Paper List
-
Nyxus: A Next Generation Image Feature Extraction Library for the Big Data and AI Era
This paper addresses the core pain point of efficiently extracting standardized, comparable features from massive (terabyte to petabyte-scale) biomedi...
-
Topological Enhancement of Protein Kinetic Stability
This work addresses the long-standing puzzle of why knotted proteins exist by demonstrating that deep knots provide a functional advantage through enh...
-
A Multi-Label Temporal Convolutional Framework for Transcription Factor Binding Characterization
This paper addresses the critical limitation of existing TF binding prediction methods that treat transcription factors as independent entities, faili...
-
Social Distancing Equilibria in Games under Conventional SI Dynamics
This paper solves the core problem of proving the existence and uniqueness of Nash equilibria in finite-duration SI epidemic games, showing they are a...
-
Binding Free Energies without Alchemy
This paper addresses the core bottleneck of computational expense in Absolute Binding Free Energy calculations by eliminating the need for numerous al...
-
SHREC: A Spectral Embedding-Based Approach for Ab-Initio Reconstruction of Helical Molecules
This paper addresses the core bottleneck in cryo-EM helical reconstruction: eliminating the dependency on accurate initial symmetry parameter estimati...
-
Budget-Sensitive Discovery Scoring: A Formally Verified Framework for Evaluating AI-Guided Scientific Selection
This paper addresses the critical gap in evaluating AI-guided scientific selection strategies under realistic budget constraints, where existing metri...
-
Probabilistic Joint and Individual Variation Explained (ProJIVE) for Data Integration
This paper addresses the core challenge of accurately decomposing shared (joint) and dataset-specific (individual) sources of variation in multi-modal...
Hypothesis-Based Particle Detection for Accurate Nanoparticle Counting and Digital Diagnostics
