Paper List
-
GOPHER: Optimization-based Phenotype Randomization for Genome-Wide Association Studies with Differential Privacy
This paper addresses the core challenge of balancing rigorous privacy protection with data utility when releasing full GWAS summary statistics, overco...
-
Real-time Cricket Sorting By Sex A low-cost embedded solution using YOLOv8 and Raspberry Pi
This paper addresses the critical bottleneck in industrial insect farming: the lack of automated, real-time sex sorting systems for Acheta domesticus ...
-
Training Dynamics of Learning 3D-Rotational Equivariance
This work addresses the core dilemma of whether to use computationally expensive equivariant architectures or faster symmetry-agnostic models with dat...
-
Fast and Accurate Node-Age Estimation Under Fossil Calibration Uncertainty Using the Adjusted Pairwise Likelihood
This paper addresses the dual challenge of computational inefficiency and sensitivity to fossil calibration errors in Bayesian divergence time estimat...
-
Few-shot Protein Fitness Prediction via In-context Learning and Test-time Training
This paper addresses the core challenge of accurately predicting protein fitness with only a handful of experimental observations, where data collecti...
-
scCluBench: Comprehensive Benchmarking of Clustering Algorithms for Single-Cell RNA Sequencing
This paper addresses the critical gap of fragmented and non-standardized benchmarking in single-cell RNA-seq clustering, which hinders objective compa...
-
Simulation and inference methods for non-Markovian stochastic biochemical reaction networks
This paper addresses the computational bottleneck of simulating and performing Bayesian inference for non-Markovian biochemical systems with history-d...
-
Assessment of Simulation-based Inference Methods for Stochastic Compartmental Models
This paper addresses the core challenge of performing accurate Bayesian parameter inference for stochastic epidemic models when the likelihood functio...
Imperfect molecular detection renormalizes apparent kinetic rates in stochastic gene regulatory networks
Department of Mathematical Analysis and Numerical Mathematics, Comenius University, Slovakia | University of Edinburgh, UK
30秒速读
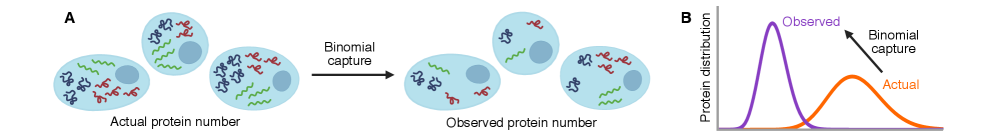
IN SHORT: This paper addresses the core challenge of distinguishing genuine stochastic dynamics of gene regulatory networks from artifacts introduced by imperfect molecular detection in single-cell experiments.
核心创新
- Methodology Extends the binomial capture model from simple gene expression to general gene regulatory networks (GRNs) with explicit regulation, enabling analysis of technical noise in complex systems.
- Theory Establishes precise mathematical conditions under which technical noise leads to a renormalization (rescaling) of kinetic rates versus when it introduces non-absorbable distortions.
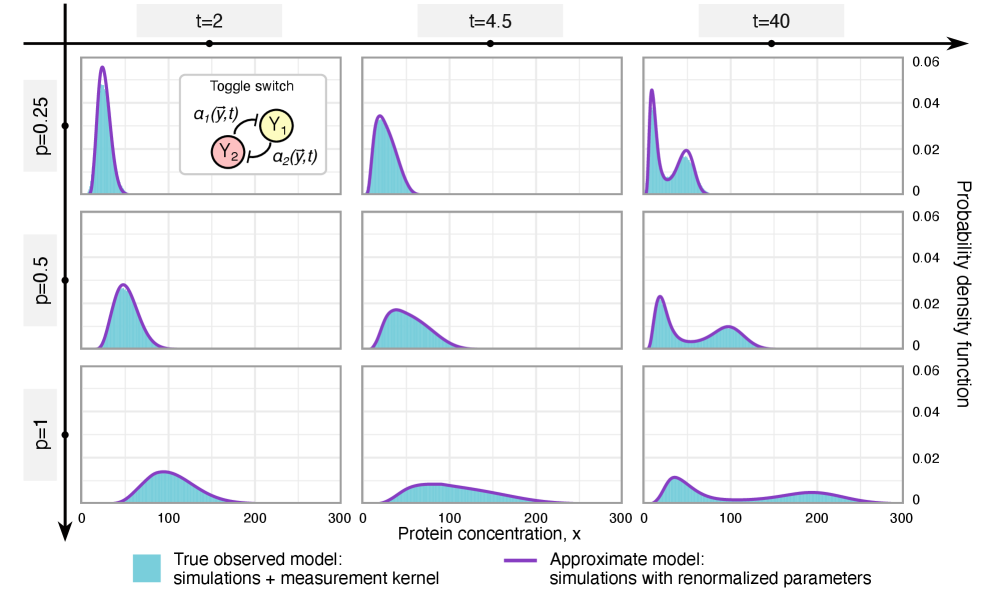
- Methodology Derives results valid for networks of arbitrary connectivity and under time-dependent kinetic rates, significantly generalizing previous steady-state analyses.
