Paper List
-
A Theoretical Framework for the Formation of Large Animal Groups: Topological Coordination, Subgroup Merging, and Velocity Inheritance
This paper addresses the core problem of how large, coordinated animal groups form in nature, challenging the classical view of gradual aggregation by...
-
CONFIDE: Hallucination Assessment for Reliable Biomolecular Structure Prediction and Design
This paper addresses the critical limitation of current protein structure prediction models (like AlphaFold3) where high-confidence scores (pLDDT) can...
-
Generative design and validation of therapeutic peptides for glioblastoma based on a potential target ATP5A
This paper addresses the critical bottleneck in therapeutic peptide design: how to efficiently optimize lead peptides with geometric constraints while...
-
Pharmacophore-based design by learning on voxel grids
This paper addresses the computational bottleneck and limited novelty in conventional pharmacophore-based virtual screening by introducing a voxel cap...
-
Human-Centred Evaluation of Text-to-Image Generation Models for Self-expression of Mental Distress: A Dataset Based on GPT-4o
This paper addresses the critical gap in evaluating how AI-generated images can effectively support cross-cultural mental distress communication, part...
-
ANNE Apnea Paper
This paper addresses the core challenge of achieving accurate, event-level sleep apnea detection and characterization using a non-intrusive, multimoda...
-
DeeDeeExperiment: Building an infrastructure for integrating and managing omics data analysis results in R/Bioconductor
This paper addresses the critical bottleneck of managing and organizing the growing volume of differential expression and functional enrichment analys...
-
Cross-Species Antimicrobial Resistance Prediction from Genomic Foundation Models
This paper addresses the core challenge of predicting antimicrobial resistance across phylogenetically distinct bacterial species, where traditional m...
DeeDeeExperiment: Building an infrastructure for integrating and managing omics data analysis results in R/Bioconductor
Institute of Medical Biostatistics, Epidemiology and Informatics (IMBEI), University Medical Center Mainz | Research Center for Immunotherapy (FZI) Mainz | Department of Nephrology, Rheumatology and Kidney Transplantation, University Medical Center Mainz
30秒速读
IN SHORT: This paper addresses the critical bottleneck of managing and organizing the growing volume of differential expression and functional enrichment analysis results from complex omics experiments, which currently lack standardized data structures for storage and contextualization.
