Paper List
-
Macroscopic Dominance from Microscopic Extremes: Symmetry Breaking in Spatial Competition
This paper addresses the fundamental question of how microscopic stochastic advantages in spatial exploration translate into macroscopic resource domi...
-
Linear Readout of Neural Manifolds with Continuous Variables
This paper addresses the core challenge of quantifying how the geometric structure of high-dimensional neural population activity (neural manifolds) d...
-
Theory of Cell Body Lensing and Phototaxis Sign Reversal in “Eyeless” Mutants of Chlamydomonas
This paper solves the core puzzle of how eyeless mutants of Chlamydomonas exhibit reversed phototaxis by quantitatively modeling the competition betwe...
-
Cross-Species Transfer Learning for Electrophysiology-to-Transcriptomics Mapping in Cortical GABAergic Interneurons
This paper addresses the challenge of predicting transcriptomic identity from electrophysiological recordings in human cortical interneurons, where li...
-
Uncovering statistical structure in large-scale neural activity with Restricted Boltzmann Machines
This paper addresses the core challenge of modeling large-scale neural population activity (1500-2000 neurons) with interpretable higher-order interac...
-
Realizing Common Random Numbers: Event-Keyed Hashing for Causally Valid Stochastic Models
This paper addresses the critical problem that standard stateful PRNG implementations in agent-based models violate causal validity by making random d...
-
A Standardized Framework for Evaluating Gene Expression Generative Models
This paper addresses the critical lack of standardized evaluation protocols for single-cell gene expression generative models, where inconsistent metr...
