Paper List
-
SpikGPT: A High-Accuracy and Interpretable Spiking Attention Framework for Single-Cell Annotation
This paper addresses the core challenge of robust single-cell annotation across heterogeneous datasets with batch effects and the critical need to ide...
-
Unlocking hidden biomolecular conformational landscapes in diffusion models at inference time
This paper addresses the core challenge of efficiently and accurately sampling the conformational landscape of biomolecules from diffusion-based struc...
-
Personalized optimization of pediatric HD-tDCS for dose consistency and target engagement
This paper addresses the critical limitation of one-size-fits-all HD-tDCS protocols in pediatric populations by developing a personalized optimization...
-
Realistic Transition Paths for Large Biomolecular Systems: A Langevin Bridge Approach
This paper addresses the core challenge of generating physically realistic and computationally efficient transition paths between distinct protein con...
-
Consistent Synthetic Sequences Unlock Structural Diversity in Fully Atomistic De Novo Protein Design
This paper addresses the core pain point of low sequence-structure alignment in existing synthetic datasets (e.g., AFDB), which severely limits the pe...
-
MoRSAIK: Sequence Motif Reactor Simulation, Analysis and Inference Kit in Python
This work addresses the computational bottleneck in simulating prebiotic RNA reactor dynamics by developing a Python package that tracks sequence moti...
-
On the Approximation of Phylogenetic Distance Functions by Artificial Neural Networks
This paper addresses the core challenge of developing computationally efficient and scalable neural network architectures that can learn accurate phyl...
-
EcoCast: A Spatio-Temporal Model for Continual Biodiversity and Climate Risk Forecasting