核心创新
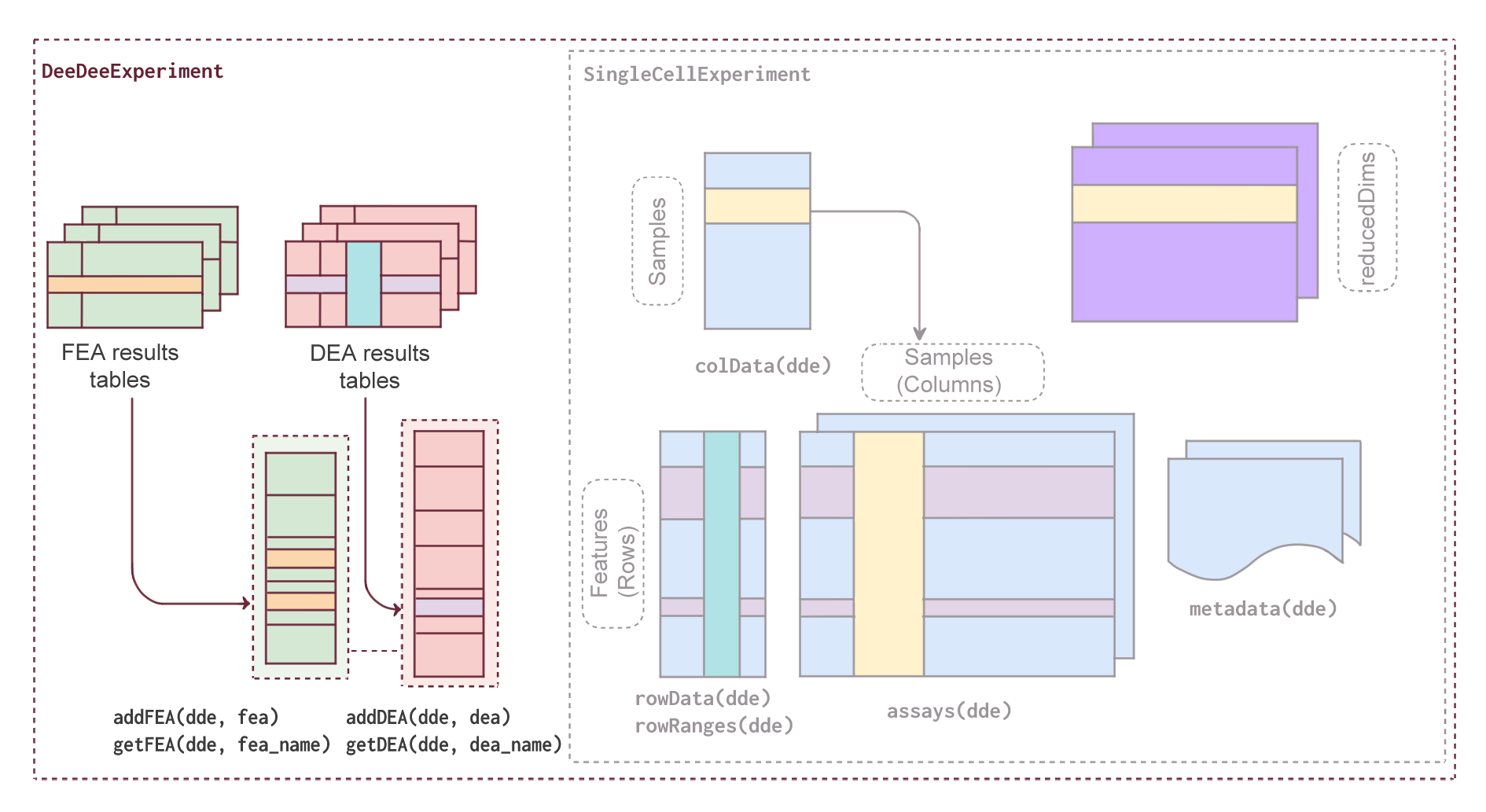
- Methodology Introduces the first standardized S4 class specifically designed to co-store DEA and FEA results with their metadata in a single, structured container within the Bioconductor ecosystem.
- Methodology Extends the widely adopted SingleCellExperiment class by adding dedicated slots for DEA and FEA results while maintaining full backward compatibility with existing Bioconductor tools.
- Methodology Implements a contrast-centric architecture that organizes results from multiple comparisons (including limma multi-contrast objects and muscat pseudobulk analyses) with efficient storage through pointer-based referencing.
主要结论
- DeeDeeExperiment provides a robust, standardized framework that enables efficient organization and retrieval of DEA/FEA results across multiple contrasts within a single data object.
- The implementation maintains full compatibility with the Bioconductor ecosystem, supporting interoperability with downstream tools like scater for visualization and iSEE for interactive exploration.
- By consolidating analysis results and metadata, the framework supports more nuanced quantitative approaches beyond simple overlap strategies, enabling trustworthy summaries of complex experimental measurements.
摘要: Summary: Modern omics experiments now involve multiple conditions and complex designs, producing an increasingly large set of differential expression and functional enrichment analysis results. However, no standardized data structure exists to store and contextualize these results together with their metadata, leaving researchers with an unmanageable and potentially non-reproducible collection of results that are difficult to navigate and/or share. Here we introduce DeeDeeExperiment, a new S4 class for managing and storing omics data analysis results, implemented within the Bioconductor ecosystem, which promotes interoperability, reproducibility and good documentation. This class extends the widely used SingleCellExperiment object by introducing dedicated slots for Differential Expression (DEA) and Functional Enrichment Analysis (FEA) results, allowing users to organize, store, and retrieve information on multiple contrasts and associated metadata within a single data object, ultimately streamlining the management and interpretation of many omics datasets. Availability and implementation: DeeDeeExperiment is available on Bioconductor under the MIT license (https://bioconductor.org/packages/DeeDeeExperiment), with its development version also available on Github (https://github.com/imbeimainz/DeeDeeExperiment).