Paper List
-
Nyxus: A Next Generation Image Feature Extraction Library for the Big Data and AI Era
This paper addresses the core pain point of efficiently extracting standardized, comparable features from massive (terabyte to petabyte-scale) biomedi...
-
Topological Enhancement of Protein Kinetic Stability
This work addresses the long-standing puzzle of why knotted proteins exist by demonstrating that deep knots provide a functional advantage through enh...
-
A Multi-Label Temporal Convolutional Framework for Transcription Factor Binding Characterization
This paper addresses the critical limitation of existing TF binding prediction methods that treat transcription factors as independent entities, faili...
-
Social Distancing Equilibria in Games under Conventional SI Dynamics
This paper solves the core problem of proving the existence and uniqueness of Nash equilibria in finite-duration SI epidemic games, showing they are a...
-
Binding Free Energies without Alchemy
This paper addresses the core bottleneck of computational expense in Absolute Binding Free Energy calculations by eliminating the need for numerous al...
-
SHREC: A Spectral Embedding-Based Approach for Ab-Initio Reconstruction of Helical Molecules
This paper addresses the core bottleneck in cryo-EM helical reconstruction: eliminating the dependency on accurate initial symmetry parameter estimati...
-
Budget-Sensitive Discovery Scoring: A Formally Verified Framework for Evaluating AI-Guided Scientific Selection
This paper addresses the critical gap in evaluating AI-guided scientific selection strategies under realistic budget constraints, where existing metri...
-