This paper addresses the critical bottleneck in conservation: the lack of timely, high-resolution, near-term forecasts of species distribution shifts ...
Beyond Bayesian Inference: The Correlation Integral Likelihood Framework and Gradient Flow Methods for Deterministic Sampling
Institute of Mathematics, Polish Academy of Sciences | Interdisciplinary Centre for Mathematical and Computational Modelling, University of Warsaw | Institute for Mathematics, Heidelberg University
30秒速读
IN SHORT: This paper addresses the core challenge of calibrating complex biological models (e.g., PDEs, agent-based models) with incomplete, noisy, or heterogeneous data, where traditional pointwise comparison methods fail due to system sensitivity and intrinsic variability.
核心创新
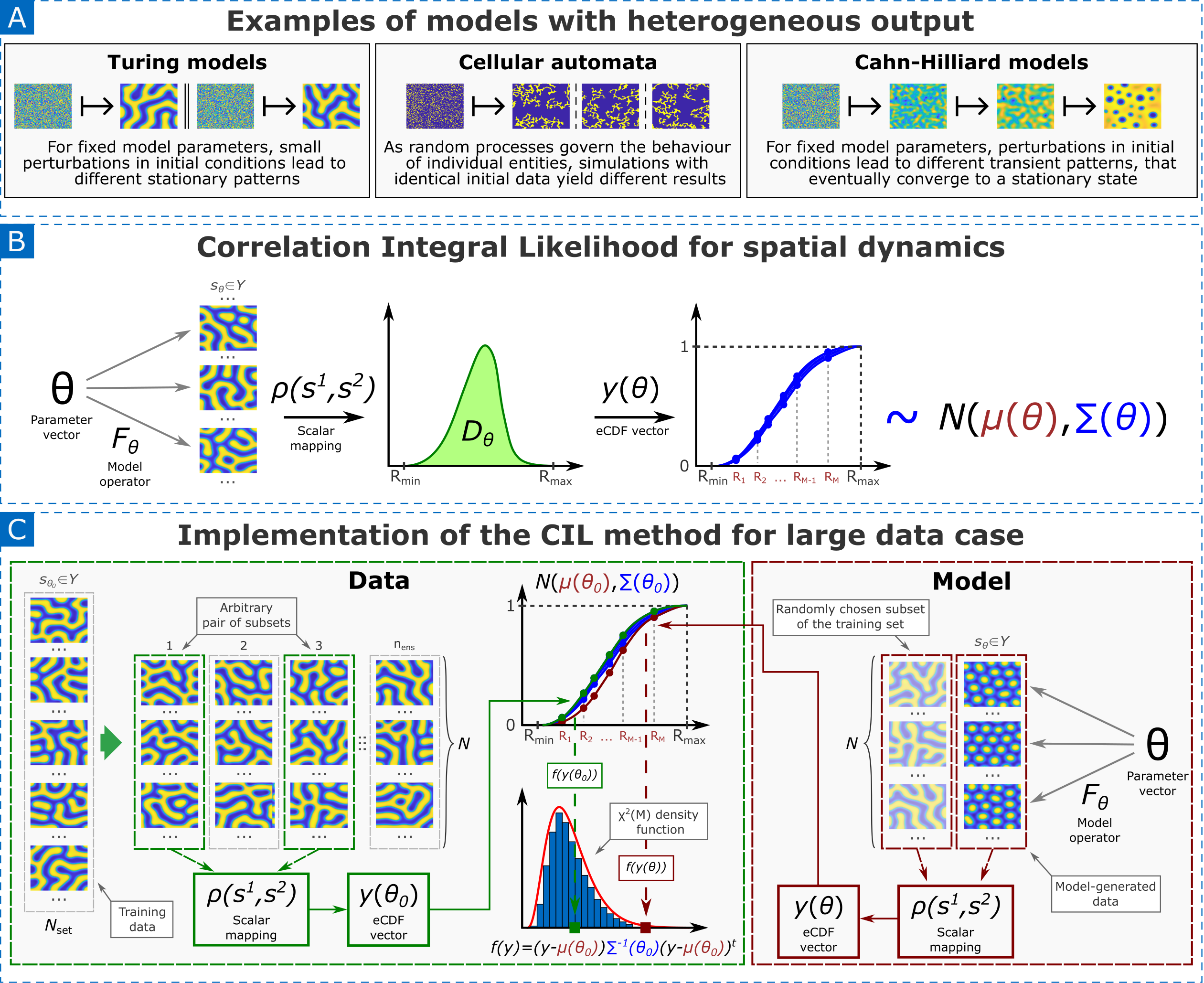
- Methodology Introduces the Correlation Integral Likelihood (CIL) framework, a unified approach for parameter estimation in systems with heterogeneous or chaotic dynamics (e.g., pattern formation, individual-based models), moving beyond classical Bayesian methods.
- Methodology Proposes integration of deterministic gradient flow methods within the CIL framework to enhance inference efficiency and accuracy, compared to traditional stochastic sampling (e.g., MCMC).
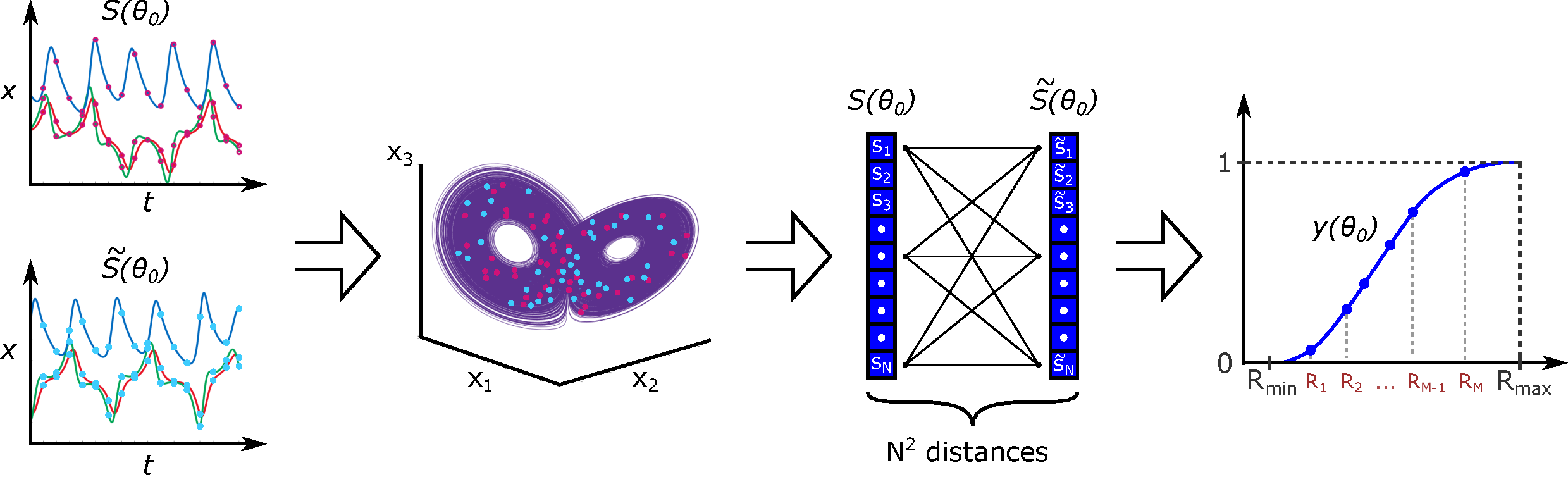
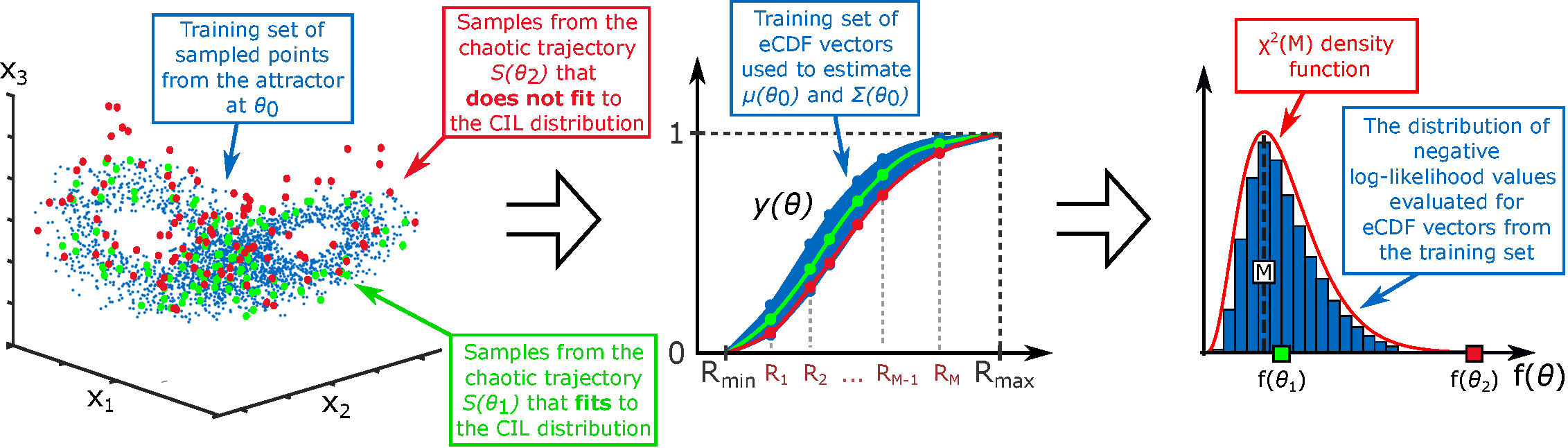
- Theory Generalizes the concept of correlation dimension from chaos theory to construct a robust metric for comparing the global geometric structure of model outputs (e.g., attractors, spatial patterns) rather than relying on unstable pointwise comparisons.
主要结论
- The CIL method provides a theoretically grounded framework for parameter estimation in systems where solution heterogeneity (e.g., in Turing patterns or chaotic attractors) makes conventional likelihoods ineffective.
