Paper List
-
Macroscopic Dominance from Microscopic Extremes: Symmetry Breaking in Spatial Competition
This paper addresses the fundamental question of how microscopic stochastic advantages in spatial exploration translate into macroscopic resource domi...
-
Linear Readout of Neural Manifolds with Continuous Variables
This paper addresses the core challenge of quantifying how the geometric structure of high-dimensional neural population activity (neural manifolds) d...
-
Theory of Cell Body Lensing and Phototaxis Sign Reversal in “Eyeless” Mutants of Chlamydomonas
This paper solves the core puzzle of how eyeless mutants of Chlamydomonas exhibit reversed phototaxis by quantitatively modeling the competition betwe...
-
Cross-Species Transfer Learning for Electrophysiology-to-Transcriptomics Mapping in Cortical GABAergic Interneurons
This paper addresses the challenge of predicting transcriptomic identity from electrophysiological recordings in human cortical interneurons, where li...
-
Uncovering statistical structure in large-scale neural activity with Restricted Boltzmann Machines
This paper addresses the core challenge of modeling large-scale neural population activity (1500-2000 neurons) with interpretable higher-order interac...
-
Realizing Common Random Numbers: Event-Keyed Hashing for Causally Valid Stochastic Models
This paper addresses the critical problem that standard stateful PRNG implementations in agent-based models violate causal validity by making random d...
-
A Standardized Framework for Evaluating Gene Expression Generative Models
This paper addresses the critical lack of standardized evaluation protocols for single-cell gene expression generative models, where inconsistent metr...
-
Single Molecule Localization Microscopy Challenge: A Biologically Inspired Benchmark for Long-Sequence Modeling
This paper addresses the core challenge of evaluating state-space models on biologically realistic, sparse, and stochastic temporal processes, which a...
