Paper List
-
Nyxus: A Next Generation Image Feature Extraction Library for the Big Data and AI Era
This paper addresses the core pain point of efficiently extracting standardized, comparable features from massive (terabyte to petabyte-scale) biomedi...
-
Topological Enhancement of Protein Kinetic Stability
This work addresses the long-standing puzzle of why knotted proteins exist by demonstrating that deep knots provide a functional advantage through enh...
-
A Multi-Label Temporal Convolutional Framework for Transcription Factor Binding Characterization
This paper addresses the critical limitation of existing TF binding prediction methods that treat transcription factors as independent entities, faili...
-
Social Distancing Equilibria in Games under Conventional SI Dynamics
This paper solves the core problem of proving the existence and uniqueness of Nash equilibria in finite-duration SI epidemic games, showing they are a...
-
Binding Free Energies without Alchemy
This paper addresses the core bottleneck of computational expense in Absolute Binding Free Energy calculations by eliminating the need for numerous al...
-
SHREC: A Spectral Embedding-Based Approach for Ab-Initio Reconstruction of Helical Molecules
This paper addresses the core bottleneck in cryo-EM helical reconstruction: eliminating the dependency on accurate initial symmetry parameter estimati...
-
Budget-Sensitive Discovery Scoring: A Formally Verified Framework for Evaluating AI-Guided Scientific Selection
This paper addresses the critical gap in evaluating AI-guided scientific selection strategies under realistic budget constraints, where existing metri...
-
Probabilistic Joint and Individual Variation Explained (ProJIVE) for Data Integration
This paper addresses the core challenge of accurately decomposing shared (joint) and dataset-specific (individual) sources of variation in multi-modal...
Hierarchical Molecular Language Models (HMLMs)
Department of Chemical Engineering, University of Arkansas, Fayetteville, AR 72701, USA
30秒速读
IN SHORT: This paper addresses the core challenge of accurately modeling context-dependent signaling, pathway cross-talk, and temporal dynamics across multiple biological scales in cellular signaling networks.
核心创新
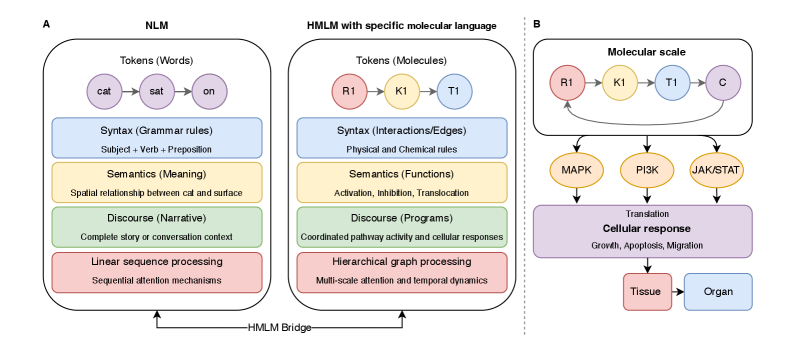
- Methodology Introduces cellular signaling as a molecular language with unique grammar and semantics, establishing a theoretical foundation for molecular artificial intelligence (MAI).
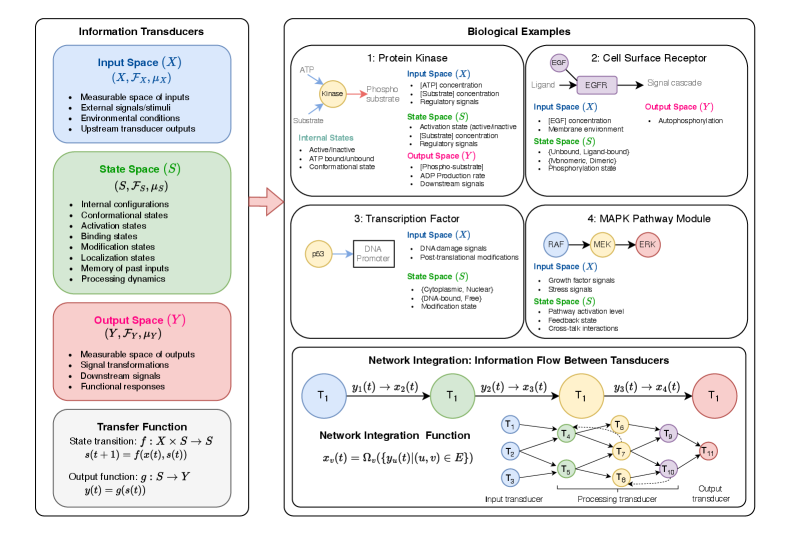
- Methodology Develops HMLMs as a novel computational architecture adapting transformer architecture to model signaling networks as information-processing systems across molecular, pathway, and cellular scales.
