Paper List
-
Evolutionarily Stable Stackelberg Equilibrium
通过要求追随者策略对突变入侵具有鲁棒性,弥合了斯塔克尔伯格领导力模型与演化稳定性之间的鸿沟。
-
Recovering Sparse Neural Connectivity from Partial Measurements: A Covariance-Based Approach with Granger-Causality Refinement
通过跨多个实验会话累积协方差统计,实现从部分记录到完整神经连接性的重建。
-
Atomic Trajectory Modeling with State Space Models for Biomolecular Dynamics
ATMOS通过提供一个基于SSM的高效框架,用于生物分子的原子级轨迹生成,弥合了计算昂贵的MD模拟与时间受限的深度生成模型之间的差距。
-
Slow evolution towards generalism in a model of variable dietary range
通过证明是种群统计噪声(而非确定性动力学)驱动了模式形成和泛化食性的演化,解决了间接竞争下物种形成的悖论。
-
Grounded Multimodal Retrieval-Augmented Drafting of Radiology Impressions Using Case-Based Similarity Search
通过将印象草稿基于检索到的历史病例,并采用明确引用和基于置信度的拒绝机制,解决放射学报告生成中的幻觉问题。
-
Unified Policy–Value Decomposition for Rapid Adaptation
通过双线性分解在策略和价值函数之间共享低维目标嵌入,实现对新颖任务的零样本适应。
-
Mathematical Modeling of Cancer–Bacterial Therapy: Analysis and Numerical Simulation via Physics-Informed Neural Networks
提供了一个严格的、无网格的PINN框架,用于模拟和分析细菌癌症疗法中复杂的、空间异质的相互作用。
-
Sample-Efficient Adaptation of Drug-Response Models to Patient Tumors under Strong Biological Domain Shift
通过从无标记分子谱中学习可迁移表征,利用最少的临床数据实现患者药物反应的有效预测。
