Paper List
-
Formation of Artificial Neural Assemblies by Biologically Plausible Inhibition Mechanisms
This work addresses the core limitation of the Assembly Calculus model—its fixed-size, biologically implausible k-WTA selection process—by introducing...
-
How to make the most of your masked language model for protein engineering
This paper addresses the critical bottleneck of efficiently sampling high-quality, diverse protein sequences from Masked Language Models (MLMs) for pr...
-
Module control in youth symptom networks across COVID-19
This paper addresses the core challenge of distinguishing whether a prolonged societal stressor (COVID-19) fundamentally reorganizes the architecture ...
-
JEDI: Jointly Embedded Inference of Neural Dynamics
This paper addresses the core challenge of inferring context-dependent neural dynamics from noisy, high-dimensional recordings using a single unified ...
-
ATP Level and Phosphorylation Free Energy Regulate Trigger-Wave Speed and Critical Nucleus Size in Cellular Biochemical Systems
This work addresses the core challenge of quantitatively predicting how the cellular energy state (ATP level and phosphorylation free energy) governs ...
-
Packaging Jupyter notebooks as installable desktop apps using LabConstrictor
This paper addresses the core pain point of ensuring Jupyter notebook reproducibility and accessibility across different computing environments, parti...
-
SNPgen: Phenotype-Supervised Genotype Representation and Synthetic Data Generation via Latent Diffusion
This paper addresses the core challenge of generating privacy-preserving synthetic genotype data that maintains both statistical fidelity and downstre...
-
Continuous Diffusion Transformers for Designing Synthetic Regulatory Elements
This paper addresses the challenge of efficiently generating novel, cell-type-specific regulatory DNA sequences with high predicted activity while min...
Assessment of Simulation-based Inference Methods for Stochastic Compartmental Models
Bonn Center for Mathematical Life Sciences, University of Bonn | Life and Medical Science Institute, University of Bonn | Institute of Software Technology, German Aerospace Center (DLR)
30秒速读
IN SHORT: This paper addresses the core challenge of performing accurate Bayesian parameter inference for stochastic epidemic models when the likelihood function is intractable, a common bottleneck for real-time forecasting.
核心创新
- Methodology Provides the first comprehensive, praxis-driven comparison between Particle Filters (PF) and Conditional Normalizing Flows (CNF) for inference on stochastic compartmental models, benchmarking their performance head-to-head.
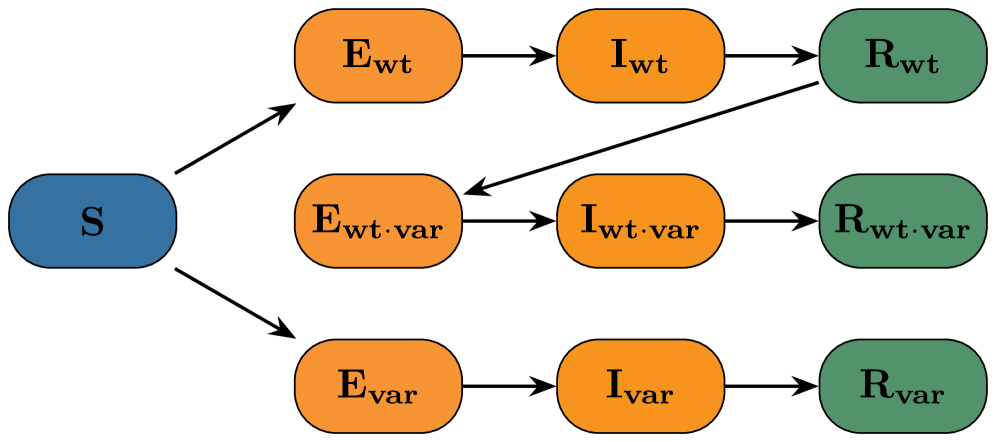
- Methodology Demonstrates the application and robustness of these likelihood-free methods on a complex, non-identifiable two-variant SEIR model with real-world data from an Ethiopian COVID-19 cohort, including scenarios with irregular sampling and missing data.
- Theory Shows that parameter space reparameterization (e.g., using R0, e0, s0) can mitigate ill-conditioning in complex models, improving posterior alignment between PF and CNF methods.
