Paper List
-
GOPHER: Optimization-based Phenotype Randomization for Genome-Wide Association Studies with Differential Privacy
This paper addresses the core challenge of balancing rigorous privacy protection with data utility when releasing full GWAS summary statistics, overco...
-
Real-time Cricket Sorting By Sex A low-cost embedded solution using YOLOv8 and Raspberry Pi
This paper addresses the critical bottleneck in industrial insect farming: the lack of automated, real-time sex sorting systems for Acheta domesticus ...
-
Training Dynamics of Learning 3D-Rotational Equivariance
This work addresses the core dilemma of whether to use computationally expensive equivariant architectures or faster symmetry-agnostic models with dat...
-
Fast and Accurate Node-Age Estimation Under Fossil Calibration Uncertainty Using the Adjusted Pairwise Likelihood
This paper addresses the dual challenge of computational inefficiency and sensitivity to fossil calibration errors in Bayesian divergence time estimat...
-
Few-shot Protein Fitness Prediction via In-context Learning and Test-time Training
This paper addresses the core challenge of accurately predicting protein fitness with only a handful of experimental observations, where data collecti...
-
scCluBench: Comprehensive Benchmarking of Clustering Algorithms for Single-Cell RNA Sequencing
This paper addresses the critical gap of fragmented and non-standardized benchmarking in single-cell RNA-seq clustering, which hinders objective compa...
-
Simulation and inference methods for non-Markovian stochastic biochemical reaction networks
This paper addresses the computational bottleneck of simulating and performing Bayesian inference for non-Markovian biochemical systems with history-d...
-
Assessment of Simulation-based Inference Methods for Stochastic Compartmental Models
This paper addresses the core challenge of performing accurate Bayesian parameter inference for stochastic epidemic models when the likelihood functio...
STAR-GO: Improving Protein Function Prediction by Learning to Hierarchically Integrate Ontology-Informed Semantic Embeddings
Department of Computer Engineering, Bogazici University, Istanbul, Turkiye
30秒速读
IN SHORT: This paper addresses the core challenge of generalizing protein function prediction to unseen or newly introduced Gene Ontology (GO) terms by overcoming the limitations of existing models that either prioritize graph structure at the expense of semantic meaning or vice versa.
核心创新
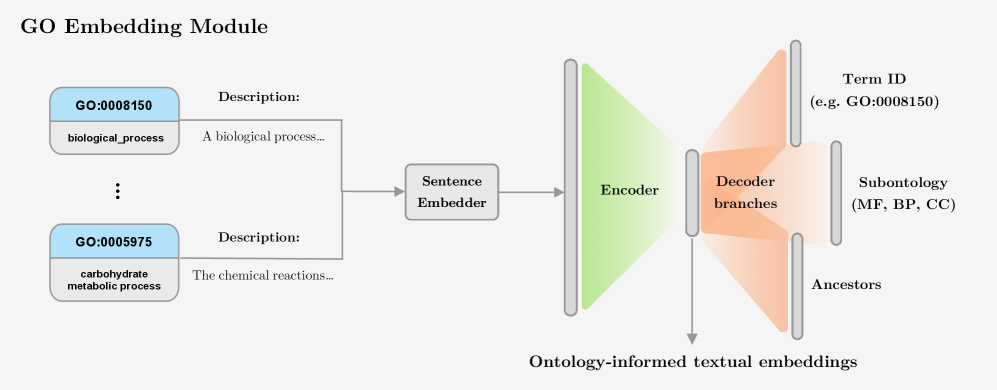
- Methodology Introduces a novel GO embedding module that integrates textual definitions (via SBERT-BioBERT) with ontology graph structure through a multi-task autoencoder, learning unified representations that preserve both semantic similarity and hierarchical dependencies.
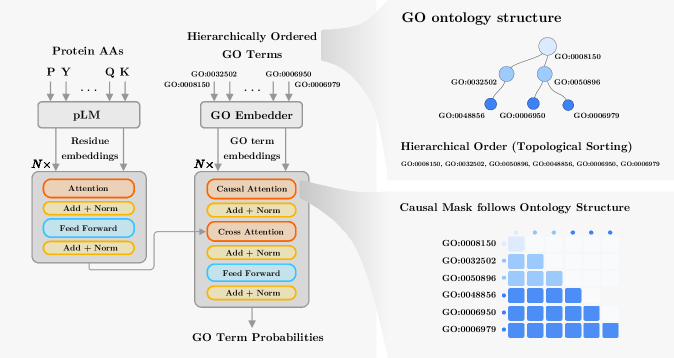
- Methodology Proposes a hierarchical Transformer decoder that processes GO terms in topological order (ancestors to descendants) using causal self-attention, enabling information propagation across ontology levels and capturing functional dependencies.
- Biology Demonstrates superior zero-shot generalization to unseen GO terms, particularly for Molecular Function and Biological Process terms, by effectively leveraging semantic information from textual definitions, which transfers better to novel ontology concepts than purely structural embeddings.
主要结论
- STAR-GO achieves state-of-the-art or competitive performance across all three GO subontologies (BP, CC, MF), with the highest AUC scores (e.g., 0.989 for BP, 0.988 for CC, 0.995 for MF), indicating strong term-level discriminability.
- In zero-shot evaluation on 16 held-out GO terms, STAR-GO variants achieve the highest AUCs in 13 cases, significantly outperforming baselines like DeepGOZero and DeepGO-SE, demonstrating superior generalization to unseen functions.
- Ablation studies reveal that semantic embeddings (STAR_T) achieve the best zero-shot results for most MF and BP terms (e.g., AUC of 0.949 for GO:0001228), while structural embeddings (STAR_S) perform best for a few terms but poorly for MF, highlighting the critical role of semantic information for generalization.
摘要: Motivation: Accurate prediction of protein function is essential for elucidating molecular mechanisms and advancing biological and therapeutic discovery. Yet experimental annotation lags far behind the rapid growth of protein sequence data. Computational approaches address this gap by associating proteins with Gene Ontology (GO) terms, which encode functional knowledge through hierarchical relations and textual definitions. However, existing models often emphasize one modality over the other, limiting their ability to generalize, particularly to unseen or newly introduced GO terms that frequently arise as the ontology evolves, and making the previously trained models outdated. Results: We present STAR-GO, a Transformer-based framework that jointly models the semantic and structural characteristics of GO terms to enhance zero-shot protein function prediction. STAR-GO integrates textual definitions with ontology graph structure to learn unified GO representations, which are processed in hierarchical order to propagate information from general to specific terms. These representations are then aligned with protein sequence embeddings to capture sequence–function relationships. STAR-GO achieves state-of-the-art performance and superior zero-shot generalization, demonstrating the utility of integrating semantics and structure for robust and adaptable protein function prediction. Availability: Code and pre-trained models are available at https://github.com/boun-tabi-lifelu/stargo.