Generative design and validation of therapeutic peptides for glioblastoma based on a potential target ATP5A
Shanghai Jiao Tong University | QuietD Biotech
30秒速读
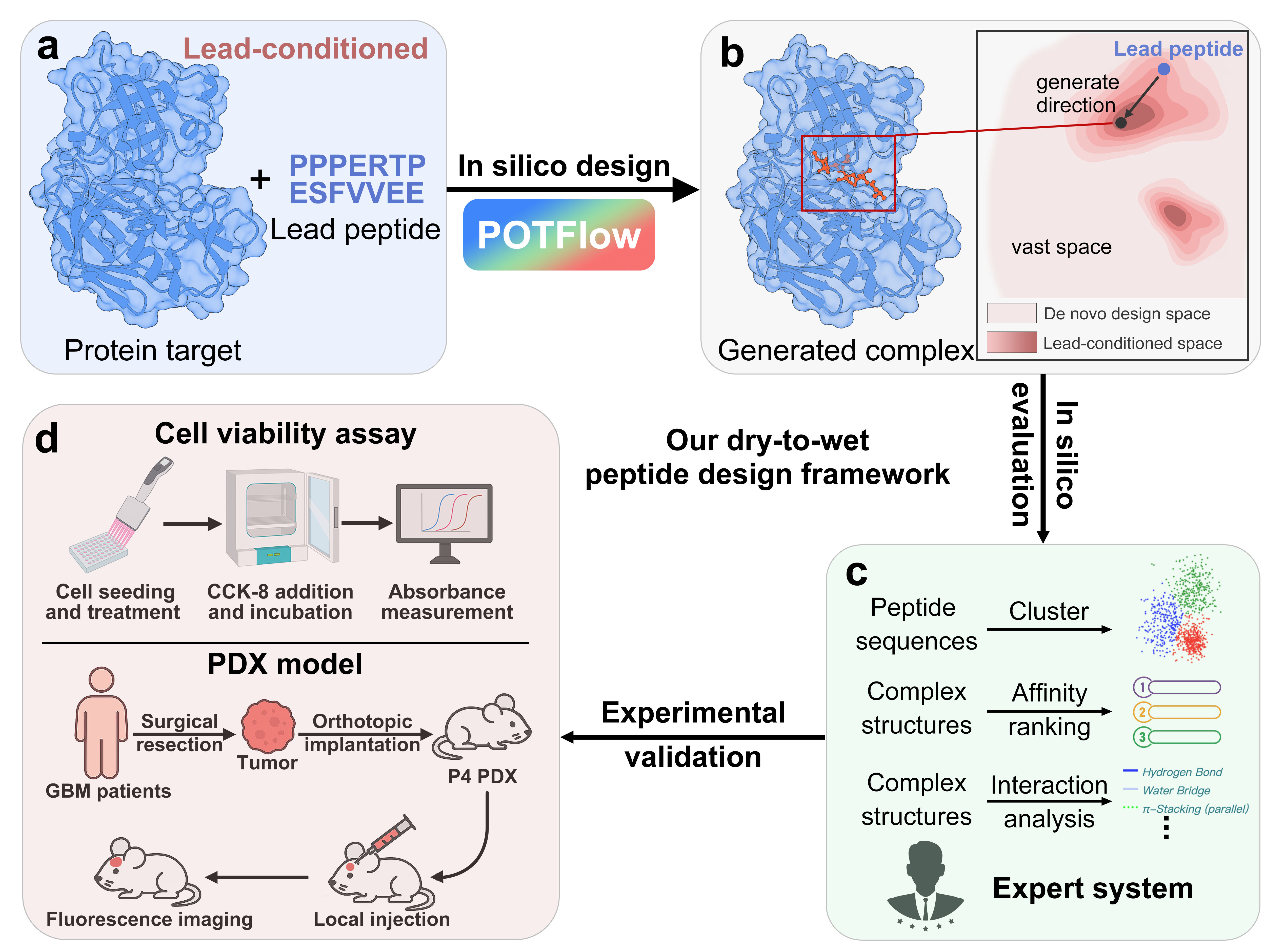
IN SHORT: This paper addresses the critical bottleneck in therapeutic peptide design: how to efficiently optimize lead peptides with geometric constraints while bridging the gap between computational generation and experimental validation.
核心创新
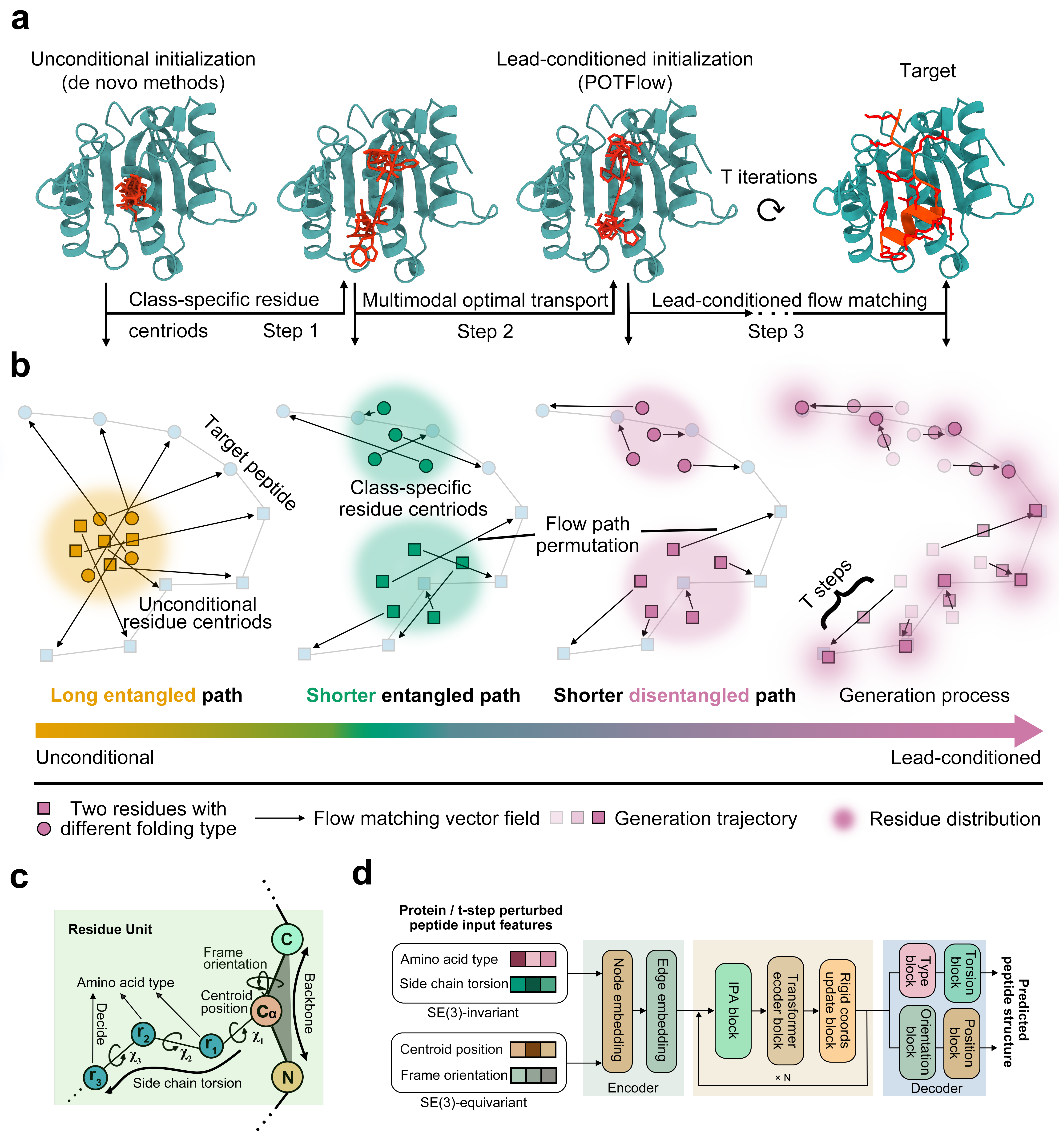
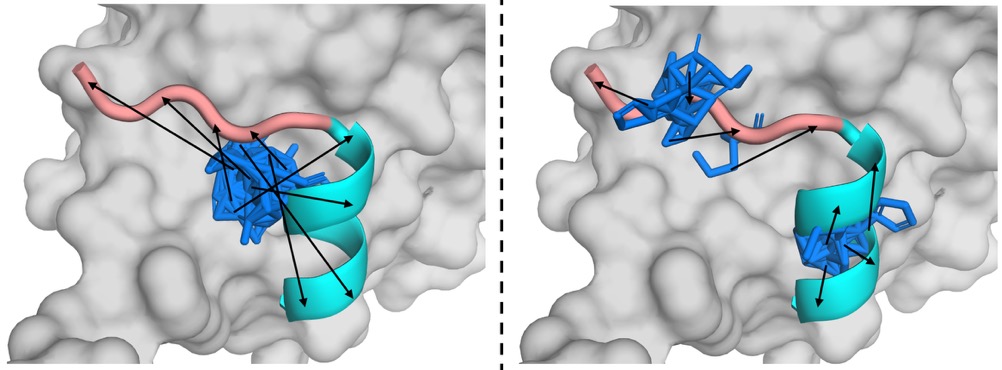
- Methodology Introduces POTFlow, the first lead peptide-conditioned flow matching model that incorporates secondary structure priors and optimal transport for shorter, disentangled generation paths
- Methodology Proposes a dry-to-wet framework that integrates computational design with experimental validation spanning in vitro assays and in vivo PDX models
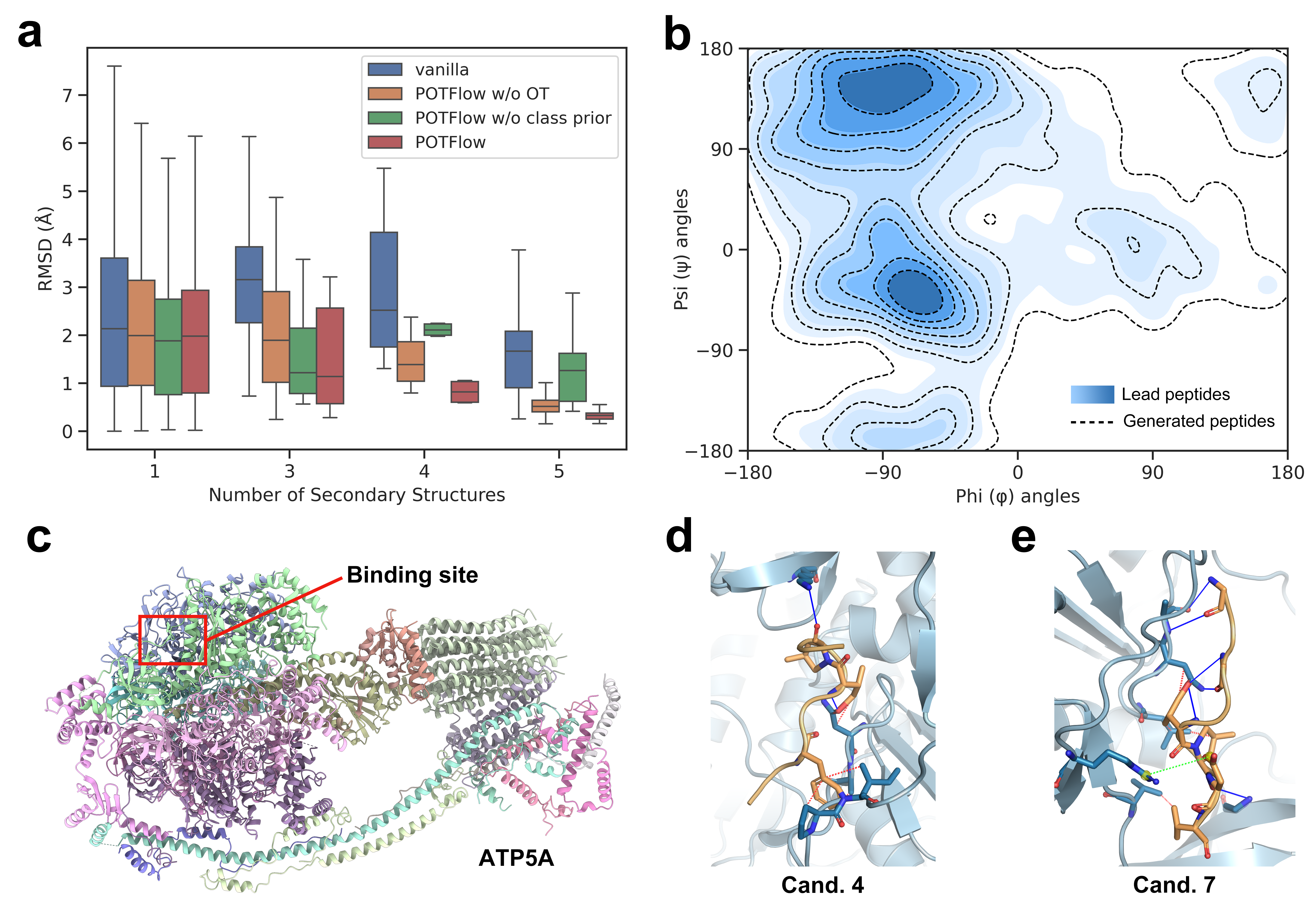
