Paper List
-
A Theoretical Framework for the Formation of Large Animal Groups: Topological Coordination, Subgroup Merging, and Velocity Inheritance
This paper addresses the core problem of how large, coordinated animal groups form in nature, challenging the classical view of gradual aggregation by...
-
CONFIDE: Hallucination Assessment for Reliable Biomolecular Structure Prediction and Design
This paper addresses the critical limitation of current protein structure prediction models (like AlphaFold3) where high-confidence scores (pLDDT) can...
-
Generative design and validation of therapeutic peptides for glioblastoma based on a potential target ATP5A
This paper addresses the critical bottleneck in therapeutic peptide design: how to efficiently optimize lead peptides with geometric constraints while...
-
Pharmacophore-based design by learning on voxel grids
This paper addresses the computational bottleneck and limited novelty in conventional pharmacophore-based virtual screening by introducing a voxel cap...
-
Human-Centred Evaluation of Text-to-Image Generation Models for Self-expression of Mental Distress: A Dataset Based on GPT-4o
This paper addresses the critical gap in evaluating how AI-generated images can effectively support cross-cultural mental distress communication, part...
-
ANNE Apnea Paper
This paper addresses the core challenge of achieving accurate, event-level sleep apnea detection and characterization using a non-intrusive, multimoda...
-
DeeDeeExperiment: Building an infrastructure for integrating and managing omics data analysis results in R/Bioconductor
This paper addresses the critical bottleneck of managing and organizing the growing volume of differential expression and functional enrichment analys...
-
Cross-Species Antimicrobial Resistance Prediction from Genomic Foundation Models
This paper addresses the core challenge of predicting antimicrobial resistance across phylogenetically distinct bacterial species, where traditional m...
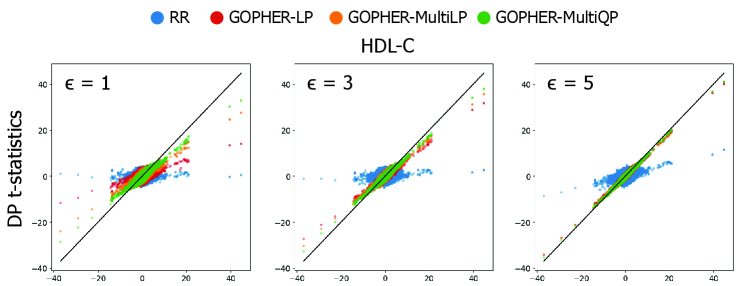
GOPHER: Optimization-based Phenotype Randomization for Genome-Wide Association Studies with Differential Privacy
Department of Biomedical Informatics & Data Science, Yale School of Medicine | Department of Technology and Operations Management, Harvard Business School | Department of Computer Science, Yale University
30秒速读
IN SHORT: This paper addresses the core challenge of balancing rigorous privacy protection with data utility when releasing full GWAS summary statistics, overcoming the limitations of prior methods that either add excessive noise or restrict output to a small subset of results.
核心创新
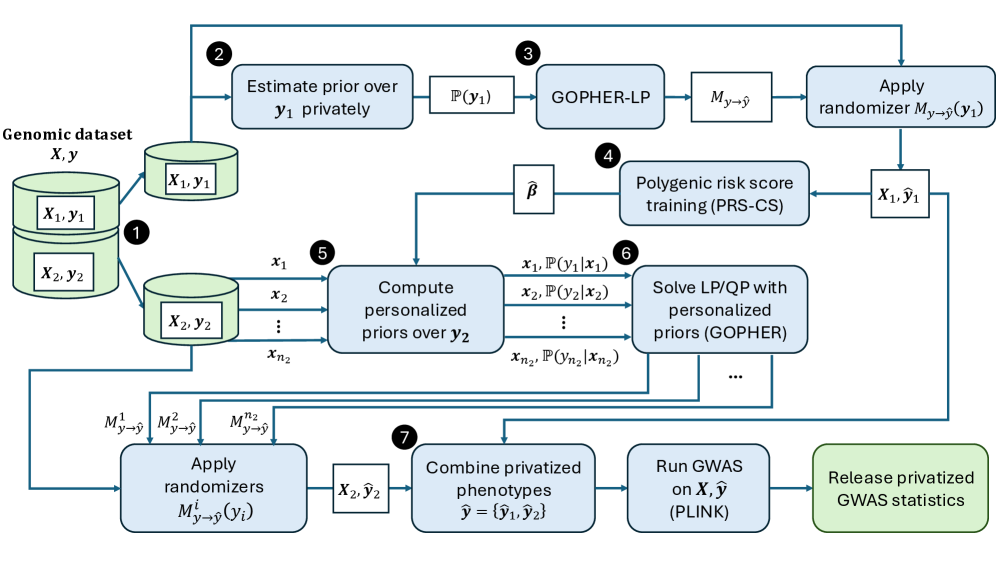
- Methodology Introduces an optimization-based phenotype randomization mechanism (GOPHER-LP) that directly minimizes expected error in GWAS statistics, formulated as a linear programming problem to enhance utility beyond baseline methods like randomized response.
