Paper List
-
SpikGPT: A High-Accuracy and Interpretable Spiking Attention Framework for Single-Cell Annotation
This paper addresses the core challenge of robust single-cell annotation across heterogeneous datasets with batch effects and the critical need to ide...
-
Unlocking hidden biomolecular conformational landscapes in diffusion models at inference time
This paper addresses the core challenge of efficiently and accurately sampling the conformational landscape of biomolecules from diffusion-based struc...
-
Personalized optimization of pediatric HD-tDCS for dose consistency and target engagement
This paper addresses the critical limitation of one-size-fits-all HD-tDCS protocols in pediatric populations by developing a personalized optimization...
-
Realistic Transition Paths for Large Biomolecular Systems: A Langevin Bridge Approach
This paper addresses the core challenge of generating physically realistic and computationally efficient transition paths between distinct protein con...
-
Consistent Synthetic Sequences Unlock Structural Diversity in Fully Atomistic De Novo Protein Design
This paper addresses the core pain point of low sequence-structure alignment in existing synthetic datasets (e.g., AFDB), which severely limits the pe...
-
MoRSAIK: Sequence Motif Reactor Simulation, Analysis and Inference Kit in Python
This work addresses the computational bottleneck in simulating prebiotic RNA reactor dynamics by developing a Python package that tracks sequence moti...
-
On the Approximation of Phylogenetic Distance Functions by Artificial Neural Networks
This paper addresses the core challenge of developing computationally efficient and scalable neural network architectures that can learn accurate phyl...
-
EcoCast: A Spatio-Temporal Model for Continual Biodiversity and Climate Risk Forecasting
This paper addresses the critical bottleneck in conservation: the lack of timely, high-resolution, near-term forecasts of species distribution shifts ...
SpikGPT: A High-Accuracy and Interpretable Spiking Attention Framework for Single-Cell Annotation
Department of Biomedical Informatics, Emory University | Department of Surgery, Duke University
30秒速读
IN SHORT: This paper addresses the core challenge of robust single-cell annotation across heterogeneous datasets with batch effects and the critical need to identify previously unseen cell populations.
核心创新
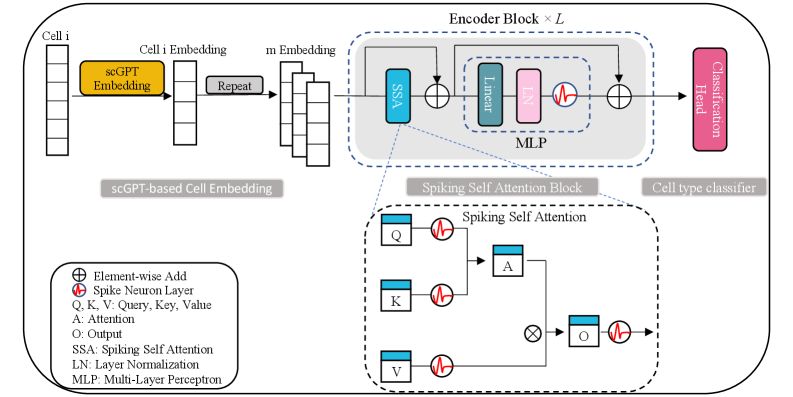
- Methodology First integration of spiking neural networks with transformer architecture for single-cell analysis, using Leaky Integrate-and-Fire (LIF) neurons in a multi-head Spiking Self-Attention mechanism for energy-efficient computation.
- Methodology Novel two-step embedding expansion strategy: repeating cell embeddings along feature channels (default m=300) and temporal dimensions (default T=4) to enhance representation richness and training stability.
