Paper List
-
SpikGPT: A High-Accuracy and Interpretable Spiking Attention Framework for Single-Cell Annotation
This paper addresses the core challenge of robust single-cell annotation across heterogeneous datasets with batch effects and the critical need to ide...
-
Unlocking hidden biomolecular conformational landscapes in diffusion models at inference time
This paper addresses the core challenge of efficiently and accurately sampling the conformational landscape of biomolecules from diffusion-based struc...
-
Personalized optimization of pediatric HD-tDCS for dose consistency and target engagement
This paper addresses the critical limitation of one-size-fits-all HD-tDCS protocols in pediatric populations by developing a personalized optimization...
-
Realistic Transition Paths for Large Biomolecular Systems: A Langevin Bridge Approach
This paper addresses the core challenge of generating physically realistic and computationally efficient transition paths between distinct protein con...
-
Consistent Synthetic Sequences Unlock Structural Diversity in Fully Atomistic De Novo Protein Design
This paper addresses the core pain point of low sequence-structure alignment in existing synthetic datasets (e.g., AFDB), which severely limits the pe...
-
MoRSAIK: Sequence Motif Reactor Simulation, Analysis and Inference Kit in Python
This work addresses the computational bottleneck in simulating prebiotic RNA reactor dynamics by developing a Python package that tracks sequence moti...
-
On the Approximation of Phylogenetic Distance Functions by Artificial Neural Networks
This paper addresses the core challenge of developing computationally efficient and scalable neural network architectures that can learn accurate phyl...
-
EcoCast: A Spatio-Temporal Model for Continual Biodiversity and Climate Risk Forecasting
This paper addresses the critical bottleneck in conservation: the lack of timely, high-resolution, near-term forecasts of species distribution shifts ...
Mapping of Lesion Images to Somatic Mutations
University of Illinois at Chicago | University of Texas MD Anderson Cancer Center
30秒速读
IN SHORT: This paper addresses the critical bottleneck of delayed genetic analysis in cancer diagnosis by predicting a patient's full somatic mutation profile directly from medical lesion images, enabling earlier targeted treatment decisions.
核心创新
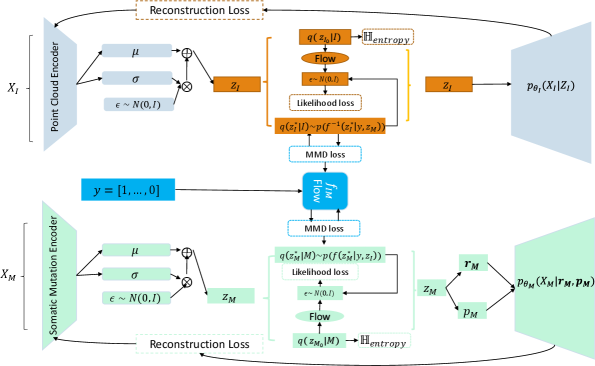
- Methodology Proposes LLOST, a novel architecture with dual VAEs and a separate, cancer-type-conditioned shared latent space, coupled with domain-specific conditional Normalizing Flow priors to handle heterogeneous data distributions.
