Paper List
-
Ill-Conditioning in Dictionary-Based Dynamic-Equation Learning: A Systems Biology Case Study
This paper addresses the critical challenge of numerical ill-conditioning and multicollinearity in library-based sparse regression methods (e.g., SIND...
-
Hybrid eTFCE–GRF: Exact Cluster-Size Retrieval with Analytical pp-Values for Voxel-Based Morphometry
This paper addresses the computational bottleneck in voxel-based neuroimaging analysis by providing a method that delivers exact cluster-size retrieva...
-
abx_amr_simulator: A simulation environment for antibiotic prescribing policy optimization under antimicrobial resistance
This paper addresses the critical challenge of quantitatively evaluating antibiotic prescribing policies under realistic uncertainty and partial obser...
-
PesTwin: a biology-informed Digital Twin for enabling precision farming
This paper addresses the critical bottleneck in precision agriculture: the inability to accurately forecast pest outbreaks in real-time, leading to su...
-
Equivariant Asynchronous Diffusion: An Adaptive Denoising Schedule for Accelerated Molecular Conformation Generation
This paper addresses the core challenge of generating physically plausible 3D molecular structures by bridging the gap between autoregressive methods ...
-
Omics Data Discovery Agents
This paper addresses the core challenge of making published omics data computationally reusable by automating the extraction, quantification, and inte...
-
Single-cell directional sensing at ultra-low chemoattractant concentrations from extreme first-passage events
This work addresses the core challenge of how a cell can rapidly and accurately determine the direction of a chemoattractant source when the signal is...
-
SDSR: A Spectral Divide-and-Conquer Approach for Species Tree Reconstruction
This paper addresses the computational bottleneck in reconstructing species trees from thousands of species and multiple genes by introducing a scalab...
SpikGPT: A High-Accuracy and Interpretable Spiking Attention Framework for Single-Cell Annotation
Department of Biomedical Informatics, Emory University | Department of Surgery, Duke University
30秒速读
IN SHORT: This paper addresses the core challenge of robust single-cell annotation across heterogeneous datasets with batch effects and the critical need to identify previously unseen cell populations.
核心创新
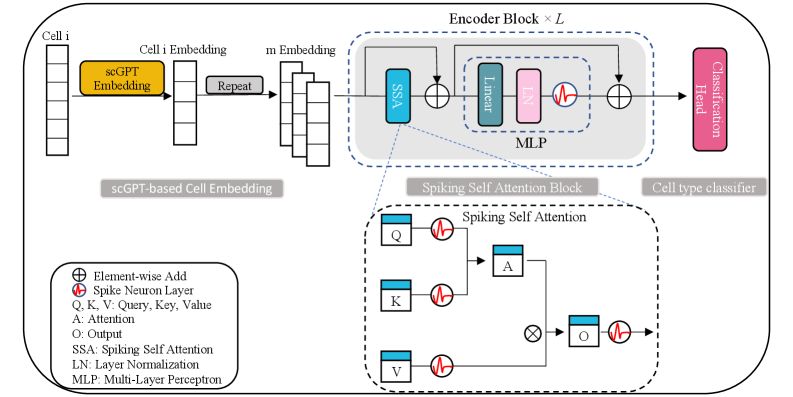
- Methodology First integration of spiking neural networks with transformer architecture for single-cell analysis, using Leaky Integrate-and-Fire (LIF) neurons in a multi-head Spiking Self-Attention mechanism for energy-efficient computation.
- Methodology Novel two-step embedding expansion strategy: repeating cell embeddings along feature channels (default m=300) and temporal dimensions (default T=4) to enhance representation richness and training stability.
