Paper List
-
Nyxus: A Next Generation Image Feature Extraction Library for the Big Data and AI Era
This paper addresses the core pain point of efficiently extracting standardized, comparable features from massive (terabyte to petabyte-scale) biomedi...
-
Topological Enhancement of Protein Kinetic Stability
This work addresses the long-standing puzzle of why knotted proteins exist by demonstrating that deep knots provide a functional advantage through enh...
-
A Multi-Label Temporal Convolutional Framework for Transcription Factor Binding Characterization
This paper addresses the critical limitation of existing TF binding prediction methods that treat transcription factors as independent entities, faili...
-
Social Distancing Equilibria in Games under Conventional SI Dynamics
This paper solves the core problem of proving the existence and uniqueness of Nash equilibria in finite-duration SI epidemic games, showing they are a...
-
Binding Free Energies without Alchemy
This paper addresses the core bottleneck of computational expense in Absolute Binding Free Energy calculations by eliminating the need for numerous al...
-
SHREC: A Spectral Embedding-Based Approach for Ab-Initio Reconstruction of Helical Molecules
This paper addresses the core bottleneck in cryo-EM helical reconstruction: eliminating the dependency on accurate initial symmetry parameter estimati...
-
Budget-Sensitive Discovery Scoring: A Formally Verified Framework for Evaluating AI-Guided Scientific Selection
This paper addresses the critical gap in evaluating AI-guided scientific selection strategies under realistic budget constraints, where existing metri...
-
Probabilistic Joint and Individual Variation Explained (ProJIVE) for Data Integration
This paper addresses the core challenge of accurately decomposing shared (joint) and dataset-specific (individual) sources of variation in multi-modal...
Modulation of DNA rheology by a transcription factor that forms aging microgels
University of Edinburgh | University of Glasgow | MRC Human Genetics Unit | WPI-SKCM2, Hiroshima University
30秒速读
IN SHORT: This work addresses the fundamental question of how the transcription factor NANOG, essential for embryonic stem cell pluripotency, physically regulates gene expression beyond simple DNA binding, by revealing its ability to form self-limiting, aging microgels that modulate DNA rheology.
核心创新
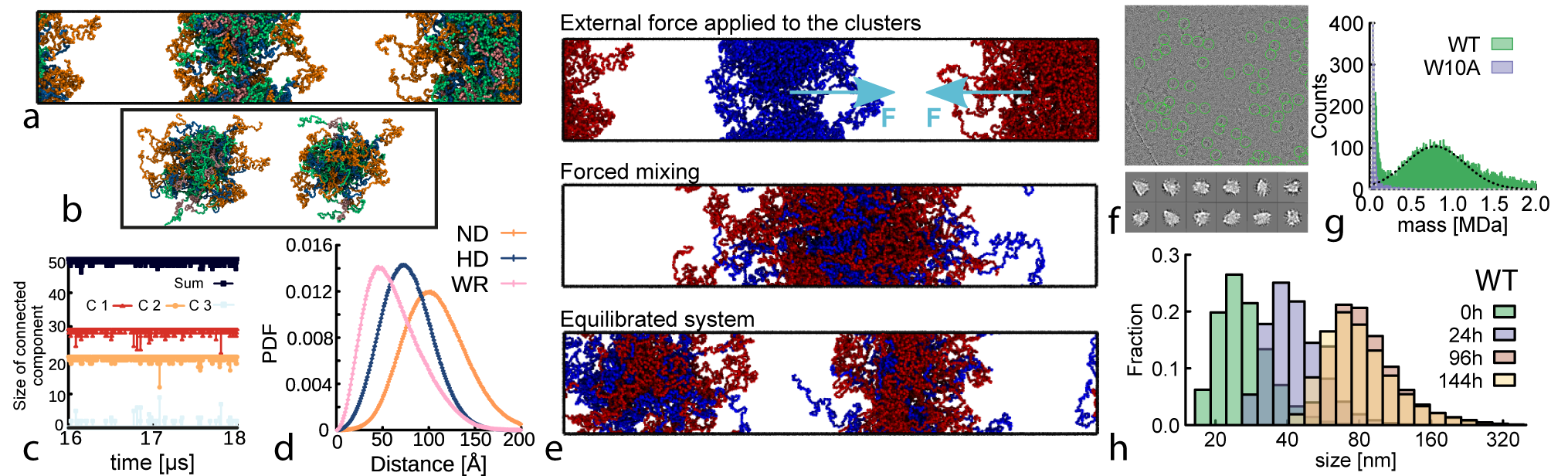
- Methodology First demonstration that a transcription factor (NANOG) forms self-limiting micelle-like clusters (~22-25 monomers) with exposed DNA-binding domains, acting as transient cross-linkers for DNA molecules.