主要结论
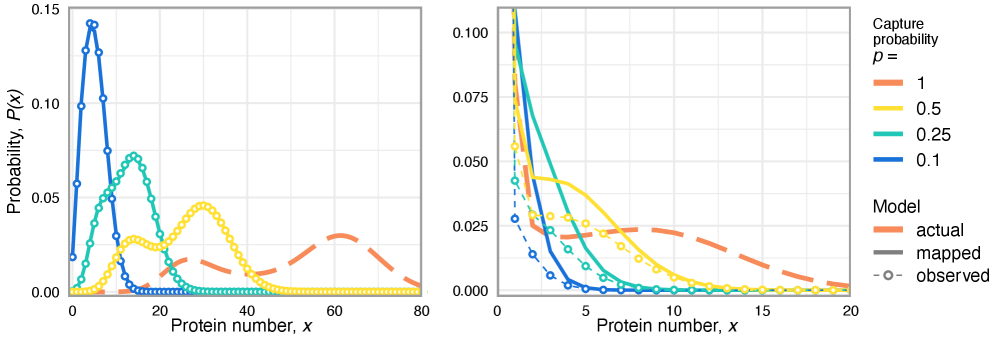
- Technical noise systematically reduces the apparent mean burst size of gene products by a factor of p (the capture probability), e.g., from b(t) to b(t)*p.
- Rate renormalization occurs when promoter-state transitions are on a distinct timescale (much slower/faster) than other reactions or under high transcription factor abundance.
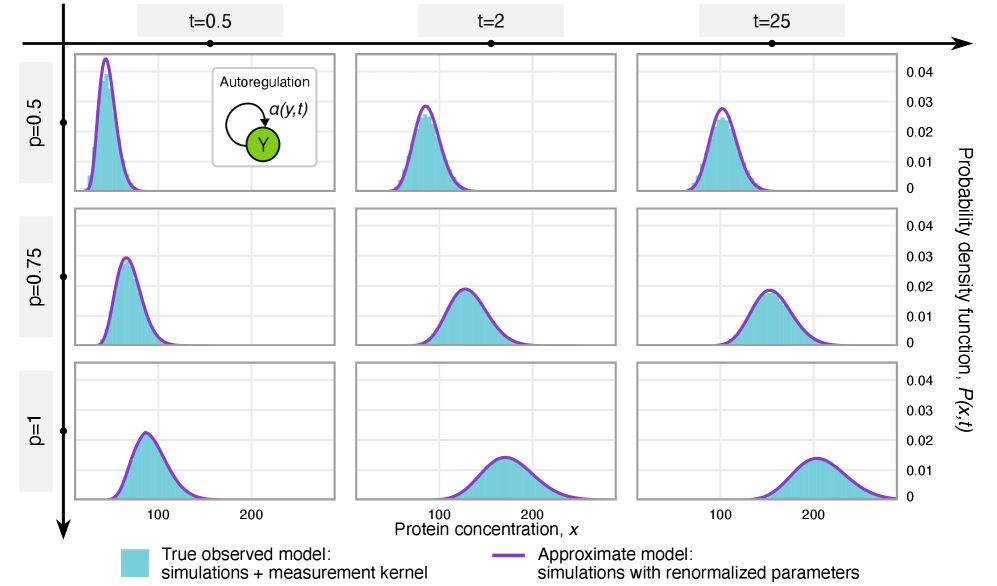
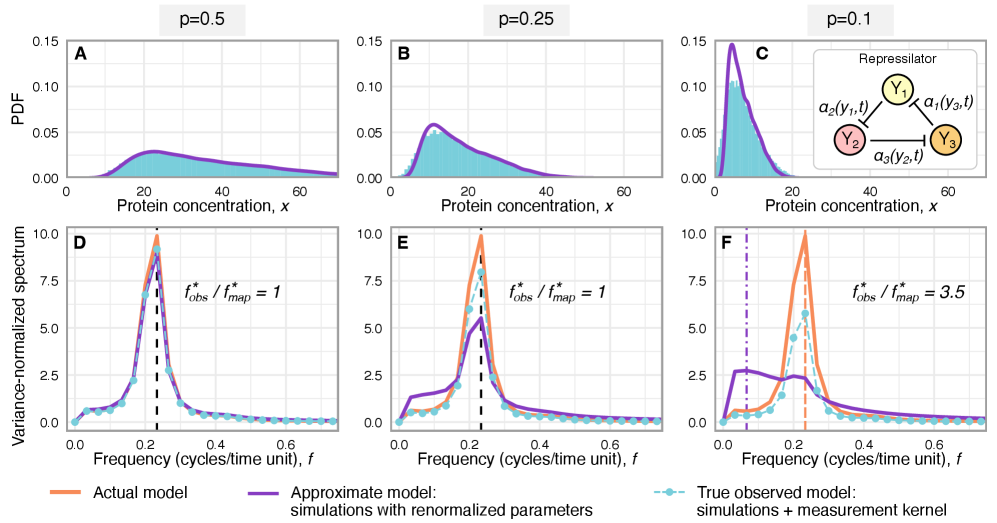
- The framework shows that for the telegraph model, the observed mRNA dynamics are equivalent to the true system with a renormalized transcription rate: k₃(t) → p*k₃(t).
摘要: Imperfect molecular detection in single-cell experiments introduces technical noise that obscures the true stochastic dynamics of gene regulatory networks. While binomial models of molecular capture provide a principled description of imperfect detection, they have so far been analyzed only for simple gene-expression models that do not explicitly account for regulation. Here, we extend binomial models of capture to general gene regulatory networks to understand how imperfect capture reshapes the observed time-dependent statistics of molecular counts. Our results reveal when capture effects correspond to a renormalization of a subset of the kinetic rates and when they cannot be absorbed into effective rates, providing a systematic basis for interpreting noisy single-cell measurements. In particular, we show that rate renormalization emerges either under significant transcription factor abundance or when promoter-state transitions occur on a distinct (much slower or faster) timescale than other reactions. In these cases, technical noise causes the apparent mean burst size of synthesized gene products to appear reduced while transcription factor binding reactions appear faster. These effects hold for gene regulatory networks of arbitrary connectivity and remain valid under time-dependent kinetic rates.