Probabilistic Joint and Individual Variation Explained (ProJIVE) for Data Integration
This paper addresses the core challenge of accurately decomposing shared (joint) and dataset-specific (individual) sources of variation in multi-modal...
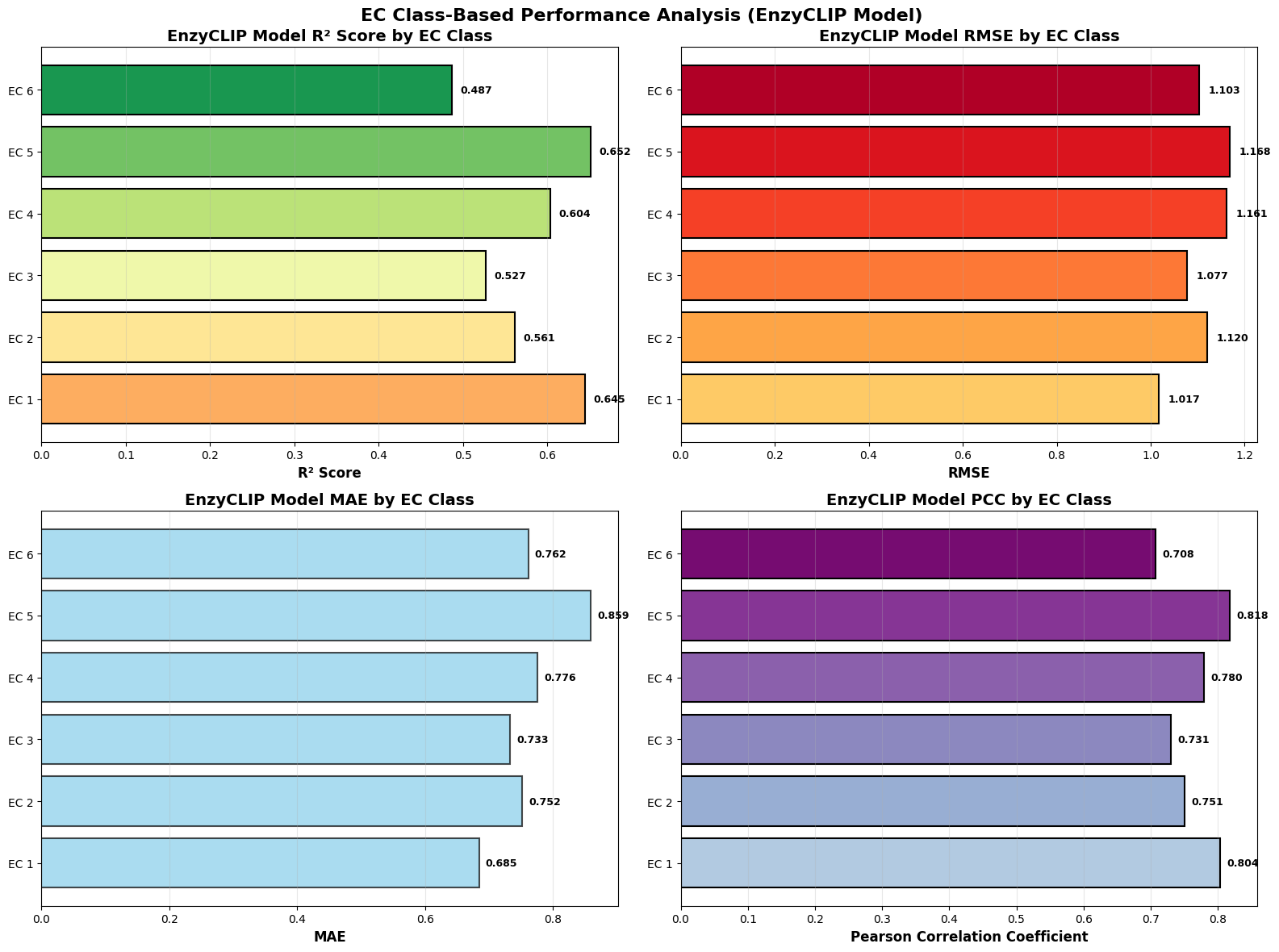
EnzyCLIP: A Cross-Attention Dual Encoder Framework with Contrastive Learning for Predicting Enzyme Kinetic Constants
Vellore Institute of Technology | BIT (Department of Computer Science) | BIT (Department of Bioengineering and Biotechnology)
30秒速读
IN SHORT: This paper addresses the core challenge of jointly predicting enzyme kinetic parameters (Kcat and Km) by modeling dynamic enzyme-substrate interactions through a multimodal contrastive learning framework.
核心创新
- Methodology Proposes a CLIP-inspired dual-encoder architecture with bidirectional cross-attention that dynamically models enzyme-substrate interactions, overcoming the limitation of separate processing in existing methods.