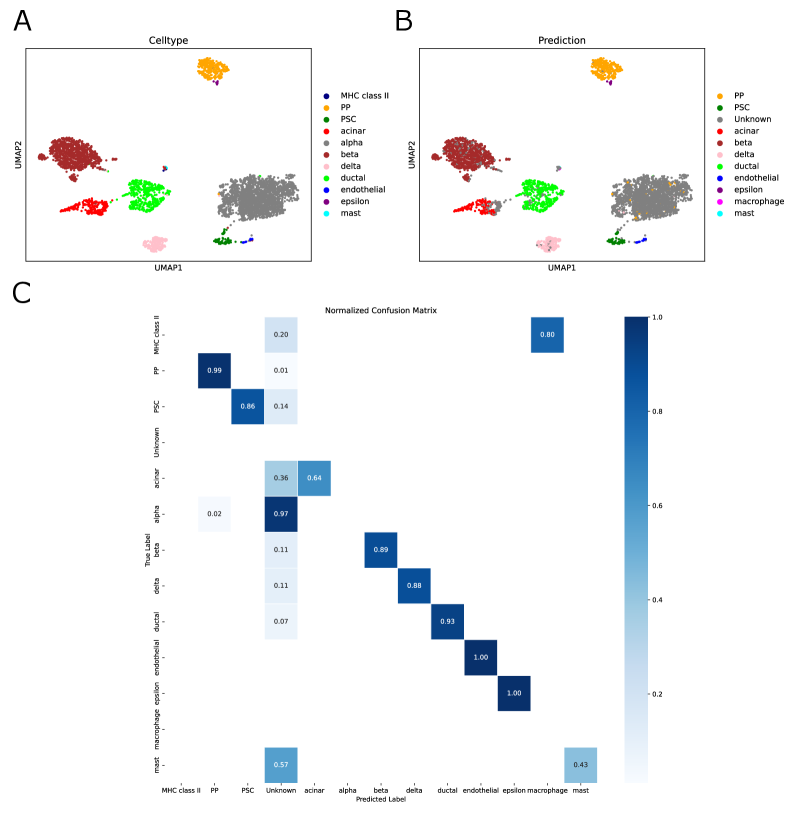
- Biology Confidence-based rejection mechanism that successfully identifies 97% of unseen 'alpha cells' as 'Unknown' in pancreas datasets, enabling robust detection of novel cell types absent from training data.
主要结论
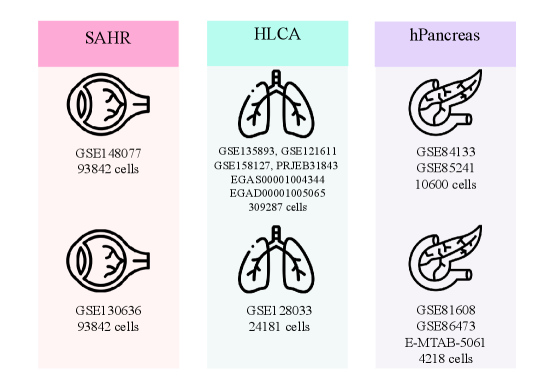
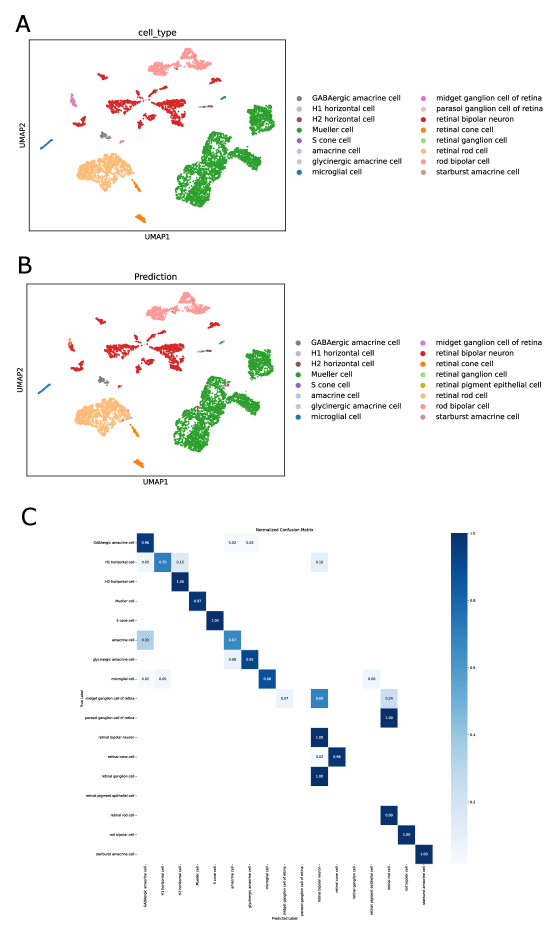
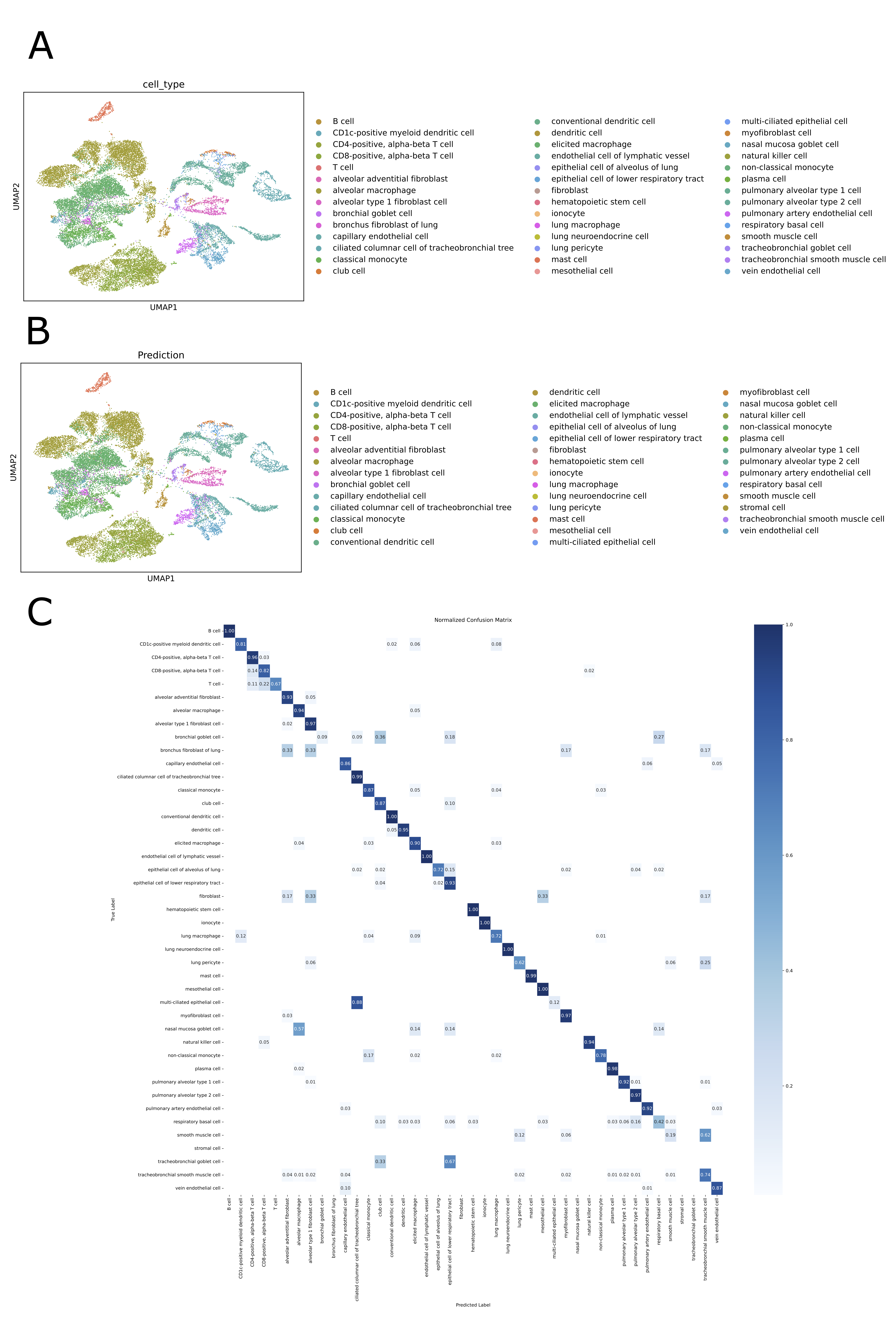
- SpikGPT achieves accuracy of 0.991 on SAHR dataset and 0.920 on HLCA dataset, outperforming or matching 8 benchmark methods including scGPT, CCA, and scPred.
- The model demonstrates superior robustness to batch effects, maintaining macro F1-score of 0.711 on heterogeneous HLCA data where traditional methods like SingleR drop to 0.207 F1-score.
- SpikGPT successfully identifies 97% of unseen 'alpha cells' as 'Unknown' using confidence thresholding (p<0.05), enabling reliable detection of novel cell populations.
摘要: Accurate and scalable cell type annotation remains a challenge in single-cell transcriptomics, especially when datasets exhibit strong batch effects or contain previously unseen cell populations. Here we introduce SpikGPT, a hybrid deep learning framework that integrates scGPT-derived cell embeddings with a spiking Transformer architecture to achieve efficient and robust annotation. scGPT provides biologically informed dense representations of each cell, which are further processed by a multi-head Spiking Self-Attention mechanism, energy-efficient feature extraction. Across multiple benchmark datasets, SpikGPT consistently matches or exceeds the performance of leading annotation tools. Notably, SpikGPT uniquely identifies unseen cell types by assigning low-confidence predictions to an 'Unknown' category, allowing accurate rejection of cell states absent from the training reference. Together, these results demonstrate that SpikGPT is a versatile and reliable annotation tool capable of generalizing across datasets, resolving complex cellular heterogeneity, and facilitating discovery of novel or disease-associated cell populations.