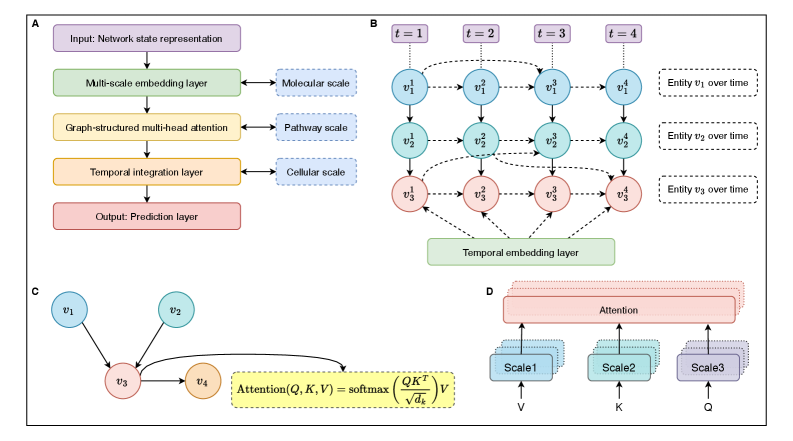
- Methodology Implements graph-structured attention mechanisms and hierarchical scale-bridging operators (aggregation, decomposition, translation) to accommodate signaling network topology and multi-scale organization.
主要结论
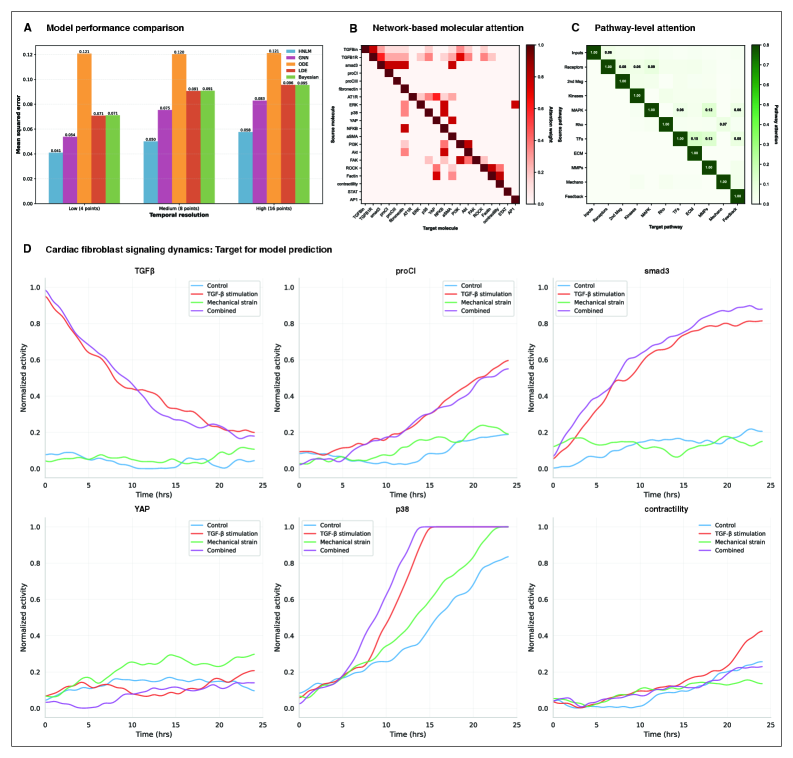
- HMLM achieved MSE of 0.058 for temporal signaling predictions, representing 30% improvement over GNNs (0.083) and 52% improvement over ODE models (0.121).
- Under sparse temporal sampling with only 4 timepoints, HMLM maintained superior performance with MSE = 0.041, demonstrating robustness to limited temporal data.
- Attention mechanisms identified biologically plausible pathway interactions including mechanotransduction-MAPK coupling and TGFβ to ERK signaling, validating the model's ability to capture meaningful biological relationships.
摘要: Cellular signaling networks represent complex information processing systems that have been modeled via traditional mathematical or statistical approaches. However, these methods often struggle to capture context-dependent signaling, pathway cross-talk, and temporal dynamics across multiple biological scales. Here, we introduce hierarchical molecular language models (HMLMs), a novel architecture that proposes a molecular network-specifiac large language model (LLM) to use in intracellular communication as a specialized molecular language, which includes molecules as tokens, protein interactions, post-translational modifications, and regulatory events modeled as semantic relationships within an adapted transformer architecture. HMLMs employ graph-structured attention mechanisms to accommodate signaling network topology while integrating information across the molecular, pathway, and cellular scales through hierarchical attention patterns. We demonstrate HMLM superiority using a cardiac fibroblast signaling network comprising over 100 molecular species across functional modules connected by regulatory edges. HMLM achieved a mean squared error (MSE) of 0.058 for temporal signaling predictions, representing 30% improvement over graph neural networks (GNNs: 0.083) and 52% improvement over ordinary differential equation models (ODEs: 0.121), with particular advantages under sparse temporal sampling conditions where HMLM maintained MSE = 0.041 with only 4 timepoints. The attention-based computational analysis identified key inter-pathway cross-talk patterns through learned attention mechanisms, including mechanotransduction-MAPK coupling and TGFβ to ERK signaling, demonstrating the model's capability to capture biologically plausible pathway interactions from network topology and temporal dynamics and convergent regulatory mechanisms controlling fibrosis markers in simulated cardiac fibroblast networks. The HMLMs offer a foundation for AI-driven biology and medicine with predictable scaling characteristics suitable for interactive applications. By bridging molecular mechanisms with cellular phenotypes through AI-driven molecular language representation, HMLMs provide a powerful paradigm for systems biology that advances precision medicine applications and therapeutic discovery in the era of AI.