- Methodology Integrates contrastive learning (InfoNCE loss) with multi-task regression (Huber loss) to learn aligned multimodal representations while jointly predicting both Kcat and Km parameters.
- Biology Addresses the critical gap in existing literature that typically focuses on single parameter prediction (mainly Kcat) by providing a unified framework for joint prediction of both fundamental kinetic constants.
主要结论
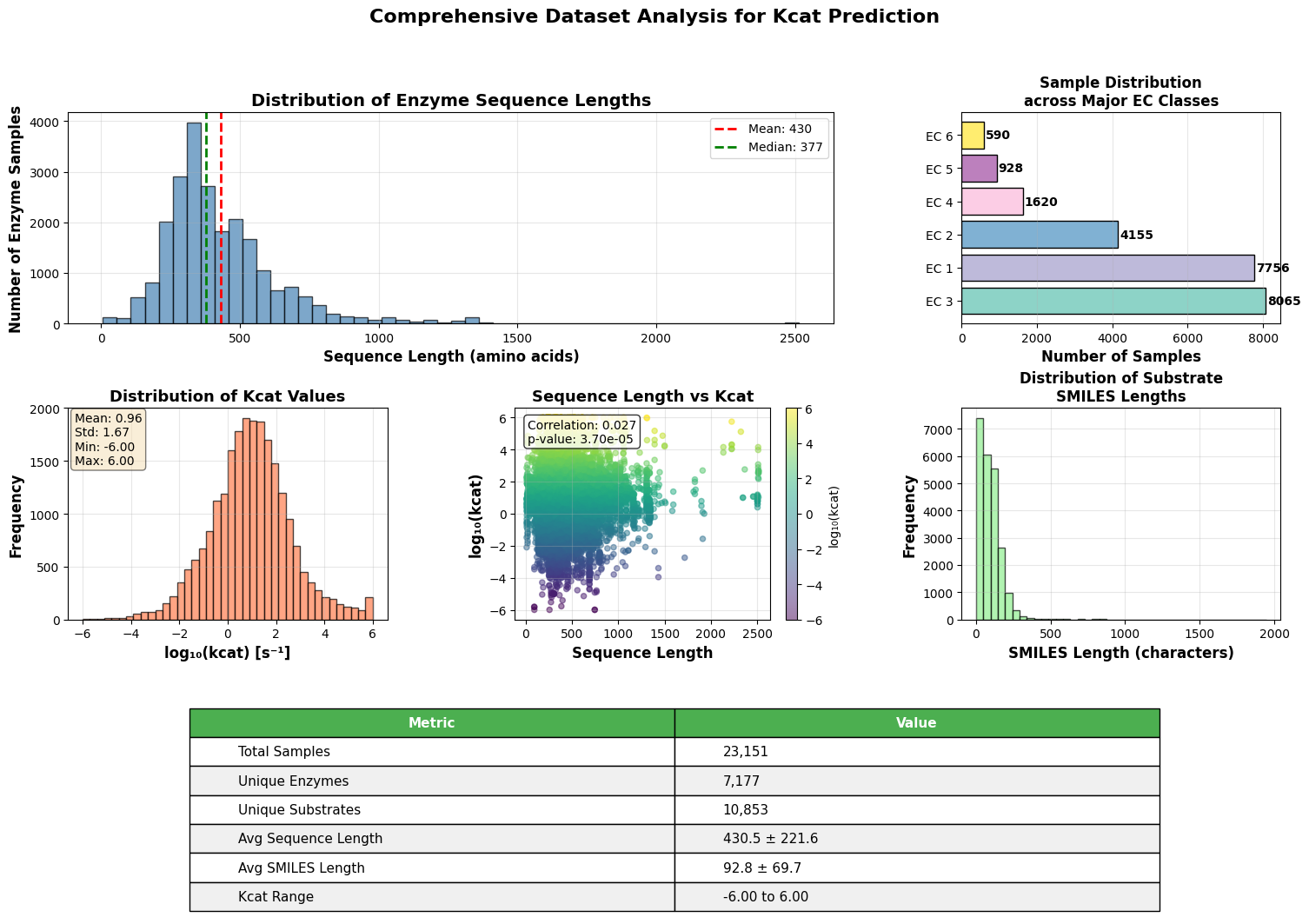
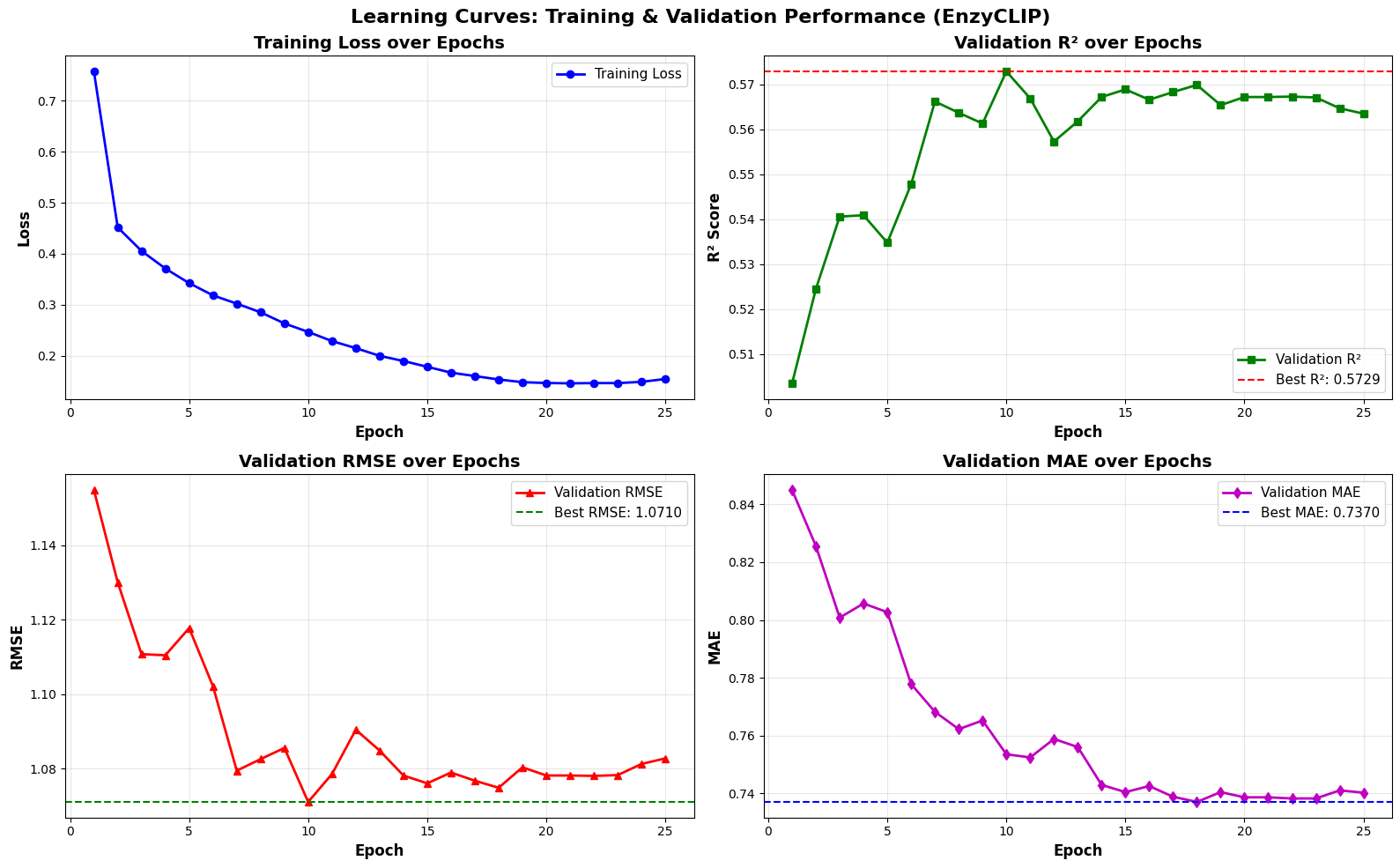
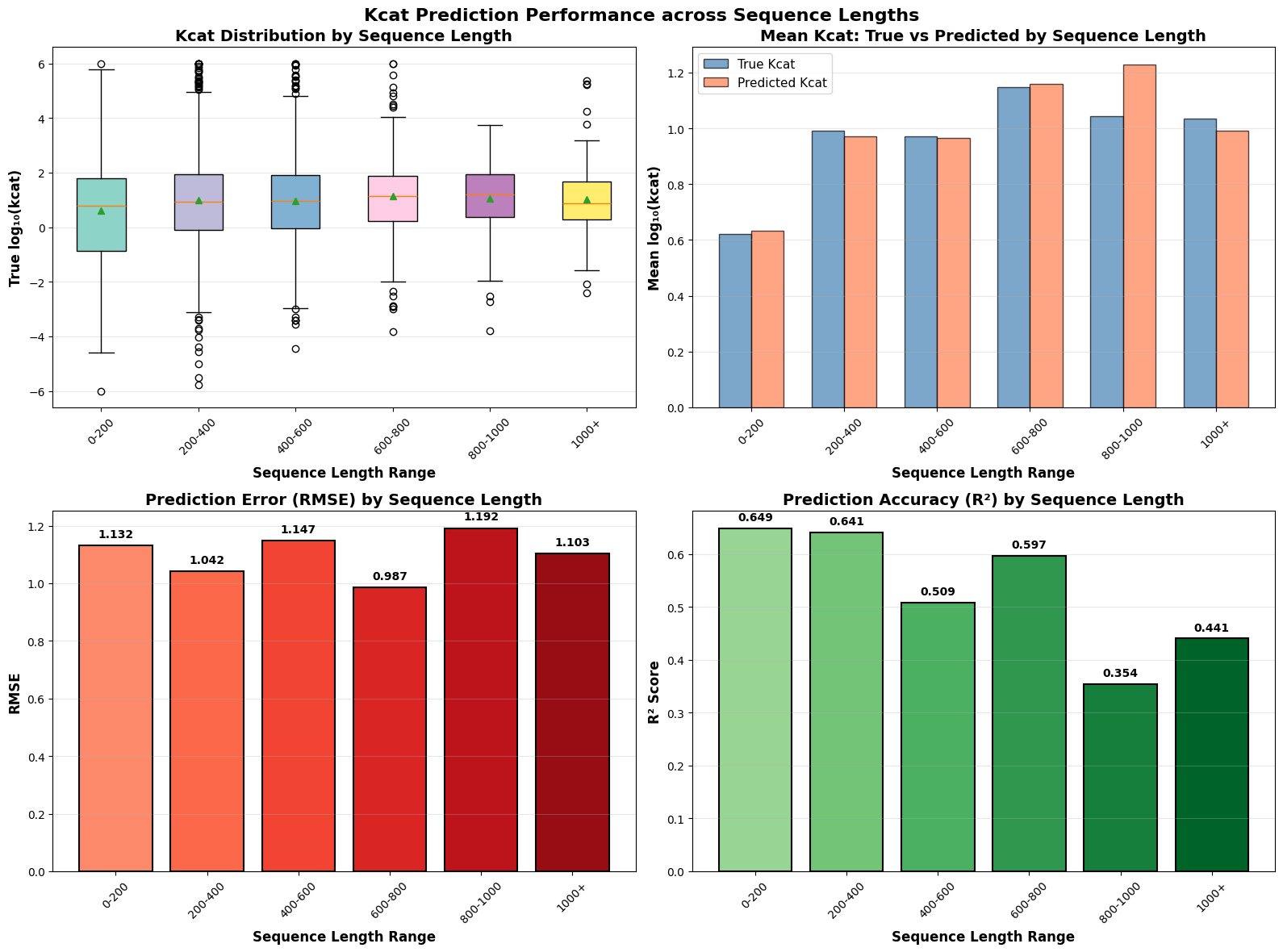
- EnzyCLIP achieves competitive baseline performance with R² scores of 0.593 for Kcat and 0.607 for Km prediction on the CatPred-DB dataset containing 23,151 Kcat and 41,174 Km measurements.