Beyond Bayesian Inference: The Correlation Integral Likelihood Framework and Gradient Flow Methods for Deterministic Sampling
Institute of Mathematics, Polish Academy of Sciences | Interdisciplinary Centre for Mathematical and Computational Modelling, University of Warsaw | Institute for Mathematics, Heidelberg University
30秒速读
IN SHORT: This paper addresses the core challenge of calibrating complex biological models (e.g., PDEs, agent-based models) with incomplete, noisy, or heterogeneous data, where traditional pointwise comparison methods fail due to system sensitivity and intrinsic variability.
核心创新
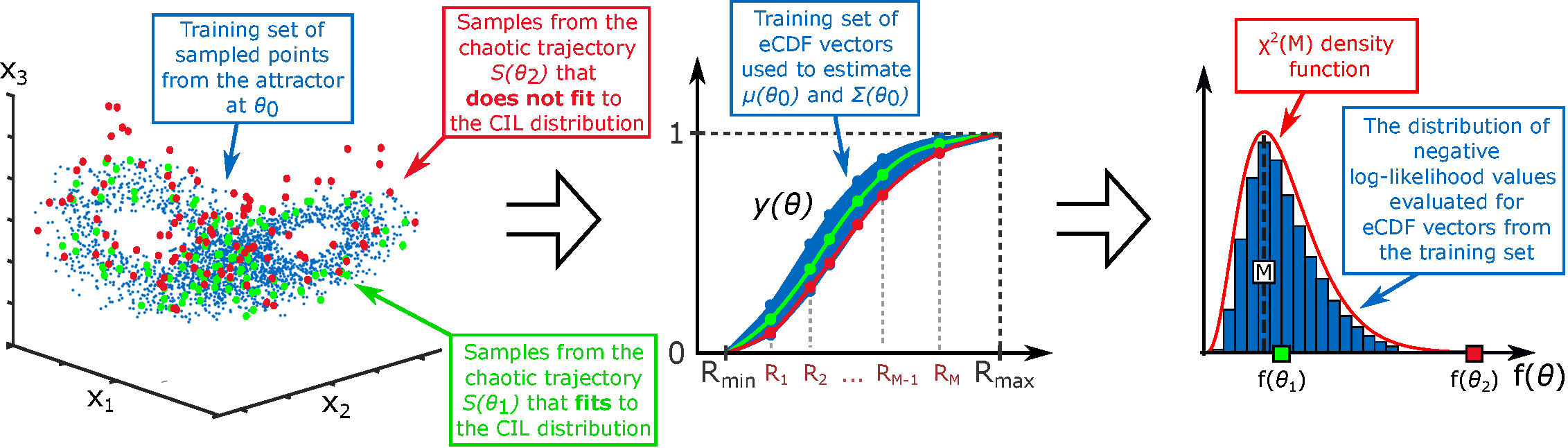
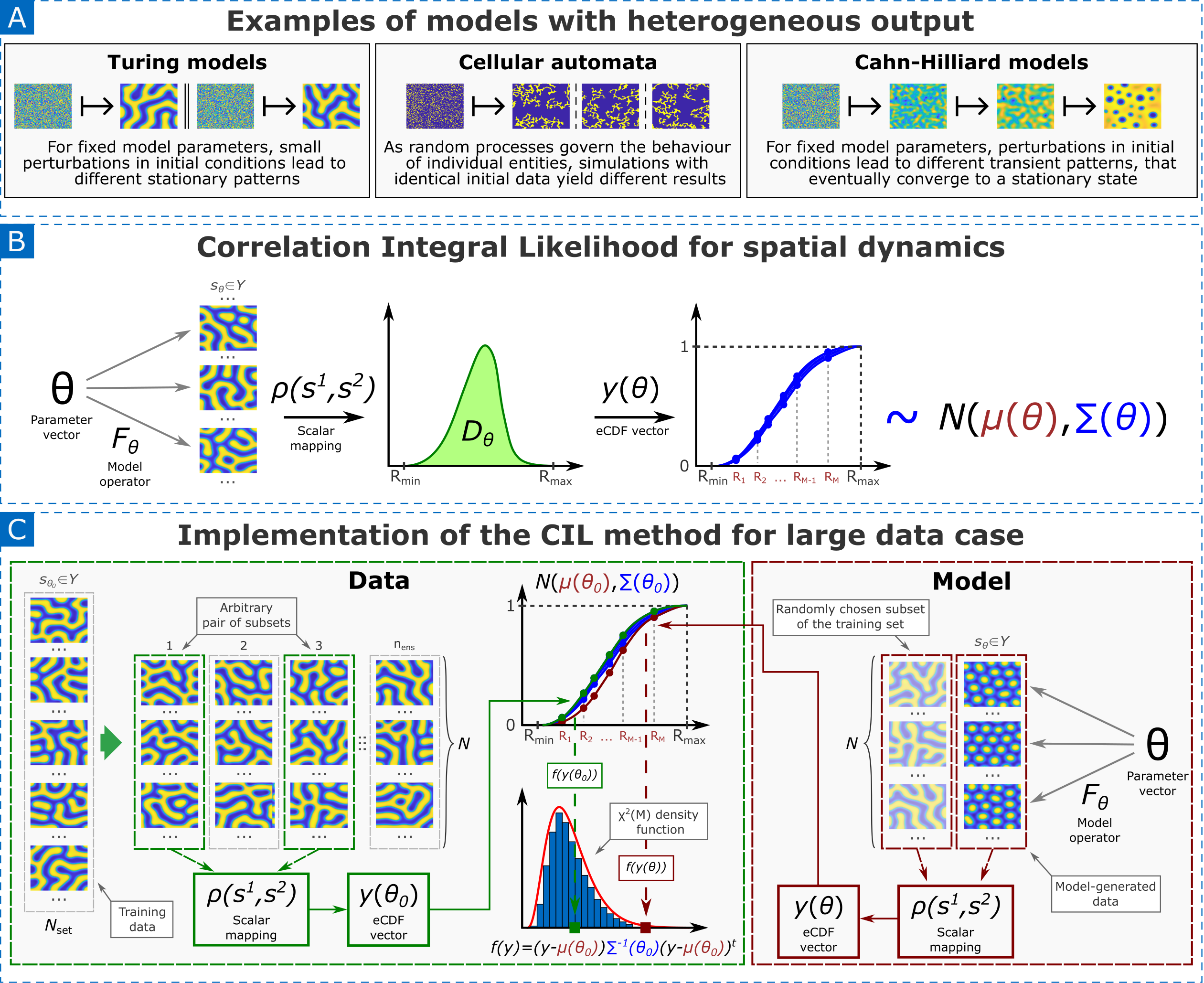
- Methodology Introduces the Correlation Integral Likelihood (CIL) framework, a unified approach for parameter estimation in systems with heterogeneous or chaotic dynamics (e.g., pattern formation, individual-based models), moving beyond classical Bayesian methods.
- Methodology Proposes integration of deterministic gradient flow methods within the CIL framework to enhance inference efficiency and accuracy, compared to traditional stochastic sampling (e.g., MCMC).