- Biology Discovery of an aging microgel formation by NANOG, where viscoelasticity increases over time (10,000-fold viscosity increase over 12h), driven by its intrinsically disordered tryptophan-rich (WR) domain.
- Theory Proposes a novel 'rheological gene regulation' paradigm: NANOG may regulate gene expression not by large-scale chromatin reorganization, but by stabilizing and restricting the *dynamics* of key regulatory sites via aging condensates, potentially ingraining mechanical memory.
主要结论
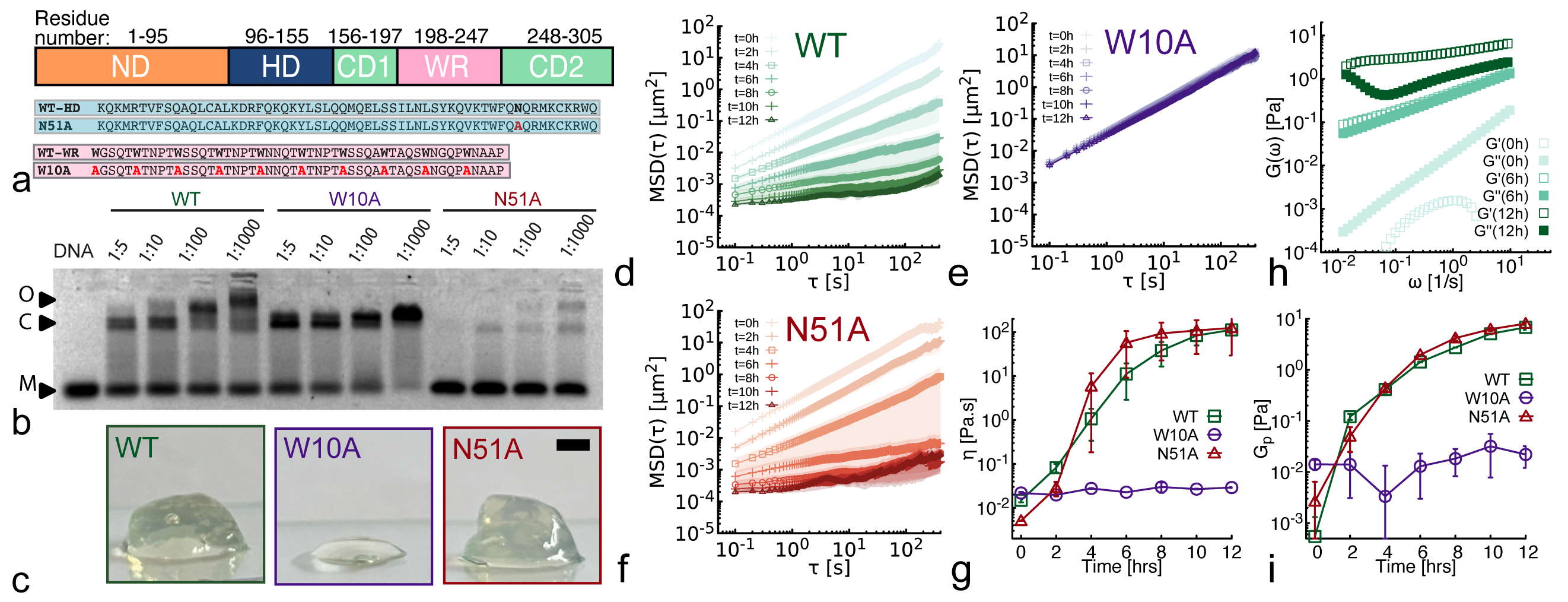
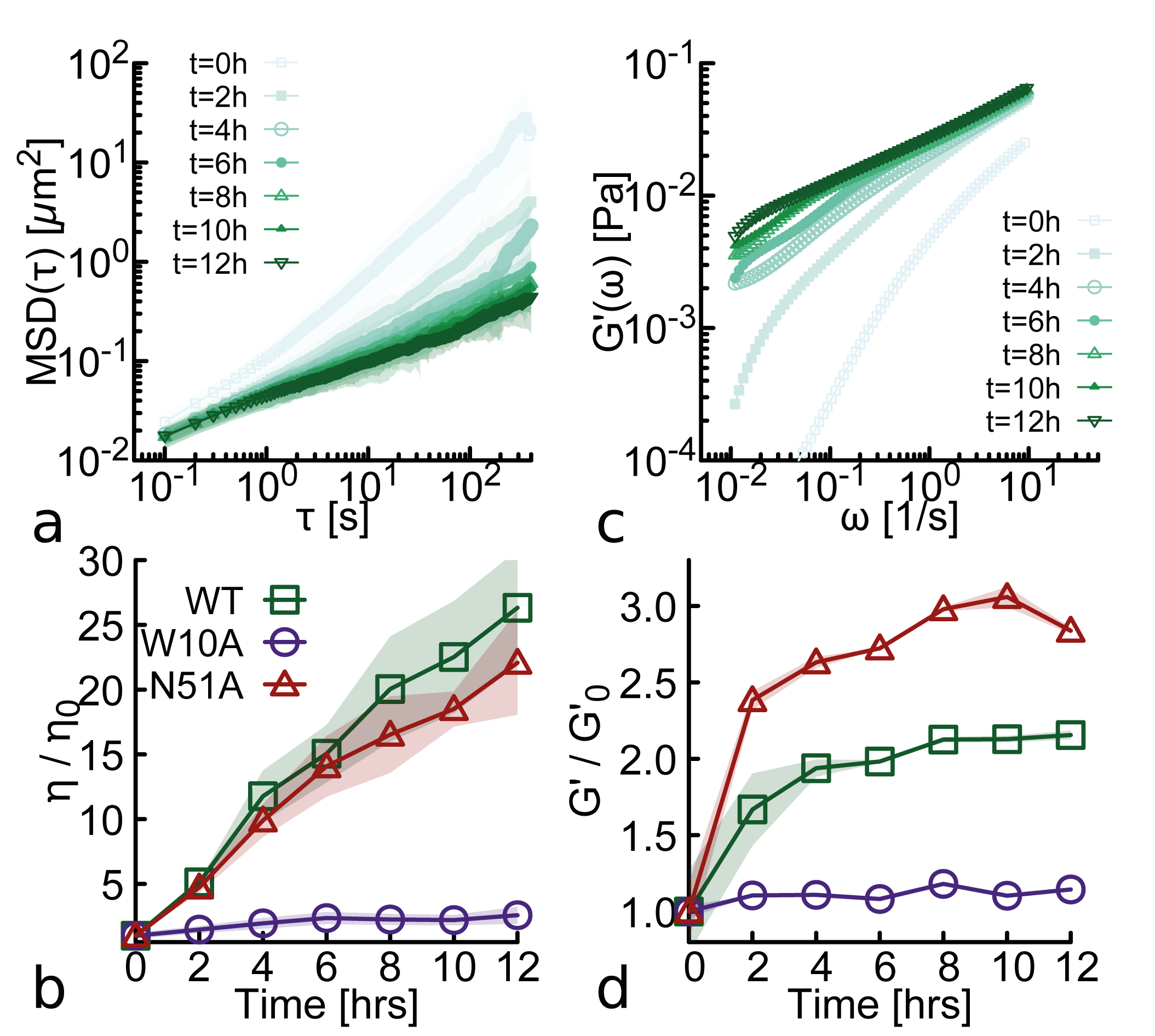
- Wild-type NANOG forms macroscopic aging gels (10,000-fold viscosity increase over 12h at 37°C) and self-limiting micelle-like clusters (~22-25 proteins), while the oligomerization-deficient mutant (W10A) does not.
