Paper List
-
A Theoretical Framework for the Formation of Large Animal Groups: Topological Coordination, Subgroup Merging, and Velocity Inheritance
This paper addresses the core problem of how large, coordinated animal groups form in nature, challenging the classical view of gradual aggregation by...
-
CONFIDE: Hallucination Assessment for Reliable Biomolecular Structure Prediction and Design
This paper addresses the critical limitation of current protein structure prediction models (like AlphaFold3) where high-confidence scores (pLDDT) can...
-
Generative design and validation of therapeutic peptides for glioblastoma based on a potential target ATP5A
This paper addresses the critical bottleneck in therapeutic peptide design: how to efficiently optimize lead peptides with geometric constraints while...
-
Pharmacophore-based design by learning on voxel grids
This paper addresses the computational bottleneck and limited novelty in conventional pharmacophore-based virtual screening by introducing a voxel cap...
-
Human-Centred Evaluation of Text-to-Image Generation Models for Self-expression of Mental Distress: A Dataset Based on GPT-4o
This paper addresses the critical gap in evaluating how AI-generated images can effectively support cross-cultural mental distress communication, part...
-
ANNE Apnea Paper
This paper addresses the core challenge of achieving accurate, event-level sleep apnea detection and characterization using a non-intrusive, multimoda...
-
DeeDeeExperiment: Building an infrastructure for integrating and managing omics data analysis results in R/Bioconductor
This paper addresses the critical bottleneck of managing and organizing the growing volume of differential expression and functional enrichment analys...
-
Cross-Species Antimicrobial Resistance Prediction from Genomic Foundation Models
This paper addresses the core challenge of predicting antimicrobial resistance across phylogenetically distinct bacterial species, where traditional m...
Imperfect molecular detection renormalizes apparent kinetic rates in stochastic gene regulatory networks
Department of Mathematical Analysis and Numerical Mathematics, Comenius University, Slovakia | University of Edinburgh, UK
30秒速读
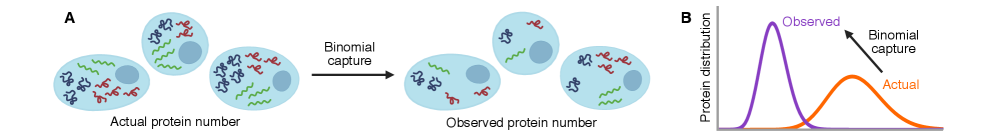
IN SHORT: This paper addresses the core challenge of distinguishing genuine stochastic dynamics of gene regulatory networks from artifacts introduced by imperfect molecular detection in single-cell experiments.
核心创新
- Methodology Extends the binomial capture model from simple gene expression to general gene regulatory networks (GRNs) with explicit regulation, enabling analysis of technical noise in complex systems.
- Theory Establishes precise mathematical conditions under which technical noise leads to a renormalization (rescaling) of kinetic rates versus when it introduces non-absorbable distortions.
- Methodology Derives results valid for networks of arbitrary connectivity and under time-dependent kinetic rates, significantly generalizing previous steady-state analyses.
主要结论
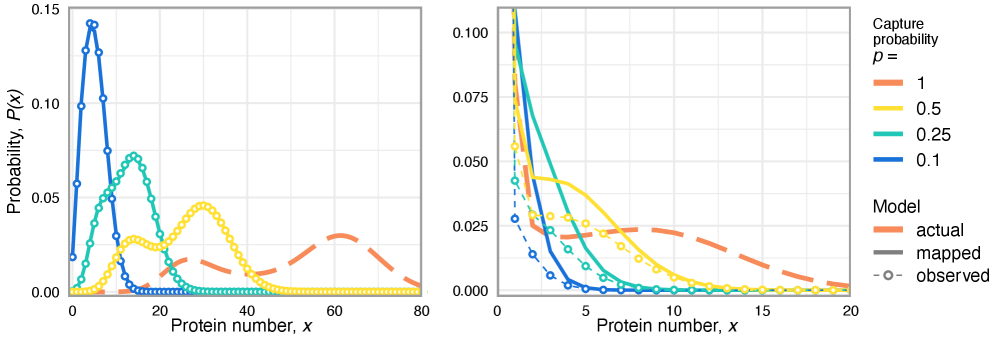
- Technical noise systematically reduces the apparent mean burst size of gene products by a factor of p (the capture probability), e.g., from b(t) to b(t)*p.