- Integrating deterministic gradient flow sampling with the CIL framework can potentially enhance computational efficiency and inference accuracy compared to purely stochastic methods like MCMC, especially for high-dimensional parameter spaces.
- The approach enables reliable model calibration and validation even with incomplete, noisy, or single-snapshot data, advancing the predictive capability and mechanistic understanding of complex biological systems.
摘要: Calibrating mathematical models of biological processes is essential for achieving predictive accuracy and gaining mechanistic insight. However, this task remains challenging due to limited and noisy data, significant biological variability, and the computational complexity of the models themselves. In this method's article, we explore a range of approaches for parameter inference in partial differential equation (PDE) models of biological systems. We introduce a unified mathematical framework, the Correlation Integral Likelihood (CIL) method, for parameter estimation in systems exhibiting heterogeneous or chaotic dynamics, encompassing both pattern formation models and individual-based models. Departing from classical Bayesian inverse problem methodologies, we motivate the development of the CIL method, demonstrate its versatility, and highlight illustrative applications within mathematical biology. Furthermore, we compare stochastic sampling strategies, such as Markov Chain Monte Carlo (MCMC), with deterministic gradient flow approaches, highlighting how these methods can be integrated within the proposed framework to enhance inference performance. Our work provides a practical and theoretically grounded toolbox for researchers seeking to calibrate complex biological models using incomplete, noisy, or heterogeneous data, thereby advancing both the predictive capability and mechanistic understanding of such systems.