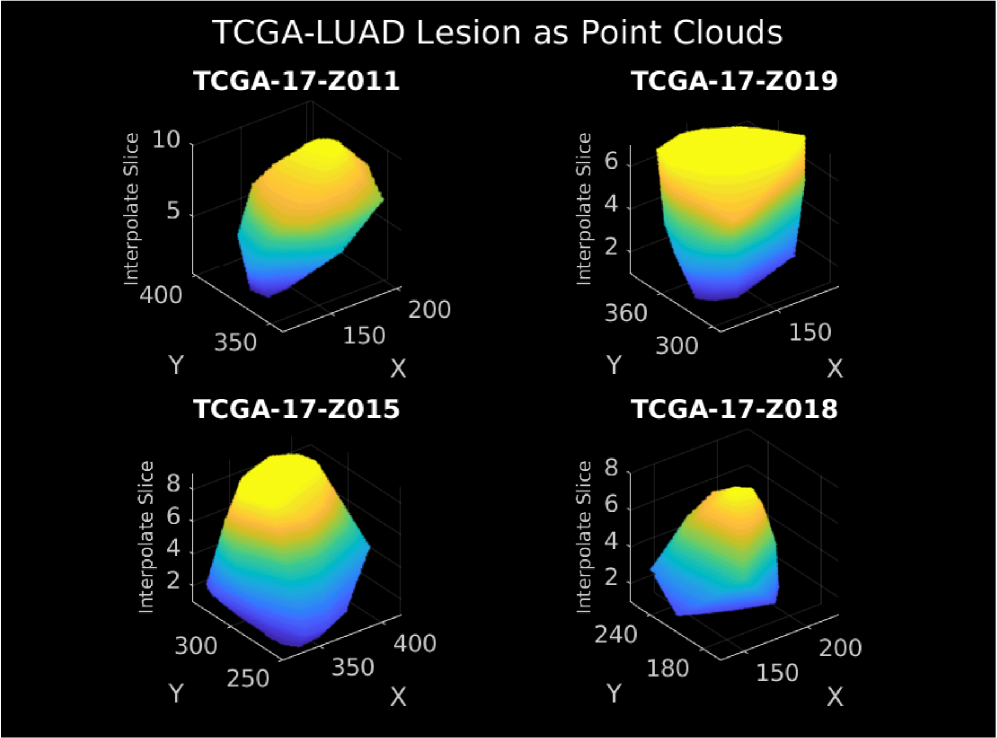
- Methodology Introduces a modality-invariant point cloud representation for lesion images, overcoming challenges of multi-slice, multi-modal (CT/MRI) medical imaging data.
- Methodology Employs a Negative-Binomial likelihood within the mutation VAE to effectively model the high-dimensional, sparse, and discrete nature of somatic mutation count data.
主要结论
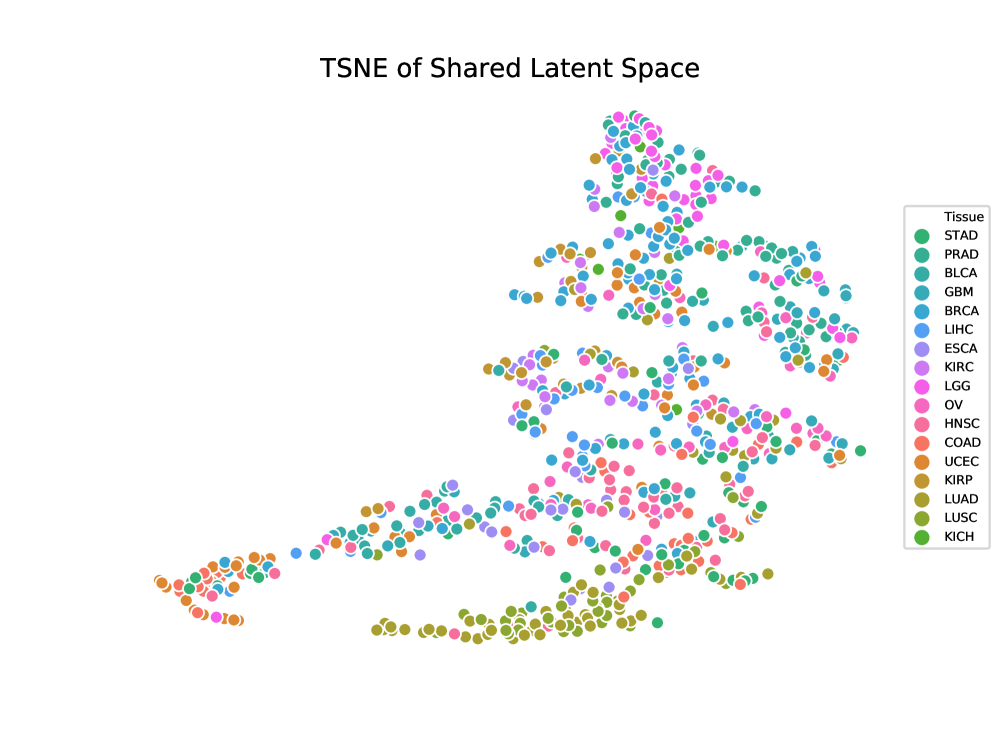
- LLOST successfully learns a shared latent representation between lesion point clouds and somatic mutation counts, capturing cancer-type-specific patterns across these disparate domains.
