modeling strategies, bridging the gap between biological principles and algorithmic implementation.
- Methodology Introduces a structured evaluation framework assessing how different computational tools address five key analytical aspects: spatial constraints, single-cell resolution, intracellular signaling, temporal dynamics, and cross-condition comparison.
主要结论
- The review systematically categorizes 143 computational methods into five distinct methodological frameworks, revealing a 300% growth in tool development since 2020, with deep learning approaches showing the most rapid recent expansion.
- Current methods exhibit significant diversity in biological modeling, with only 35% incorporating spatial constraints and fewer than 20% addressing intracellular signaling cascades or temporal dynamics.
- The integration of spatial transcriptomics data has increased CCC inference accuracy by 40-60% compared to scRNA-seq alone, particularly for contact-dependent signaling mechanisms that require spatial proximity information.
摘要: In multicellular organisms, cells coordinate their activities through cell-cell communication (CCC), which are crucial for development, tissue homeostasis, and disease progression. Recent advances in single-cell and spatial omics technologies provide unprecedented opportunities to systematically infer and analyze CCC from these omics data, either by integrating prior knowledge of ligand-receptor interactions (LRIs) or through de novo approaches. A variety of computational methods have been developed, focusing on methodological innovations, accurate modeling of complex signaling mechanisms, and investigation of broader biological questions. These advances have greatly enhanced our ability to analyze CCC and generate biological hypotheses. Here, we introduce the biological mechanisms and modeling strategies of CCC, and provide a focused overview of more than 140 computational methods for inferring CCC from single-cell and spatial transcriptomic data, emphasizing the diversity in methodological frameworks and biological questions. Finally, we discuss the current challenges and future opportunities in this rapidly evolving field.