-
Single Molecule Localization Microscopy Challenge: A Biologically Inspired Benchmark for Long-Sequence Modeling
This paper addresses the core challenge of evaluating state-space models on biologically realistic, sparse, and stochastic temporal processes, which a...
Beyond Bayesian Inference: The Correlation Integral Likelihood Framework and Gradient Flow Methods for Deterministic Sampling
Institute of Mathematics, Polish Academy of Sciences | Interdisciplinary Centre for Mathematical and Computational Modelling, University of Warsaw | Institute for Mathematics, Heidelberg University
30秒速读
IN SHORT: This paper addresses the core challenge of calibrating complex biological models (e.g., PDEs, agent-based models) with incomplete, noisy, or heterogeneous data, where traditional pointwise comparison methods fail due to system sensitivity and intrinsic variability.
核心创新
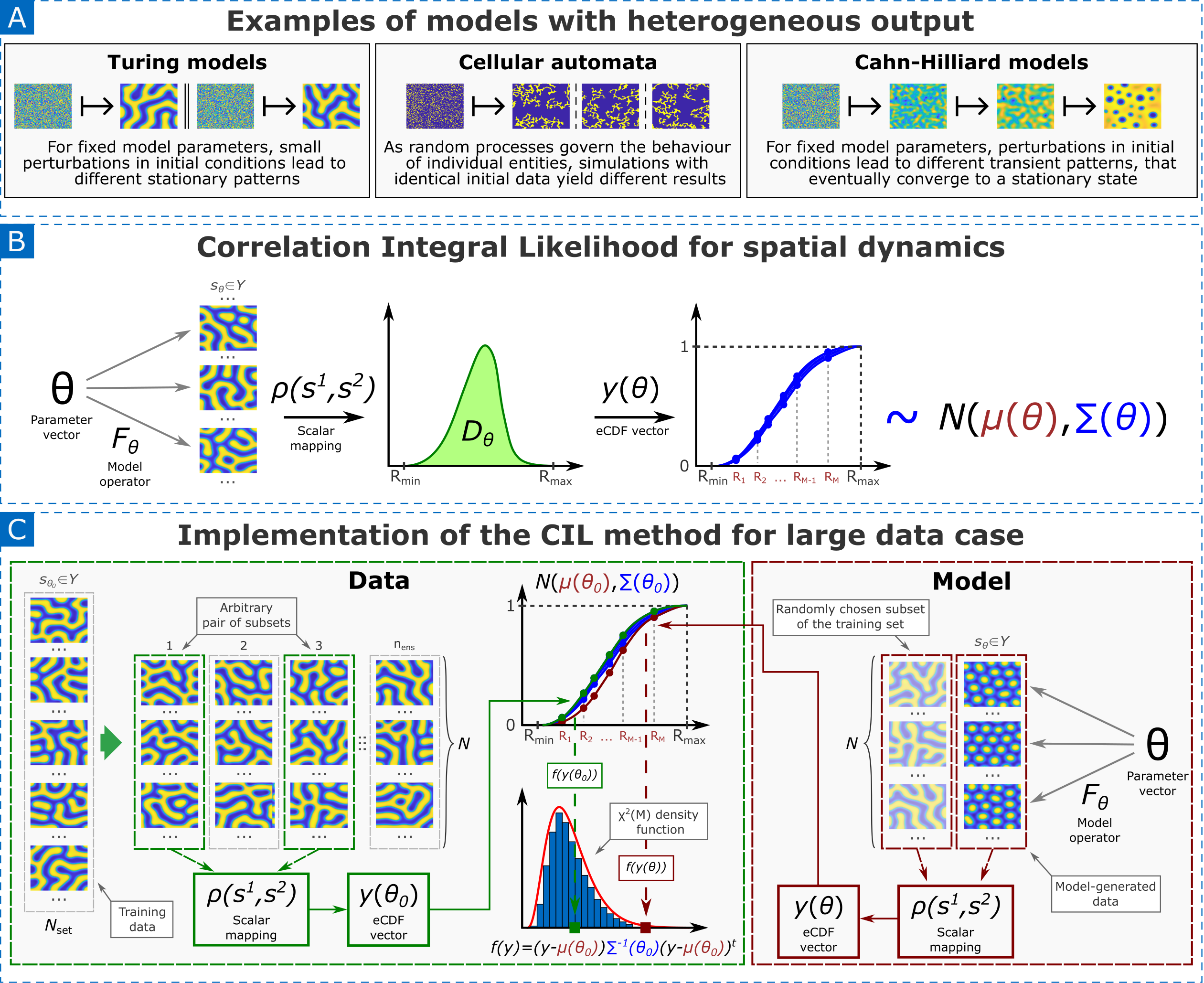
- Methodology Introduces the Correlation Integral Likelihood (CIL) framework, a unified approach for parameter estimation in systems with heterogeneous or chaotic dynamics (e.g., pattern formation, individual-based models), moving beyond classical Bayesian methods.
- Methodology Proposes integration of deterministic gradient flow methods within the CIL framework to enhance inference efficiency and accuracy, compared to traditional stochastic sampling (e.g., MCMC).
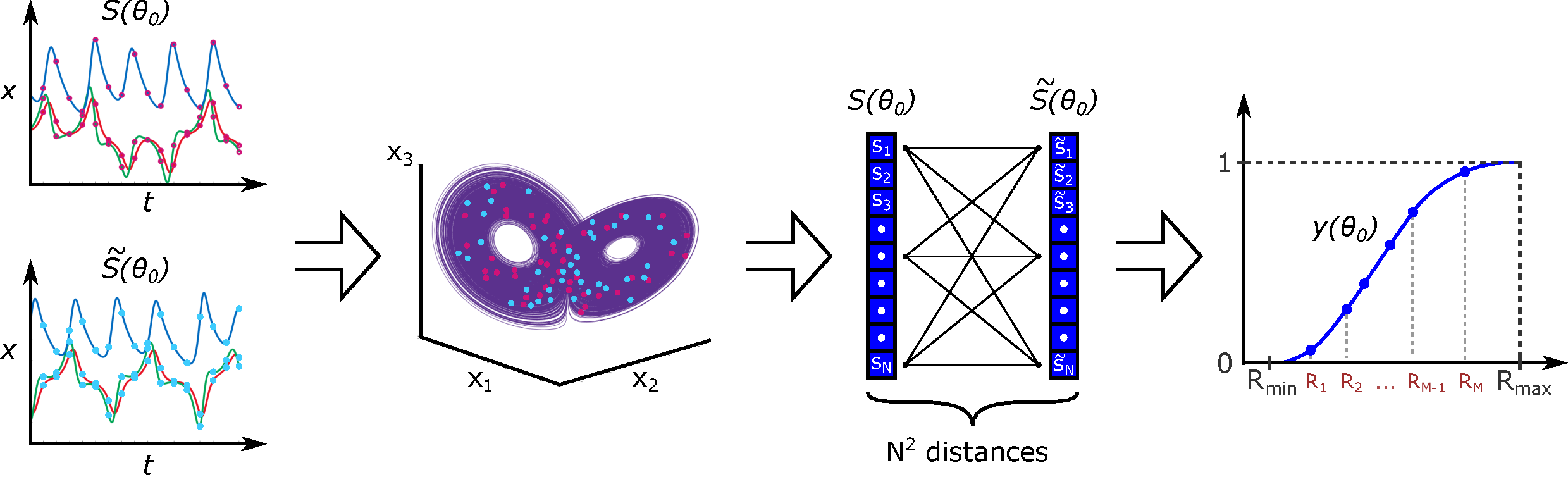
- Theory Generalizes the concept of correlation dimension from chaos theory to construct a robust metric for comparing the global geometric structure of model outputs (e.g., attractors, spatial patterns) rather than relying on unstable pointwise comparisons.