主要结论
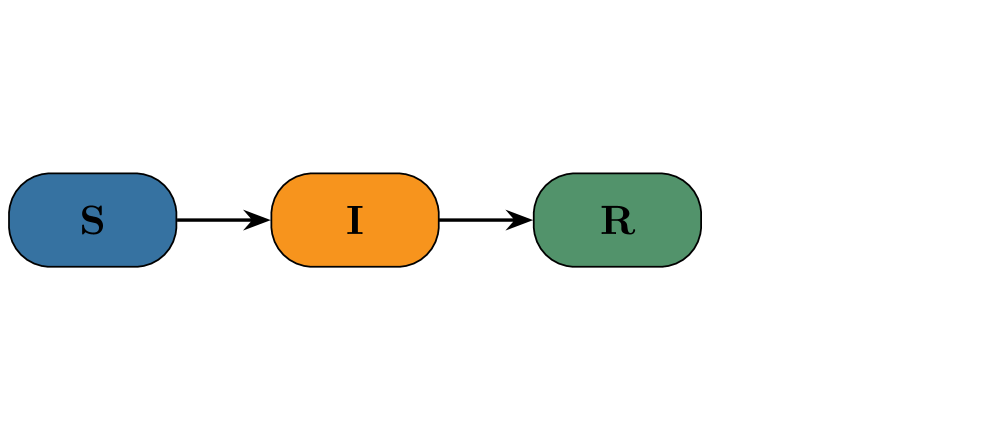
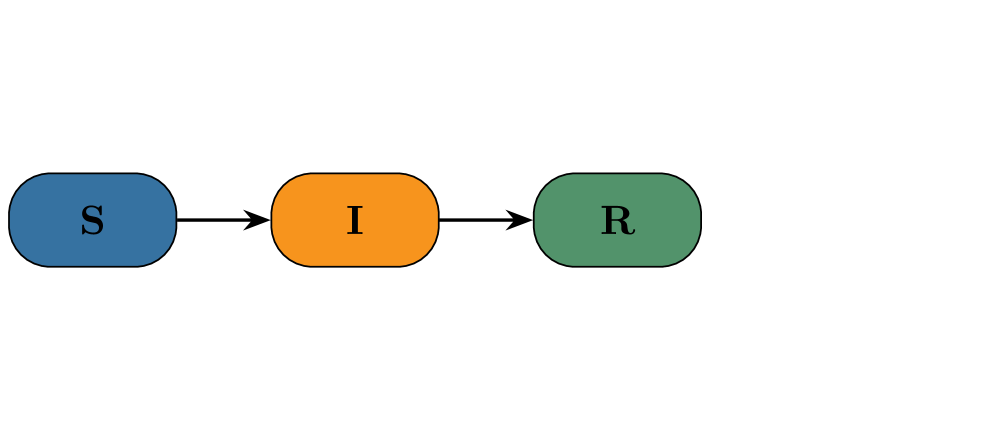
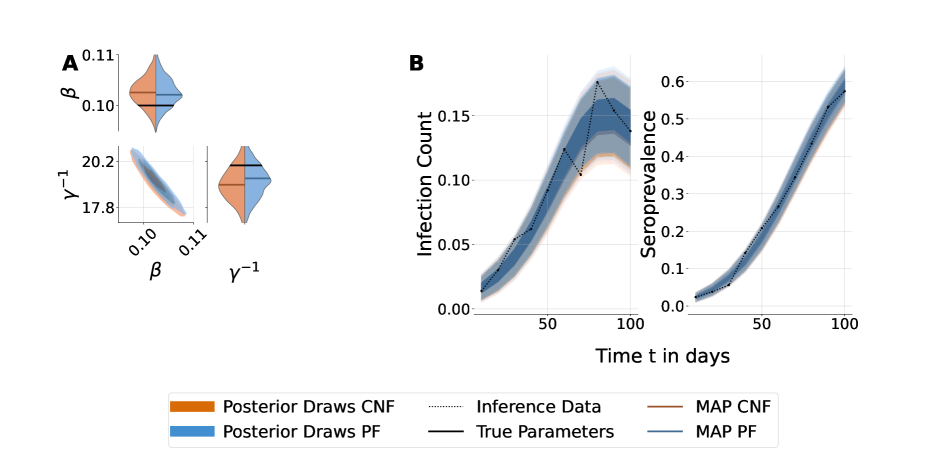
- Both PF and CNF provided robust and reliable inference on the stochastic SIR model with synthetic data, validating the implementation framework.
- For the complex two-variant SEIR model, both methods yielded good fits to synthetic data, but ill-conditioning led to differences in marginal posterior shapes; reparameterization with dimension reduction improved posterior alignment.
- Application to real Ethiopian cohort data demonstrated the operational robustness of both PF and CNF under conditions of real-world noise and irregular data sampling, proving their practical utility.
摘要: Global pandemics, such as the recent COVID-19 crisis, highlight the need for stochastic epidemic models that can capture the randomness inherent in the spread of disease. Such models must be accompanied by methods for estimating parameters in order to generate fast nowcasts and short-term forecasts that can inform public health decisions. This paper presents a comparison of two advanced Bayesian inference methods: 1) pseudo-marginal particle Markov chain Monte Carlo, short Particle Filters (PF), and 2) Conditional Normalizing Flows (CNF). We investigate their performance on two commonly used compartmental models: a classical Susceptible-Infected-Recovered (SIR) model and a two-variant Susceptible-Exposed-Infected-Recovered (SEIR) model, complemented by an observation model that maps latent trajectories to empirical data. Addressing the challenges of intractable likelihoods for parameter inference in stochastic settings, our analysis highlights how these likelihood-free methods provide accurate and robust inference capabilities. The results of our simulation study further underscore the effectiveness of these approaches in capturing the stochastic dynamics of epidemics, providing prediction capabilities for the control of epidemic outbreaks. Results on an Ethiopian cohort study demonstrate operational robustness under real‑world noise and irregular data sampling. To facilitate reuse and to enable building pipelines that ultimately contribute to better informed decision making in public health, we make code and synthetic datasets publicly available.