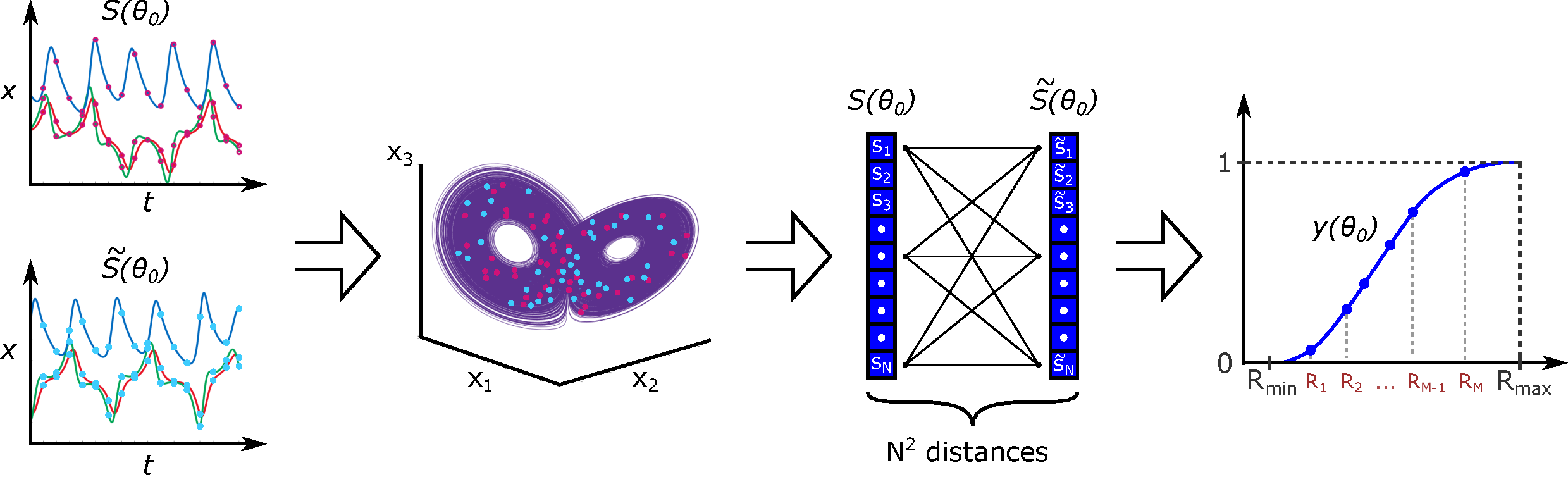
- Theory Generalizes the concept of correlation dimension from chaos theory to construct a robust metric for comparing the global geometric structure of model outputs (e.g., attractors, spatial patterns) rather than relying on unstable pointwise comparisons.
主要结论
- The CIL method provides a theoretically grounded framework for parameter estimation in systems where solution heterogeneity (e.g., in Turing patterns or chaotic attractors) makes conventional likelihoods ineffective.
- Integrating deterministic gradient flow sampling with the CIL framework can potentially enhance computational efficiency and inference accuracy compared to purely stochastic methods like MCMC, especially for high-dimensional parameter spaces.
- The approach enables reliable model calibration and validation even with incomplete, noisy, or single-snapshot data, advancing the predictive capability and mechanistic understanding of complex biological systems.
摘要: Calibrating mathematical models of biological processes is essential for achieving predictive accuracy and gaining mechanistic insight. However, this task remains challenging due to limited and noisy data, significant biological variability, and the computational complexity of the models themselves. In this method's article, we explore a range of approaches for parameter inference in partial differential equation (PDE) models of biological systems. We introduce a unified mathematical framework, the Correlation Integral Likelihood (CIL) method, for parameter estimation in systems exhibiting heterogeneous or chaotic dynamics, encompassing both pattern formation models and individual-based models. Departing from classical Bayesian inverse problem methodologies, we motivate the development of the CIL method, demonstrate its versatility, and highlight illustrative applications within mathematical biology. Furthermore, we compare stochastic sampling strategies, such as Markov Chain Monte Carlo (MCMC), with deterministic gradient flow approaches, highlighting how these methods can be integrated within the proposed framework to enhance inference performance. Our work provides a practical and theoretically grounded toolbox for researchers seeking to calibrate complex biological models using incomplete, noisy, or heterogeneous data, thereby advancing both the predictive capability and mechanistic understanding of such systems.