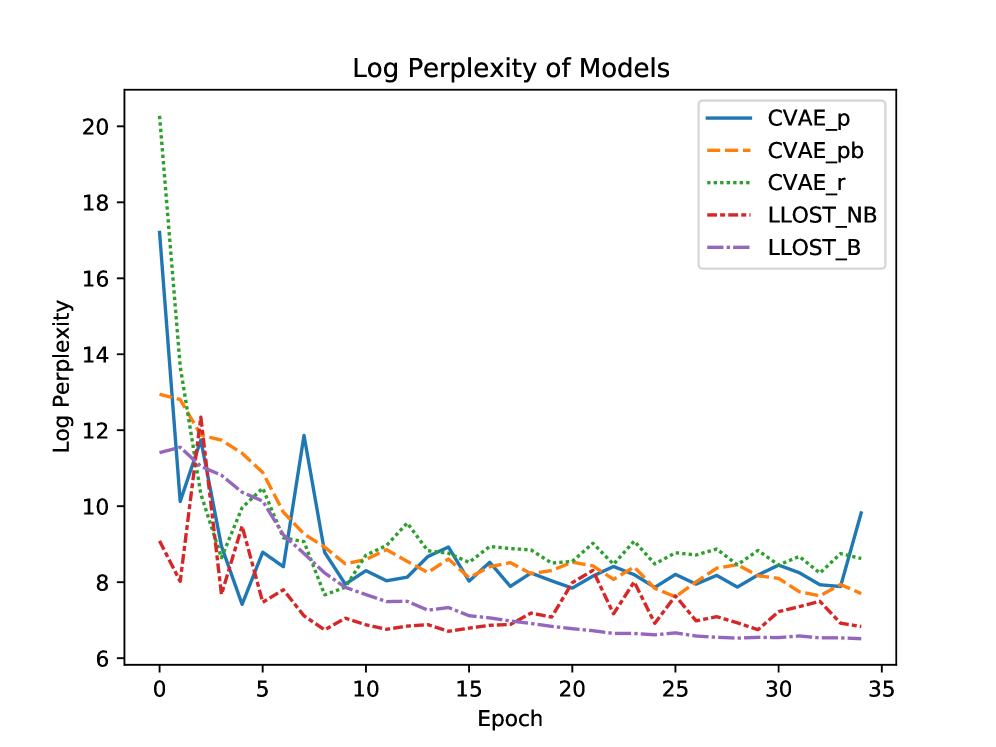
- The model demonstrates predictive capability for both mutation occurrence (binary prediction) and mutation counts, validated on a dataset of 1342 patients across 18 cancer types from TCGA/TCIA.
- The use of conditional Normalizing Flow priors and a separate shared latent space allows the model to account for and bridge the complex, distinct distributions of imaging and genomic data.
摘要: Medical imaging is a critical initial tool used by clinicians to determine a patient’s cancer diagnosis, allowing for faster intervention and more reliable patient prognosis. At subsequent stages of patient diagnosis, genetic information is extracted to help select specific patient treatment options. As the efficacy of cancer treatment often relies on early diagnosis and treatment, we build a deep latent variable model to determine patients’ somatic mutation profiles based on their corresponding medical images. We first introduce a point cloud representation of lesions images to allow for invariance to the imaging modality. We then propose, LLOST, a model with dual variational autoencoders coupled together by a separate shared latent space that unifies features from the lesion point clouds and counts of distinct somatic mutations. Therefore our model consists of three latent space, each of which is learned with a conditional normalizing flow prior to account for the diverse distributions of each domain. We conduct qualitative and quantitative experiments on de-identified medical images from The Cancer Imaging Archive and the corresponding somatic mutations from the Pan Cancer dataset of The Cancer Genomic Archive. We show the model’s predictive performance on the counts of specific mutations as well as it’s ability to accurately predict the occurrence of mutations. In particular, shared patterns between the imaging and somatic mutation domain that reflect cancer type. We conclude with a remark on how to improve the model and possible future avenues of research to include other genetic domains.