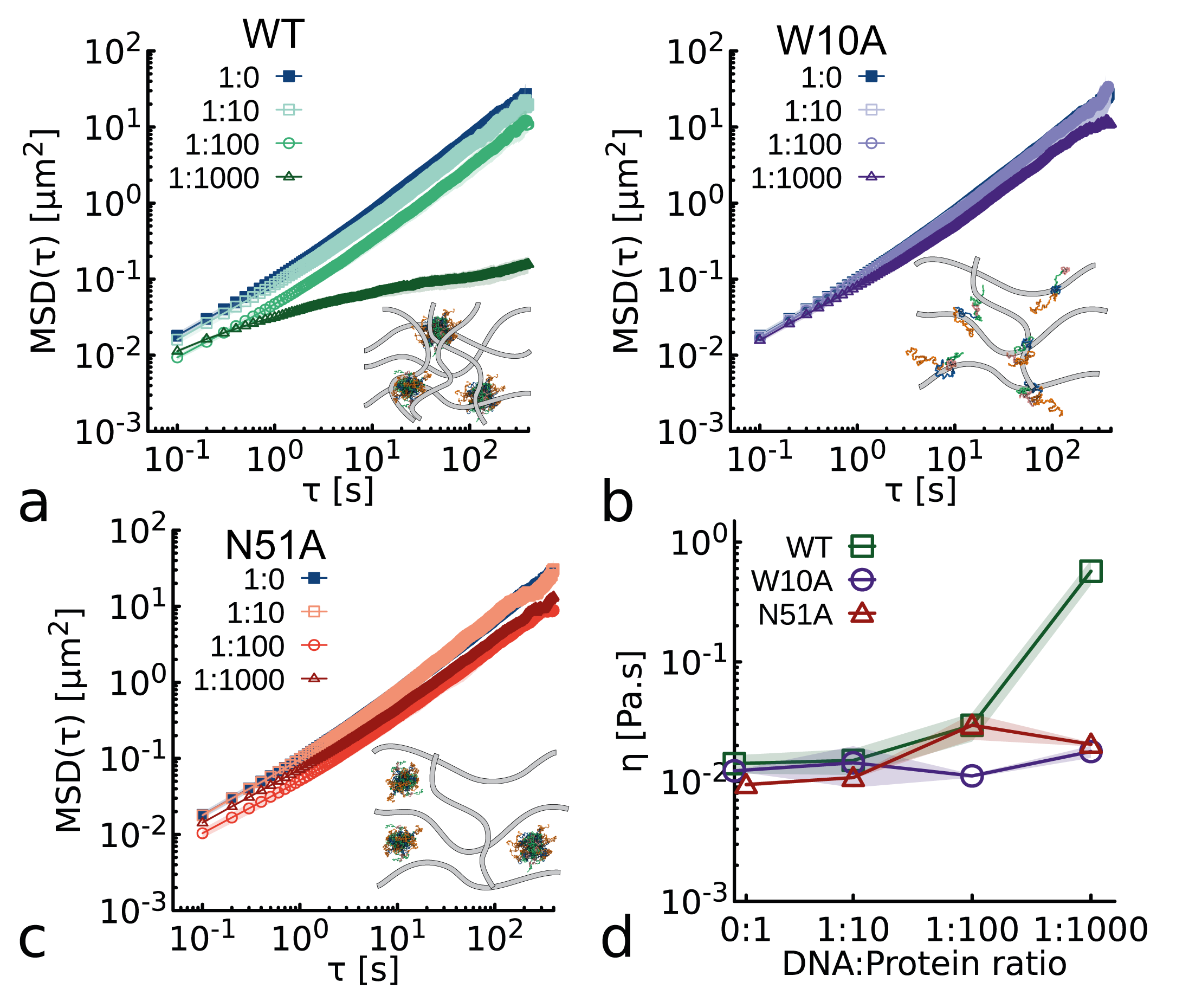
- Both clustering (via WR domain) and DNA binding (via homeodomain) are required for NANOG to act as an effective DNA cross-linker, significantly enhancing the viscoelasticity of entangled DNA solutions (observed in WT but not in W10A or DNA-binding-deficient N51A mutants).
- Aging (increasing viscoelasticity over time) occurs in NANOG-DNA solutions for both WT and the DNA-binding-deficient N51A mutant, indicating that oligomerization alone is sufficient to drive this slow restructuring toward gel-like states.
摘要: Proteins and nucleic acids form non-Newtonian liquids with complex rheological properties that contribute to their function in vivo. Here we investigate the rheology of the transcription factor NANOG, a key protein in sustaining embryonic stem cell self-renewal. We discover that at high concentrations NANOG forms macroscopic aging gels through its intrinsically disordered tryptophan-rich domain. By combining molecular dynamics simulations, mass photometry and Cryo-EM, we also discover that NANOG forms self-limiting micelle-like clusters which expose their DNA-binding domains. In dense solutions of DNA, NANOG micelle-like structures stabilize inter-molecular entanglements and crosslinks, forming microgel-like structures. Our findings suggest that NANOG may contribute to regulate gene expression in a unconventional way: by restricting and stabilizing genome dynamics at key transcriptional sites through the formation of an aging microgel-like structure, potentially enabling mechanical memory in the gene network.