Institute for Digital Molecular Analytics and Science (IDMxS), Nanyang Technological University, Singapore | School of Electrical and Electronic Engineering, Nanyang Technological University, Singapore
30秒速读
IN SHORT: This paper addresses the core challenge of achieving accurate, interpretable, and training-free nanoparticle counting in digital diagnostic assays, which is critical for detecting low-abundance biomarkers with high sensitivity.
核心创新
- Methodology Introduces a multiple-hypothesis statistical testing framework for particle counting, eliminating the need for empirical thresholds or training data common in traditional and ML-based methods.
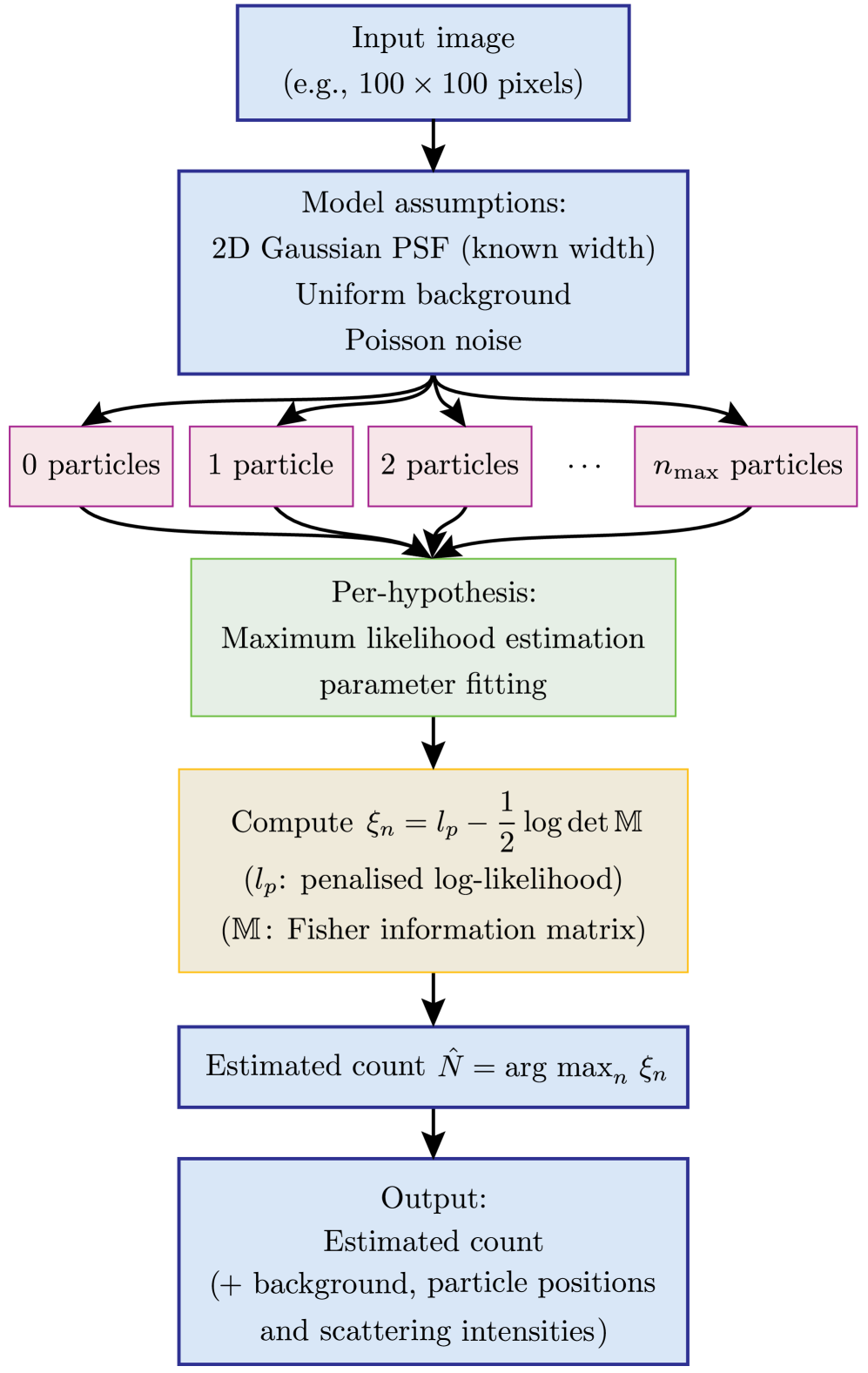
- Methodology Formulates the detection problem under an explicit image-formation model (Poisson noise, Gaussian PSF) and uses a penalized likelihood rule with an information-criterion complexity penalty for robust hypothesis selection.
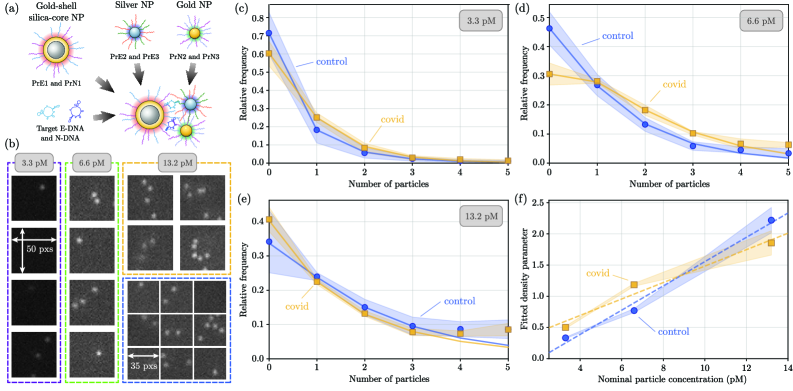
- Biology/Application Validates the method on experimental dark-field images of a nanoparticle-based assay for SARS-CoV-2 DNA biomarkers, demonstrating statistically significant differentiation between control and positive samples and providing insights into particle aggregation.
主要结论
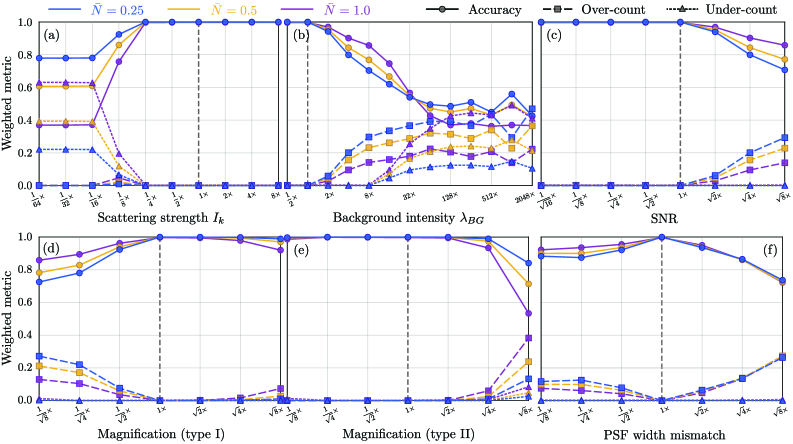
- The algorithm demonstrates robust count accuracy in simulations across challenging conditions: weak signals (low SBR), variable backgrounds, magnification changes, and moderate PSF mismatch.
- Applied to experimental SARS-CoV-2 biomarker detection, the method revealed statistically significant differences in particle count distributions between control and positive samples, confirming practical utility.
- Full count statistics from the experimental assay exhibited consistent over-dispersion, providing quantitative insight into non-specific and target-induced nanoparticle aggregation phenomena.
