Paper List
-
SpikGPT: A High-Accuracy and Interpretable Spiking Attention Framework for Single-Cell Annotation
This paper addresses the core challenge of robust single-cell annotation across heterogeneous datasets with batch effects and the critical need to ide...
-
Unlocking hidden biomolecular conformational landscapes in diffusion models at inference time
This paper addresses the core challenge of efficiently and accurately sampling the conformational landscape of biomolecules from diffusion-based struc...
-
Personalized optimization of pediatric HD-tDCS for dose consistency and target engagement
This paper addresses the critical limitation of one-size-fits-all HD-tDCS protocols in pediatric populations by developing a personalized optimization...
-
Realistic Transition Paths for Large Biomolecular Systems: A Langevin Bridge Approach
This paper addresses the core challenge of generating physically realistic and computationally efficient transition paths between distinct protein con...
-
Consistent Synthetic Sequences Unlock Structural Diversity in Fully Atomistic De Novo Protein Design
This paper addresses the core pain point of low sequence-structure alignment in existing synthetic datasets (e.g., AFDB), which severely limits the pe...
-
MoRSAIK: Sequence Motif Reactor Simulation, Analysis and Inference Kit in Python
This work addresses the computational bottleneck in simulating prebiotic RNA reactor dynamics by developing a Python package that tracks sequence moti...
-
On the Approximation of Phylogenetic Distance Functions by Artificial Neural Networks
This paper addresses the core challenge of developing computationally efficient and scalable neural network architectures that can learn accurate phyl...
-
EcoCast: A Spatio-Temporal Model for Continual Biodiversity and Climate Risk Forecasting
This paper addresses the critical bottleneck in conservation: the lack of timely, high-resolution, near-term forecasts of species distribution shifts ...
Training Dynamics of Learning 3D-Rotational Equivariance
Genentech Computational Sciences | New York University
30秒速读
IN SHORT: This work addresses the core dilemma of whether to use computationally expensive equivariant architectures or faster symmetry-agnostic models with data augmentation, by quantifying the speed and extent to which the latter learn 3D rotational symmetry.
核心创新
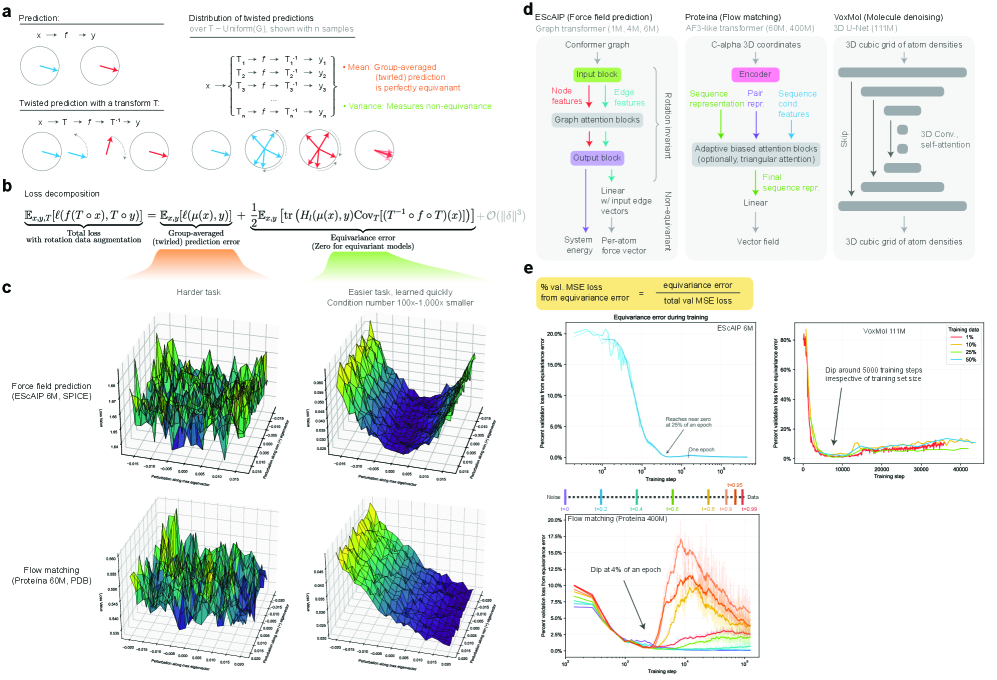
- Methodology Introduces a principled, generalizable framework to decompose total loss into a 'twirled prediction error' (ℒ_mean) and an 'equivariance error' (ℒ_equiv), enabling precise measurement of the percent of loss attributable to imperfect symmetry learning.