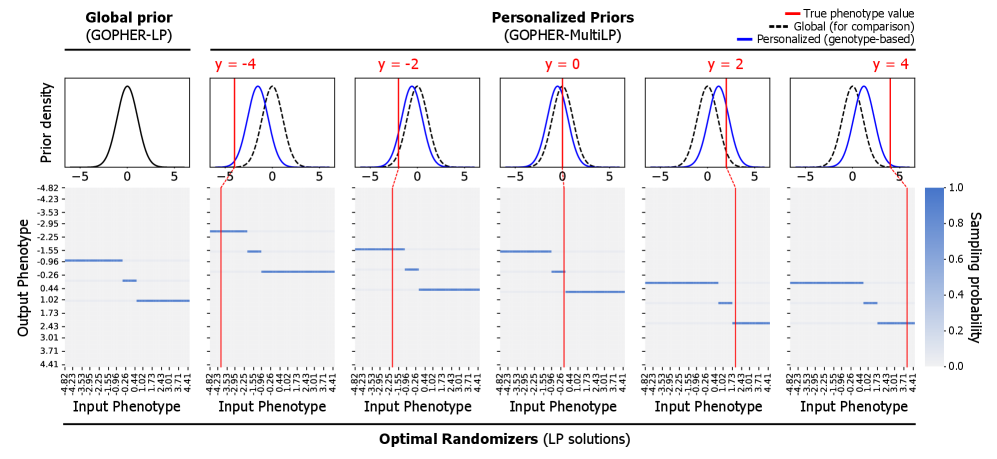
- Methodology Proposes GOPHER-MultiLP, which incorporates personalized priors derived from predictive models (e.g., polygenic risk scores) trained on a held-out subset, enabling sample-specific optimization that leverages genotype information to further reduce noise.
- Theory Adopts and extends the concept of phenotypic differential privacy (analogous to label DP), focusing protection on sensitive phenotypes while treating genotypes as public, providing a practical middle ground between full DP and unrestricted release.
主要结论
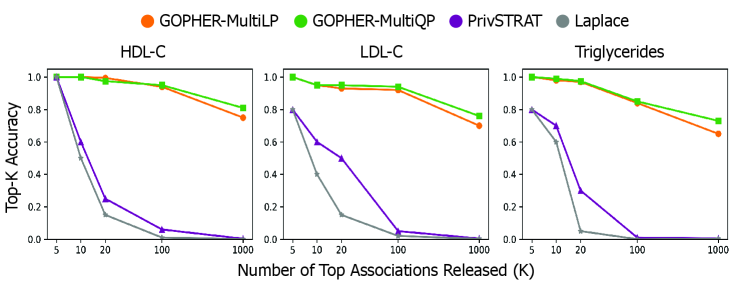
- The GOPHER framework enables the release of complete GWAS statistics (e.g., over 500,000 variants) with provable privacy guarantees, a significant scalability advance over prior methods limited to releasing only 3-5 top associations.
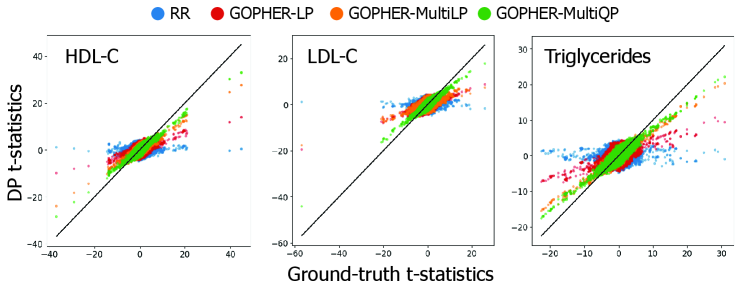
- Experiments on UK Biobank data (n=100,000) demonstrate that the mechanisms yield association statistics that accurately match non-private GWAS results while maintaining rigorous (ε, δ)-DP guarantees.
- The phenotype-randomization approach decouples the added noise from the number of genetic variants analyzed, addressing a fundamental scalability challenge not previously solved in the DP-GWAS literature.
摘要: Genome-wide association studies (GWAS) are an essential tool in biomedical research for identifying genetic factors linked to health and disease. However, publicly releasing GWAS summary statistics poses well-recognized privacy risks, including the potential to infer an individual’s participation in the study or to reveal sensitive phenotypic information (e.g., disease status). While differential privacy (DP) offers a rigorous mathematical framework for mitigating these risks, existing DP techniques for GWAS either introduce excessive noise or restrict the release to a limited set of results. In this work, we present practical DP mechanisms for releasing the complete set of genome-wide association statistics with privacy guarantees. We demonstrate the accuracy of the privacy-preserving statistics released by our mechanisms on a range of GWAS datasets from the UK Biobank, utilizing both real and simulated phenotypes. We introduce two key techniques to overcome the limitations of prior approaches: (1) an optimization-based randomization mechanism that directly minimizes the expected error in GWAS results to enhance utility, and (2) the use of personalized priors, derived from predictive models privately trained on a subset of the dataset, to enable sample-specific optimization which further reduces the amount of noise introduced by DP. Overall, our work provides practical tools for accurately releasing comprehensive GWAS results with provable protection of study participants.