主要结论
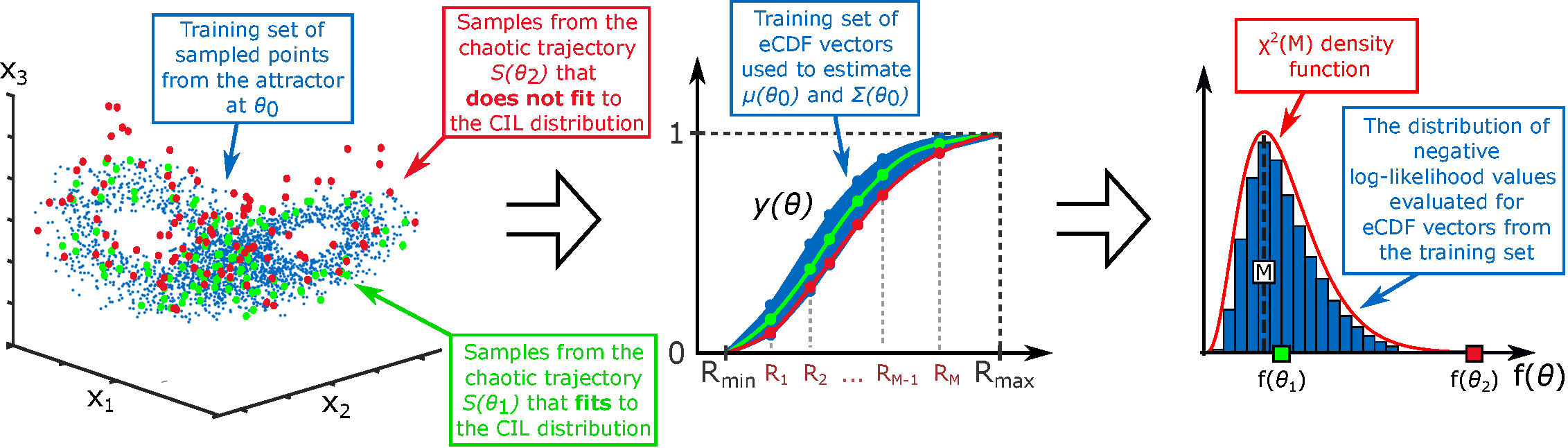
- The CIL method provides a theoretically grounded framework for parameter estimation in systems where solution heterogeneity (e.g., in Turing patterns or chaotic attractors) makes conventional likelihoods ineffective.
- Integrating deterministic gradient flow sampling with the CIL framework can potentially enhance computational efficiency and inference accuracy compared to purely stochastic methods like MCMC, especially for high-dimensional parameter spaces.
- The approach enables reliable model calibration and validation even with incomplete, noisy, or single-snapshot data, advancing the predictive capability and mechanistic understanding of complex biological systems.
摘要: Calibrating mathematical models of biological processes is essential for achieving predictive accuracy and gaining mechanistic insight. However, this task remains challenging due to limited and noisy data, significant biological variability, and the computational complexity of the models themselves. In this method's article, we explore a range of approaches for parameter inference in partial differential equation (PDE) models of biological systems. We introduce a unified mathematical framework, the Correlation Integral Likelihood (CIL) method, for parameter estimation in systems exhibiting heterogeneous or chaotic dynamics, encompassing both pattern formation models and individual-based models. Departing from classical Bayesian inverse problem methodologies, we motivate the development of the CIL method, demonstrate its versatility, and highlight illustrative applications within mathematical biology. Furthermore, we compare stochastic sampling strategies, such as Markov Chain Monte Carlo (MCMC), with deterministic gradient flow approaches, highlighting how these methods can be integrated within the proposed framework to enhance inference performance. Our work provides a practical and theoretically grounded toolbox for researchers seeking to calibrate complex biological models using incomplete, noisy, or heterogeneous data, thereby advancing both the predictive capability and mechanistic understanding of such systems.