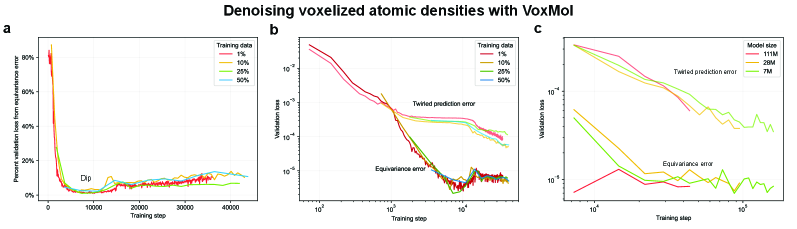
- Methodology Empirically demonstrates that models learning 3D-rotational equivariance via data augmentation achieve very low equivariance error (≤2% of total loss) remarkably quickly, within 1k-10k training steps, across diverse molecular tasks and model scales.
- Theory Provides theoretical and experimental evidence that learning equivariance is an easier task than the main prediction, characterized by a smoother and better-conditioned loss landscape (e.g., 1000x lower condition number for ℒ_equiv vs. ℒ_mean in force field prediction).
主要结论
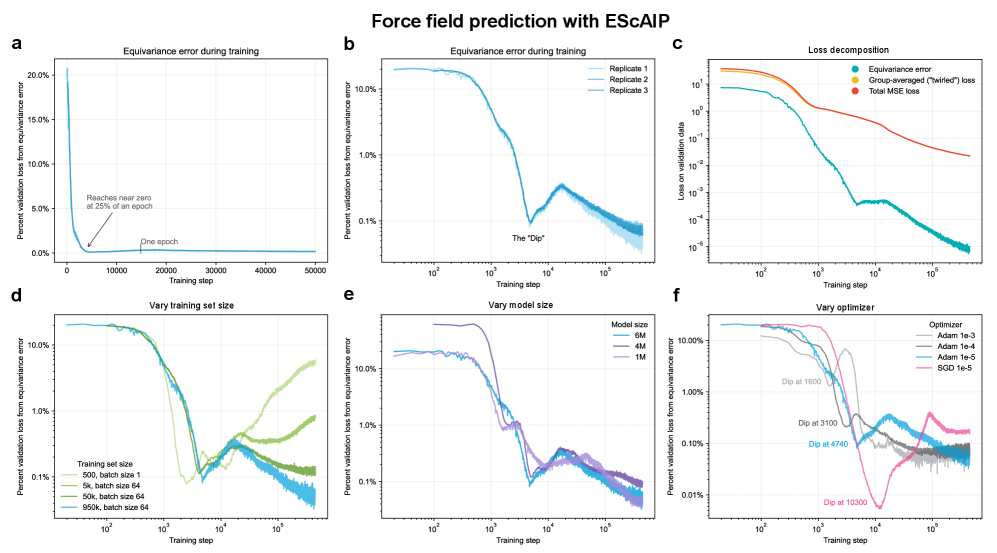
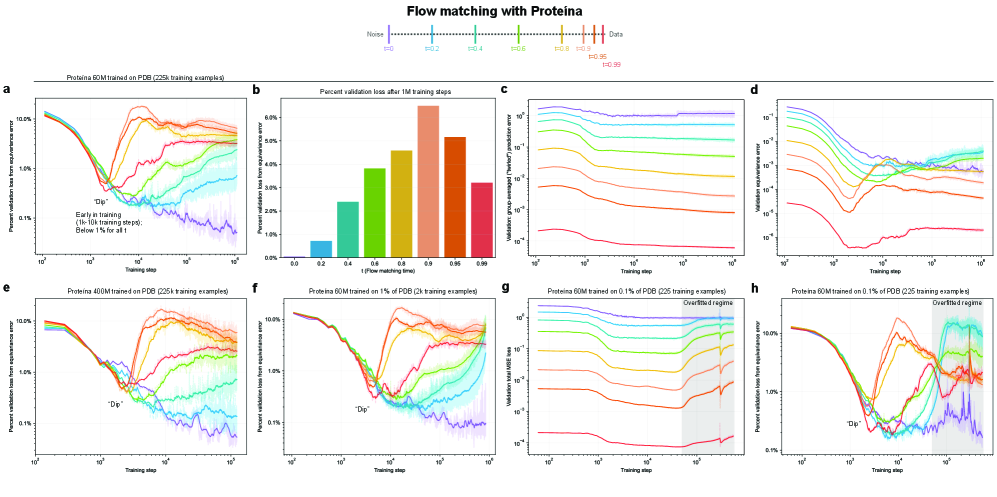
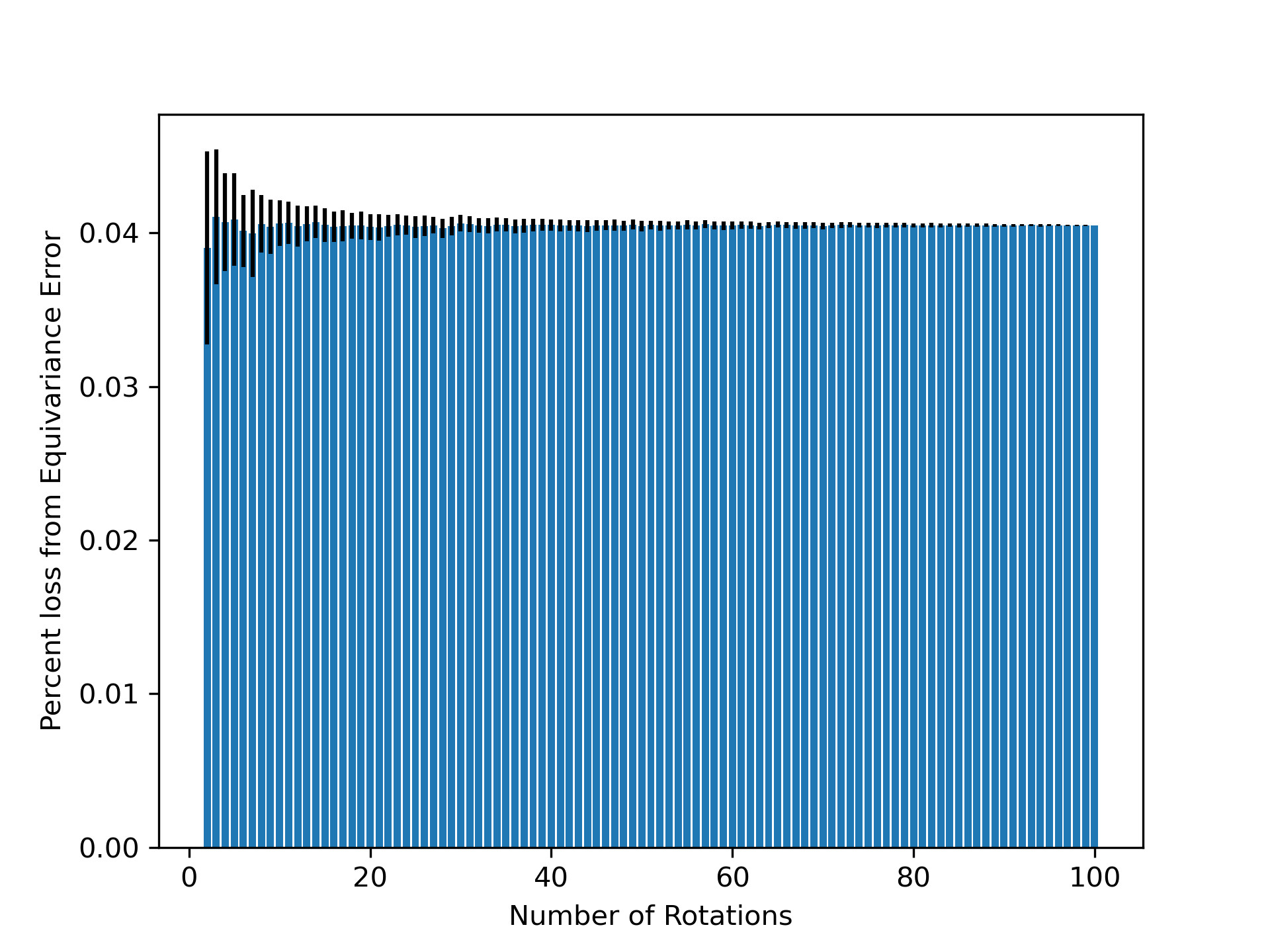
- Non-equivariant models with data augmentation learn 3D rotational equivariance rapidly and effectively, reducing the equivariance error component to ≤2% of the total validation loss within the first 1k-10k training steps.
- The loss penalty for imperfect equivariance (ℒ_equiv) is small throughout training for 3D rotations, meaning the primary trade-off is the 'efficiency gap' (slower training/inference of equivariant models) rather than a significant accuracy penalty.
- The speed of learning equivariance is robust to model size (1M to 400M parameters), dataset size (500 to 1M samples), and optimizer choice, indicating it is a fundamental property of the learning task landscape.
摘要: While data augmentation is widely used to train symmetry-agnostic models, it remains unclear how quickly and effectively they learn to respect symmetries. We investigate this by deriving a principled measure of equivariance error that, for convex losses, calculates the percent of total loss attributable to imperfections in learned symmetry. We focus our empirical investigation to 3D-rotation equivariance on high-dimensional molecular tasks (flow matching, force field prediction, denoising voxels) and find that models reduce equivariance error quickly to ≤2% held-out loss within 1k-10k training steps, a result robust to model and dataset size. This happens because learning 3D-rotational equivariance is an easier learning task, with a smoother and better-conditioned loss landscape, than the main prediction task. For 3D rotations, the loss penalty for non-equivariant models is small throughout training, so they may achieve lower test loss than equivariant models per GPU-hour unless the equivariant “efficiency gap” is narrowed. We also experimentally and theoretically investigate the relationships between relative equivariance error, learning gradients, and model parameters.