Paper List
-
Nyxus: A Next Generation Image Feature Extraction Library for the Big Data and AI Era
This paper addresses the core pain point of efficiently extracting standardized, comparable features from massive (terabyte to petabyte-scale) biomedi...
-
Topological Enhancement of Protein Kinetic Stability
This work addresses the long-standing puzzle of why knotted proteins exist by demonstrating that deep knots provide a functional advantage through enh...
-
A Multi-Label Temporal Convolutional Framework for Transcription Factor Binding Characterization
This paper addresses the critical limitation of existing TF binding prediction methods that treat transcription factors as independent entities, faili...
-
Social Distancing Equilibria in Games under Conventional SI Dynamics
This paper solves the core problem of proving the existence and uniqueness of Nash equilibria in finite-duration SI epidemic games, showing they are a...
-
Binding Free Energies without Alchemy
This paper addresses the core bottleneck of computational expense in Absolute Binding Free Energy calculations by eliminating the need for numerous al...
-
SHREC: A Spectral Embedding-Based Approach for Ab-Initio Reconstruction of Helical Molecules
This paper addresses the core bottleneck in cryo-EM helical reconstruction: eliminating the dependency on accurate initial symmetry parameter estimati...
-
Budget-Sensitive Discovery Scoring: A Formally Verified Framework for Evaluating AI-Guided Scientific Selection
This paper addresses the critical gap in evaluating AI-guided scientific selection strategies under realistic budget constraints, where existing metri...
-
Probabilistic Joint and Individual Variation Explained (ProJIVE) for Data Integration
This paper addresses the core challenge of accurately decomposing shared (joint) and dataset-specific (individual) sources of variation in multi-modal...
scCluBench: Comprehensive Benchmarking of Clustering Algorithms for Single-Cell RNA Sequencing
Not specified in provided content
30秒速读
IN SHORT: This paper addresses the critical gap of fragmented and non-standardized benchmarking in single-cell RNA-seq clustering, which hinders objective comparison and selection of appropriate methods for specific biological contexts.
核心创新
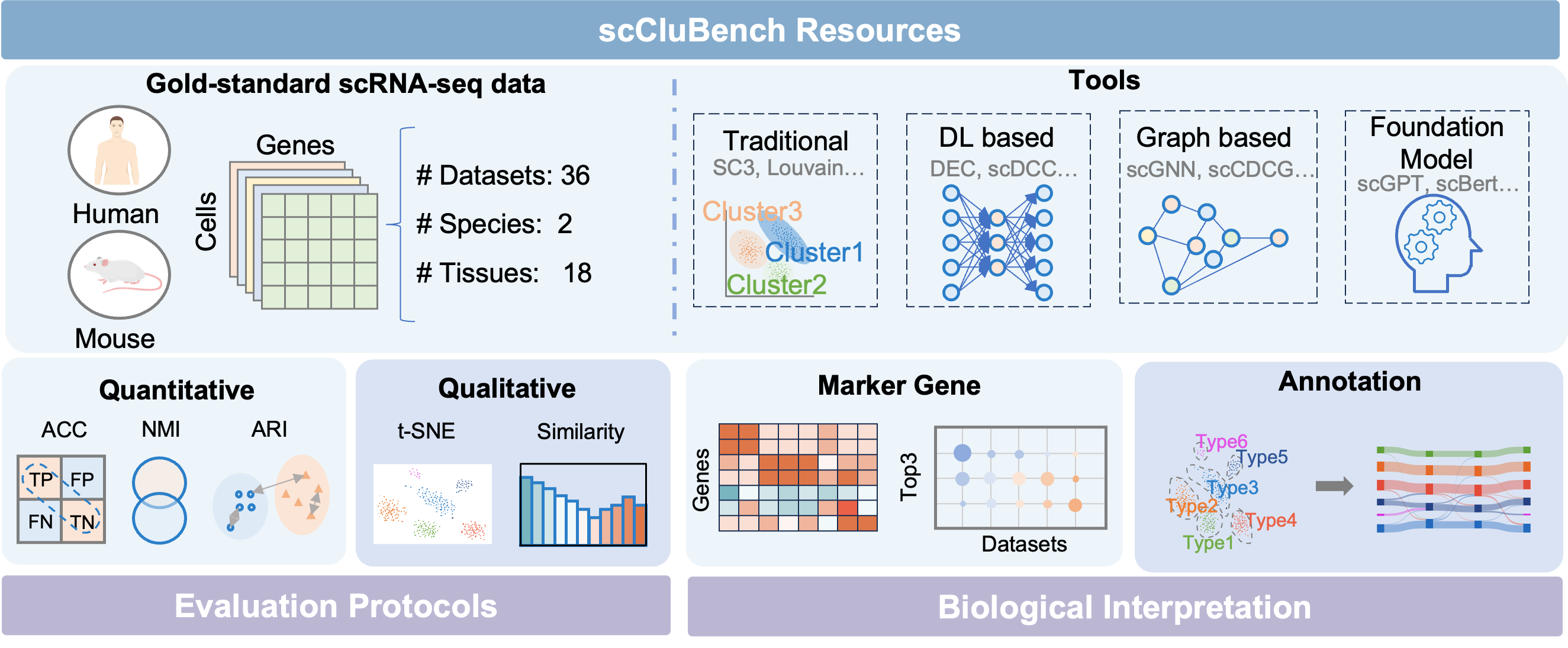
- Methodology Introduces scCluBench, the first comprehensive benchmarking framework that systematically evaluates 16 clustering methods across four categories (traditional, deep learning-based, graph-based, and foundation models) on 36 standardized datasets.
- Methodology Establishes standardized protocols for biological interpretation, including reproducible pipelines for marker gene identification and two distinct cell type annotation approaches (best-mapping and marker-overlap), validated with gold-standard references.
- Methodology Provides a unified and modular benchmarking workflow covering data preprocessing, clustering, and annotation with standardized input-output formats, ensuring reproducibility and fair comparison.