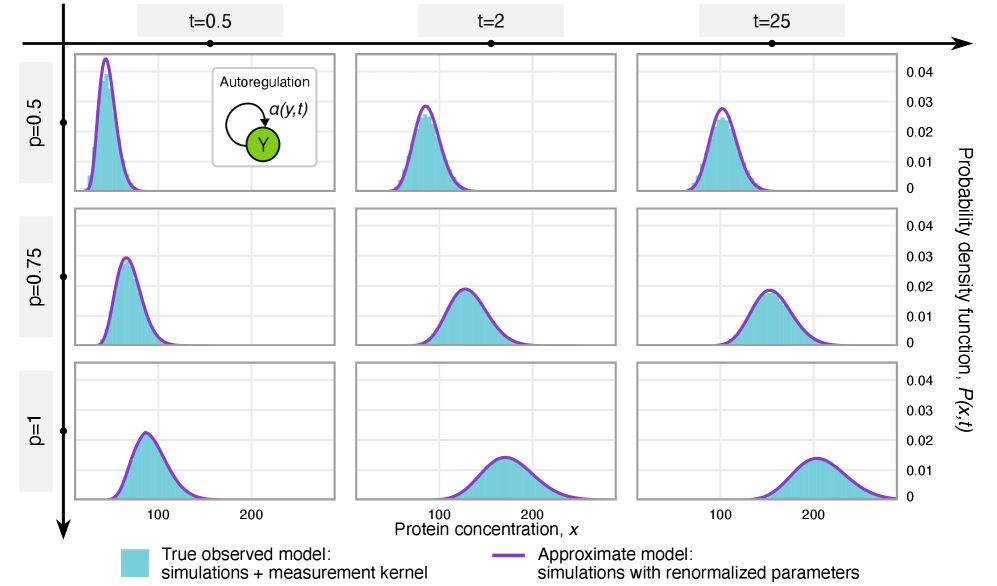
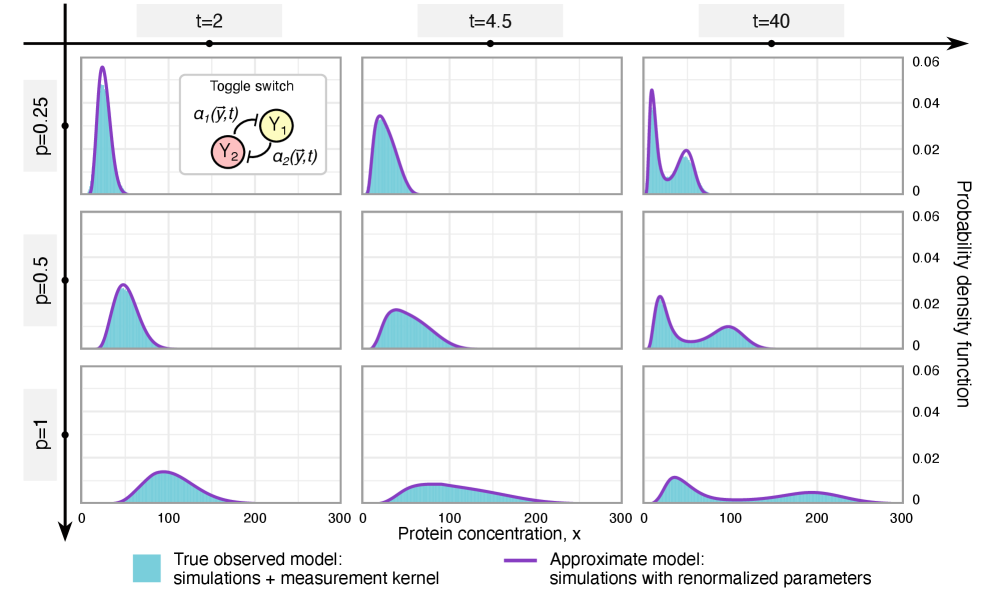
- Rate renormalization occurs when promoter-state transitions are on a distinct timescale (much slower/faster) than other reactions or under high transcription factor abundance.
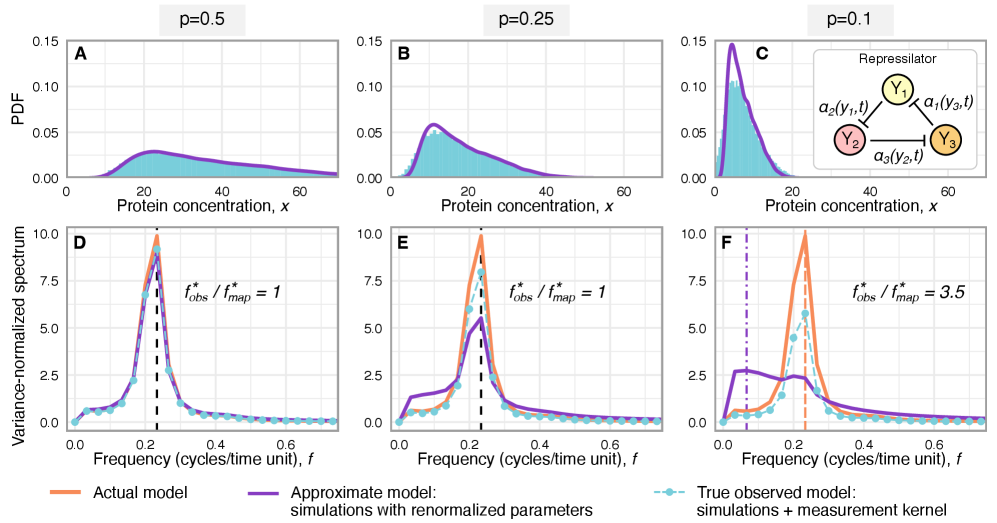
- The framework shows that for the telegraph model, the observed mRNA dynamics are equivalent to the true system with a renormalized transcription rate: k₃(t) → p*k₃(t).
摘要: Imperfect molecular detection in single-cell experiments introduces technical noise that obscures the true stochastic dynamics of gene regulatory networks. While binomial models of molecular capture provide a principled description of imperfect detection, they have so far been analyzed only for simple gene-expression models that do not explicitly account for regulation. Here, we extend binomial models of capture to general gene regulatory networks to understand how imperfect capture reshapes the observed time-dependent statistics of molecular counts. Our results reveal when capture effects correspond to a renormalization of a subset of the kinetic rates and when they cannot be absorbed into effective rates, providing a systematic basis for interpreting noisy single-cell measurements. In particular, we show that rate renormalization emerges either under significant transcription factor abundance or when promoter-state transitions occur on a distinct (much slower or faster) timescale than other reactions. In these cases, technical noise causes the apparent mean burst size of synthesized gene products to appear reduced while transcription factor binding reactions appear faster. These effects hold for gene regulatory networks of arbitrary connectivity and remain valid under time-dependent kinetic rates.