- Biology Demonstrates successful optimization of ATP5A-binding peptides for glioblastoma, achieving improved tumor selectivity and in vivo efficacy
主要结论
- POTFlow outperforms five state-of-the-art methods across multiple metrics, achieving 53.44% similarity, 95.07% compactness, 30.56% affinity, and 1.66Å RMSD on benchmark datasets
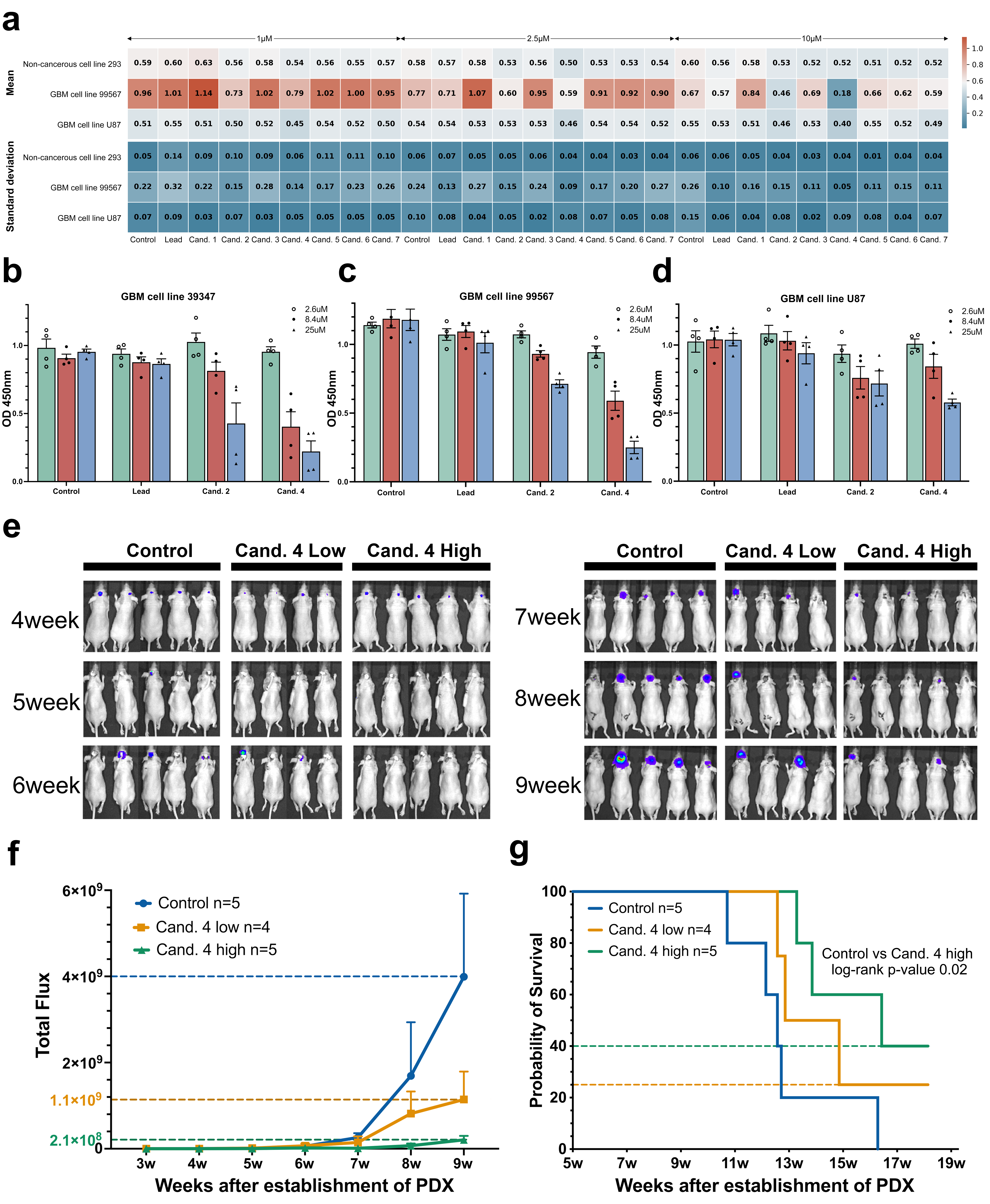
- Generated peptide candidates showed 18-68% higher inhibition of viability rate (IVR) in GBM cells compared to non-cancerous cells (<10%), demonstrating improved tumor selectivity
- High-dose candidate 4 (20mg/kg) significantly prolonged survival in PDX models (p-value = 0.02) with 40% of mice surviving beyond week 18 compared to 0% in control group
摘要: Glioblastoma (GBM) remains the most aggressive tumor, urgently requiring novel therapeutic strategies. Here, we present a dry-to-wet framework combining generative modeling and experimental validation to optimize peptides targeting ATP5A, a potential peptide-binding protein for GBM. Our framework introduces the first lead-conditioned generative model, which focuses exploration on geometrically relevant regions around lead peptides and mitigates the combinatorial complexity of de novo methods. Specifically, we propose POTFlow, a Prior and Optimal Transport-based Flow-matching model for peptide optimization. POTFlow employs secondary structure information (e.g., helix, sheet, loop) as geometric constraints, which are further refined by optimal transport to produce shorter flow paths. With this design, our method achieves state-of-the-art performance compared with five popular approaches. When applied to GBM, our method generates peptides that selectively inhibit cell viability and significantly prolong survival in a patient-derived xenograft (PDX) model. As the first lead peptide-conditioned flow matching model, POTFlow holds strong potential as a generalizable framework for therapeutic peptide design.