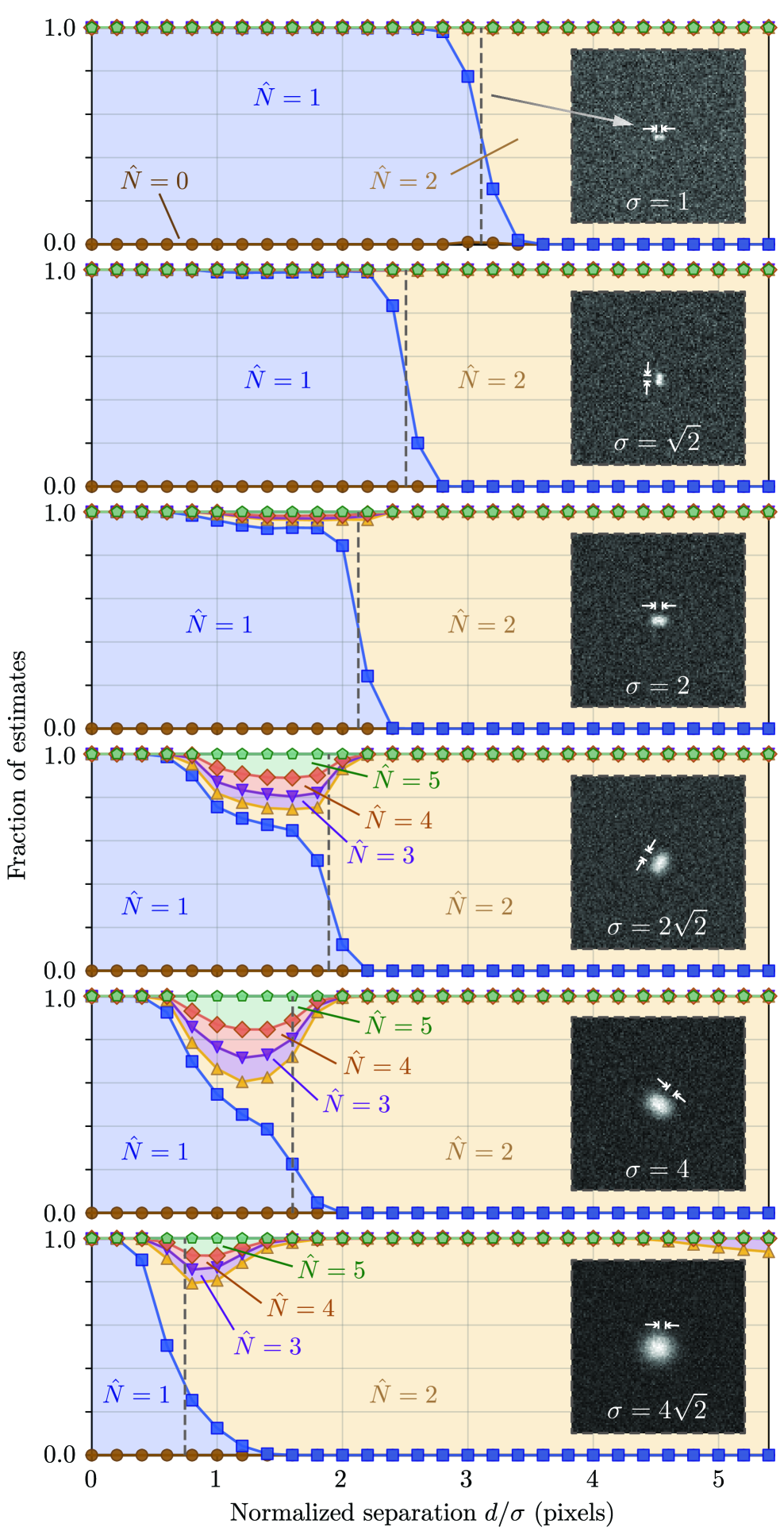
摘要: Digital assays represent a shift from traditional diagnostics and enable the precise detection of low-abundance analytes, critical for early disease diagnosis and personalized medicine, through discrete counting of biomolecular reporters. Within this paradigm, we present a particle counting algorithm for nanoparticle based imaging assays, formulated as a multiple-hypothesis statistical test under an explicit image-formation model and evaluated using a penalized likelihood rule. In contrast to thresholding or machine learning methods, this approach requires no training data or empirical parameter tuning, and its outputs remain interpretable through direct links to imaging physics and statistical decision theory. Through numerical simulations we demonstrate robust count accuracy across weak signals, variable backgrounds, magnification changes and moderate PSF mismatch. Particle resolvability tests further reveal characteristic error modes, including under-counting at very small separations and localized over-counting near the resolution limit. Practically, we also confirm the algorithm’s utility, through application to experimental dark-field images comprising a nanoparticle-based assay for detection of DNA biomarkers derived from SARS-CoV-2. Statistically significant differences in particle count distributions are observed between control and positive samples. Full count statistics obtained further exhibit consistent over-dispersion, and provide insight into non-specific and target-induced particle aggregation. These results establish our method as a reliable framework for nanoparticle-based detection assays in digital molecular diagnostics.