- The integration of contrastive learning with cross-attention mechanisms enables the model to capture biochemical relationships and substrate preferences even for unseen enzyme-substrate pairs.
- XGBoost ensemble methods applied to learned embeddings further improved Km prediction performance to R² = 0.61 while maintaining robust Kcat prediction capabilities.
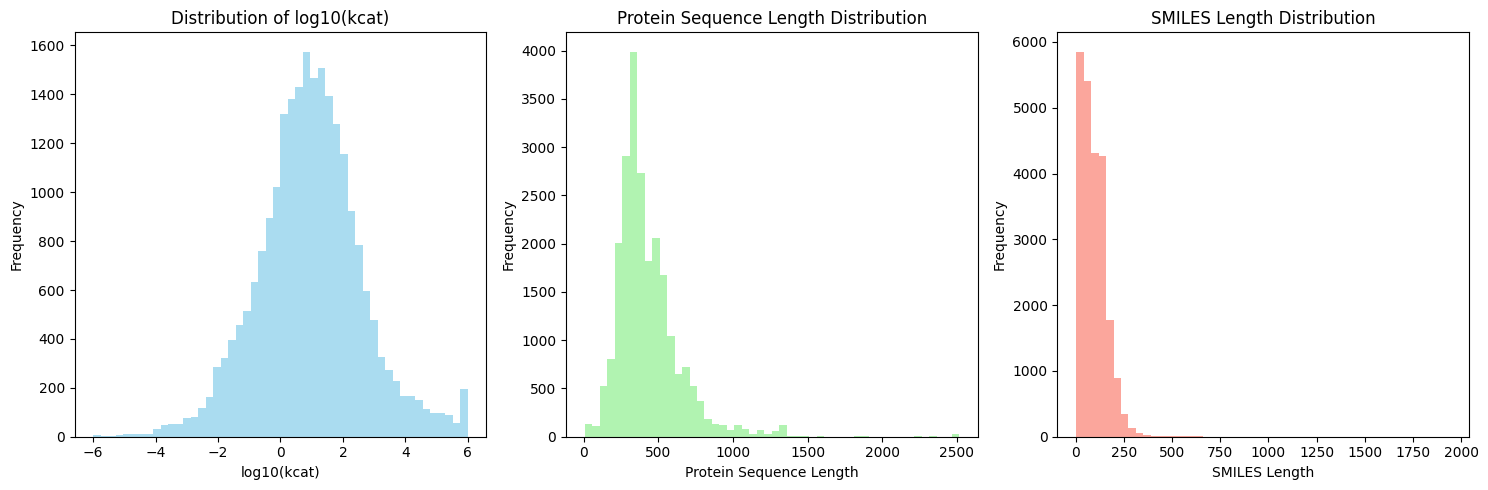
摘要: Accurate prediction of enzyme kinetic parameters is crucial for drug discovery, metabolic engineering, and synthetic biology applications. Current computational approaches face limitations in capturing complex enzyme–substrate interactions and often focus on single parameters while neglecting the joint prediction of catalytic turnover numbers (Kcat) and Michaelis–Menten constants (Km). We present EnzyCLIP, a novel dual-encoder framework that leverages contrastive learning and cross-attention mechanisms to predict enzyme kinetic parameters from protein sequences and substrate molecular structures. Our approach integrates ESM-2 protein language model embeddings with ChemBERTa chemical representations through a CLIP-inspired architecture enhanced with bidirectional cross-attention for dynamic enzyme–substrate interaction modeling. EnzyCLIP combines InfoNCE contrastive loss with Huber regression loss to learn aligned multimodal representations while predicting log10-transformed kinetic parameters. EnzyCLIP is trained on the CatPred-DB database containing 23,151 Kcat and 41,174 Km experimentally validated measurements, and achieved competitive baseline performance with R2 scores of 0.593 for Kcat and 0.607 for Km prediction. XGBoost ensemble methods on learned embeddings further improved Km prediction (R2 = 0.61) while maintaining robust Kcat performance.