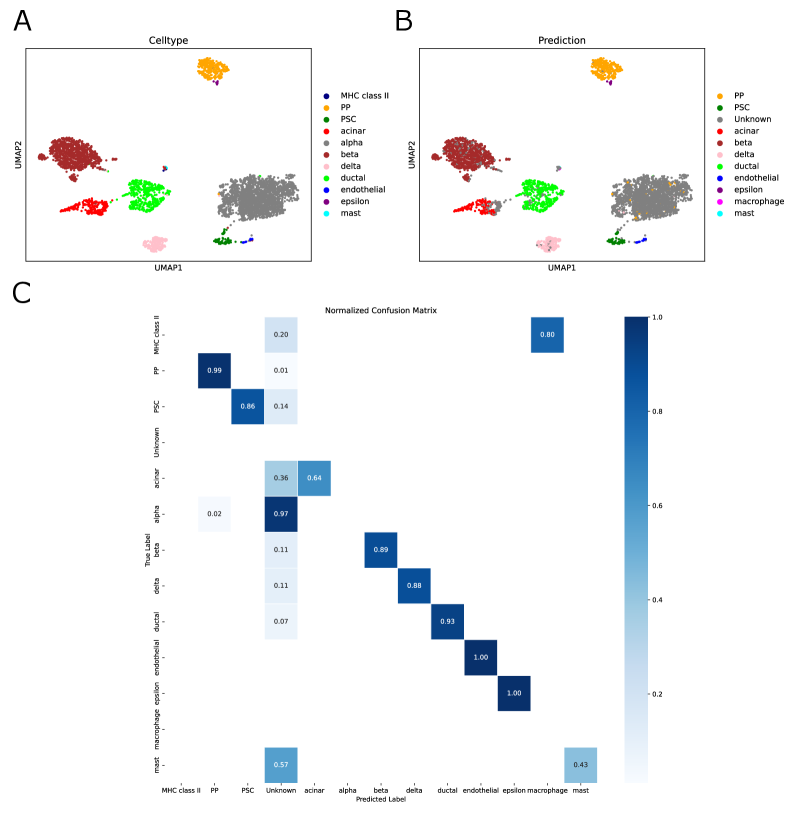
- Biology Confidence-based rejection mechanism that successfully identifies 97% of unseen 'alpha cells' as 'Unknown' in pancreas datasets, enabling robust detection of novel cell types absent from training data.
主要结论
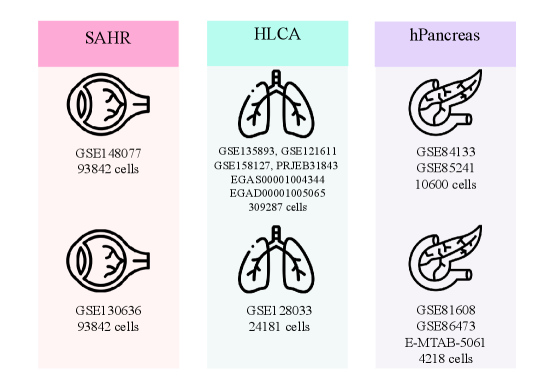
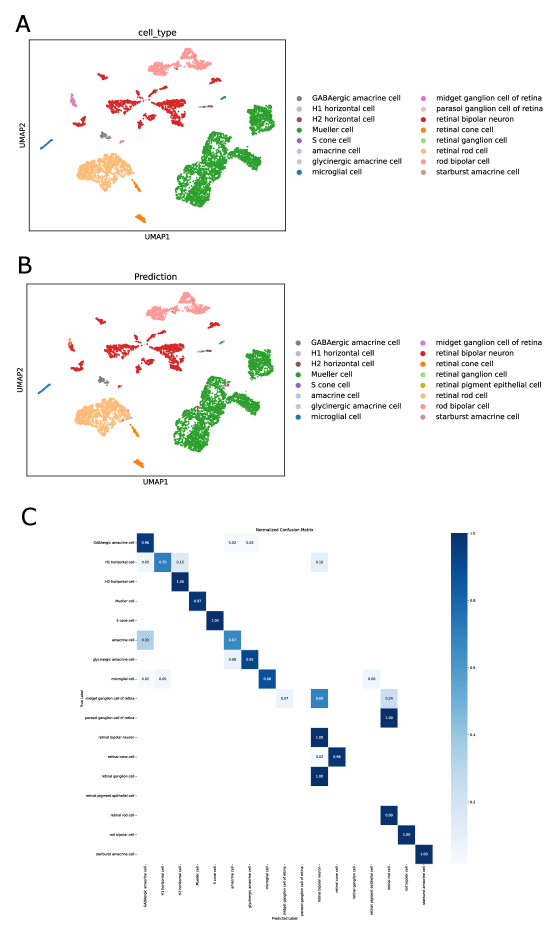
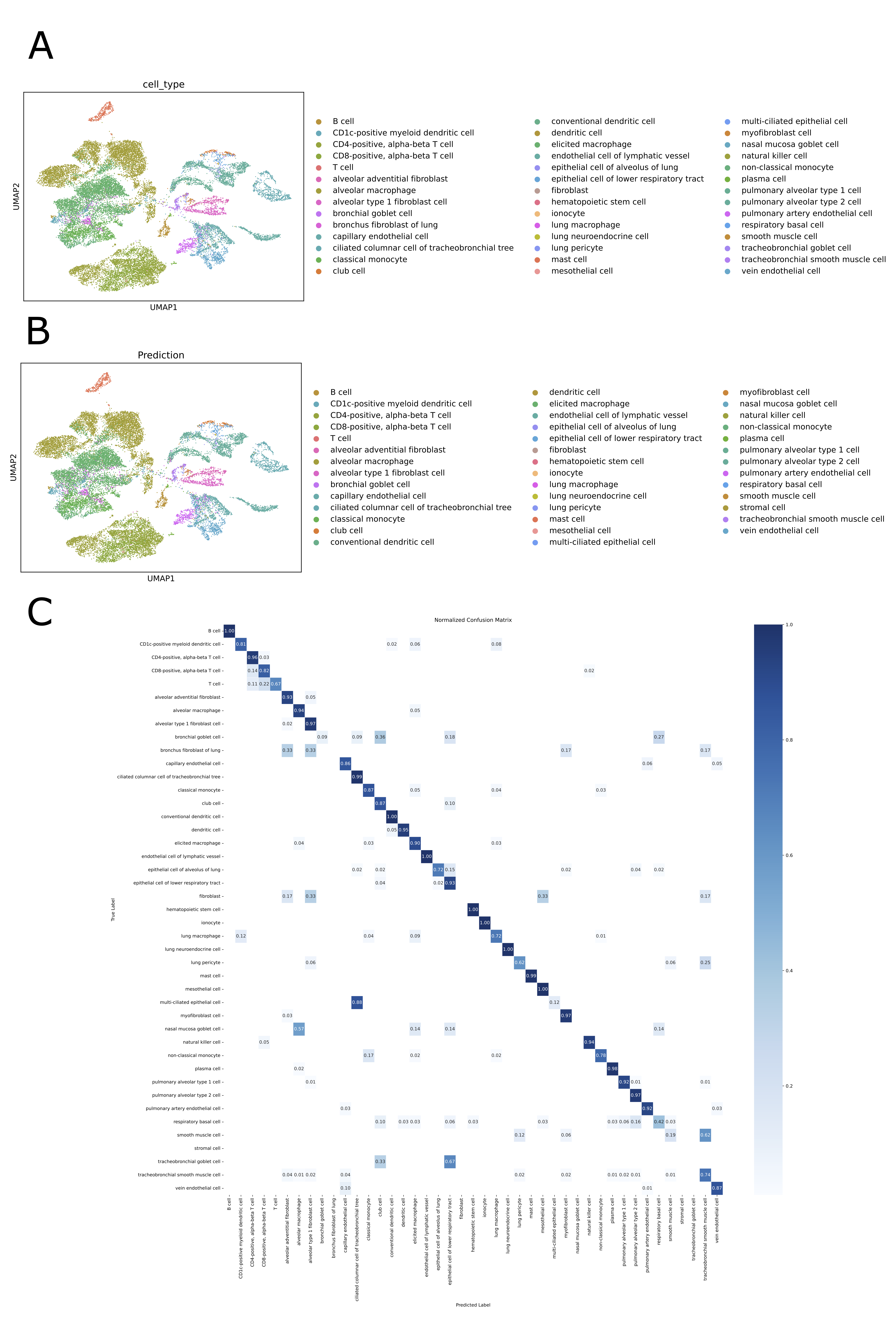
- SpikGPT achieves accuracy of 0.991 on SAHR dataset and 0.920 on HLCA dataset, outperforming or matching 8 benchmark methods including scGPT, CCA, and scPred.
- The model demonstrates superior robustness to batch effects, maintaining macro F1-score of 0.711 on heterogeneous HLCA data where traditional methods like SingleR drop to 0.207 F1-score.
- SpikGPT successfully identifies 97% of unseen 'alpha cells' as 'Unknown' using confidence thresholding (p<0.05), enabling reliable detection of novel cell populations.
摘要: Accurate and scalable cell type annotation remains a challenge in single-cell transcriptomics, especially when datasets exhibit strong batch effects or contain previously unseen cell populations. Here we introduce SpikGPT, a hybrid deep learning framework that integrates scGPT-derived cell embeddings with a spiking Transformer architecture to achieve efficient and robust annotation. scGPT provides biologically informed dense representations of each cell, which are further processed by a multi-head Spiking Self-Attention mechanism, energy-efficient feature extraction. Across multiple benchmark datasets, SpikGPT consistently matches or exceeds the performance of leading annotation tools. Notably, SpikGPT uniquely identifies unseen cell types by assigning low-confidence predictions to an 'Unknown' category, allowing accurate rejection of cell states absent from the training reference. Together, these results demonstrate that SpikGPT is a versatile and reliable annotation tool capable of generalizing across datasets, resolving complex cellular heterogeneity, and facilitating discovery of novel or disease-associated cell populations.