主要结论
- scCDCG (a cut-informed graph embedding model) achieved the highest average clustering accuracy (81.29 ± 1.45) across 36 datasets, outperforming other graph-based, deep learning, and traditional methods.
- Biological foundation models (scGPT, GeneFormer, GeneCompass) showed strong performance in classification tasks (e.g., scGPT achieved 98.14% ACC on Sapiens Ear Crista Ampullaris) but underperformed in direct clustering, highlighting a trade-off between general representation and task-specific optimization.
- The benchmark reveals method-specific limitations: traditional methods struggle with sparse data, deep learning models may fail to capture cell relationships, and graph-based models can suffer from over-smoothing, while most methods decouple embedding learning from clustering optimization.
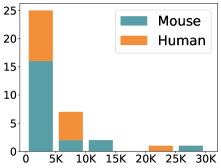
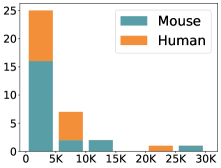
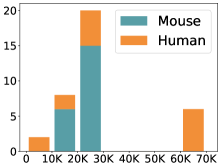
摘要: Cell clustering is crucial for uncovering cellular heterogeneity in single-cell RNA sequencing (scRNA-seq) data by identifying cell types and marker genes. Despite its importance, benchmarks for scRNA-seq clustering methods remain fragmented, often lacking standardized protocols and failing to incorporate recent advances in artificial intelligence. To fill these gaps, we present scCluBench, a comprehensive benchmark of clustering algorithms for scRNA-seq data. First, scCluBench provides 36 scRNA-seq datasets collected from diverse public sources, covering multiple tissues, which are uniformly processed and standardized to ensure consistency for systematic evaluation and downstream analyses. To evaluate performance, we collect and reproduce a range of scRNA-seq clustering methods, including traditional, deep learning-based, graph-based, and biological foundation models. We comprehensively evaluate each method both quantitatively and qualitatively, using core performance metrics as well as visualization analyses. Furthermore, we construct representative downstream biological tasks, such as marker gene identification and cell type annotation, to further assess the practical utility. scCluBench then investigates the performance differences and applicability boundaries of various clustering models across diverse analytical tasks, systematically assessing their robustness and scalability in real-world scenarios. Overall, scCluBench offers a standardized and user-friendly benchmark for scRNA-seq clustering, with curated datasets, unified evaluation protocols, and transparent analyses, facilitating informed method selection and providing valuable insights into model generalizability and application scope.222All datasets, code, and the Extended version for scCluBench are available at the link: https://github.com/XPgogogo/scCluBench. More details for each stage are provided in the extended version.