Paper List
-
A Theoretical Framework for the Formation of Large Animal Groups: Topological Coordination, Subgroup Merging, and Velocity Inheritance
This paper addresses the core problem of how large, coordinated animal groups form in nature, challenging the classical view of gradual aggregation by...
-
CONFIDE: Hallucination Assessment for Reliable Biomolecular Structure Prediction and Design
This paper addresses the critical limitation of current protein structure prediction models (like AlphaFold3) where high-confidence scores (pLDDT) can...
-
Generative design and validation of therapeutic peptides for glioblastoma based on a potential target ATP5A
This paper addresses the critical bottleneck in therapeutic peptide design: how to efficiently optimize lead peptides with geometric constraints while...
-
Pharmacophore-based design by learning on voxel grids
This paper addresses the computational bottleneck and limited novelty in conventional pharmacophore-based virtual screening by introducing a voxel cap...
-
Human-Centred Evaluation of Text-to-Image Generation Models for Self-expression of Mental Distress: A Dataset Based on GPT-4o
This paper addresses the critical gap in evaluating how AI-generated images can effectively support cross-cultural mental distress communication, part...
-
ANNE Apnea Paper
This paper addresses the core challenge of achieving accurate, event-level sleep apnea detection and characterization using a non-intrusive, multimoda...
-
DeeDeeExperiment: Building an infrastructure for integrating and managing omics data analysis results in R/Bioconductor
This paper addresses the critical bottleneck of managing and organizing the growing volume of differential expression and functional enrichment analys...
-
Cross-Species Antimicrobial Resistance Prediction from Genomic Foundation Models
This paper addresses the core challenge of predicting antimicrobial resistance across phylogenetically distinct bacterial species, where traditional m...
Assessment of Simulation-based Inference Methods for Stochastic Compartmental Models
Bonn Center for Mathematical Life Sciences, University of Bonn | Life and Medical Science Institute, University of Bonn | Institute of Software Technology, German Aerospace Center (DLR)
30秒速读
IN SHORT: This paper addresses the core challenge of performing accurate Bayesian parameter inference for stochastic epidemic models when the likelihood function is intractable, a common bottleneck for real-time forecasting.
核心创新
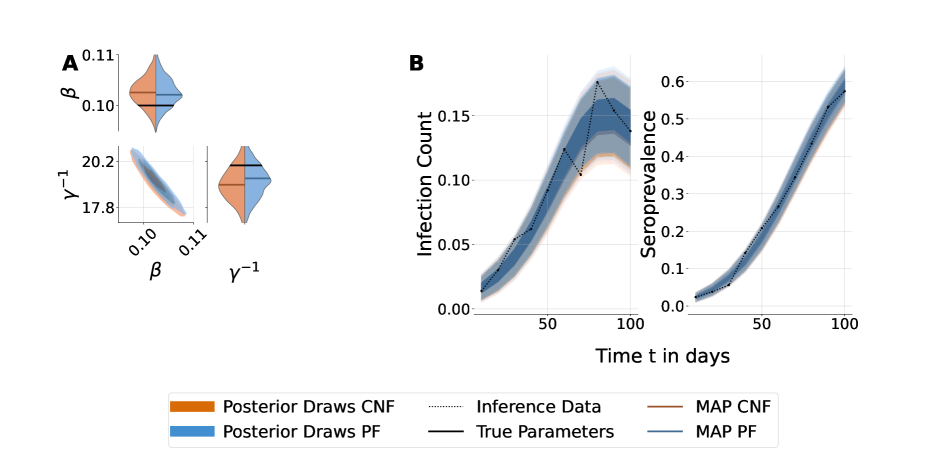
- Methodology Provides the first comprehensive, praxis-driven comparison between Particle Filters (PF) and Conditional Normalizing Flows (CNF) for inference on stochastic compartmental models, benchmarking their performance head-to-head.
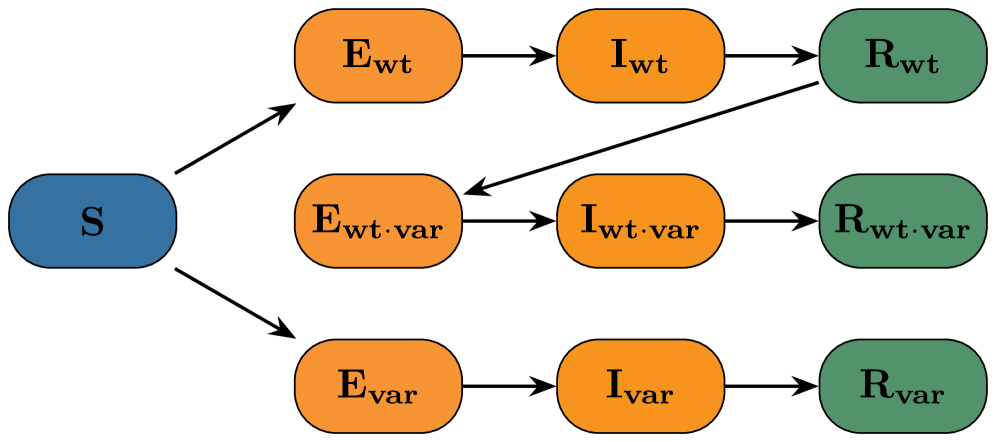
- Methodology Demonstrates the application and robustness of these likelihood-free methods on a complex, non-identifiable two-variant SEIR model with real-world data from an Ethiopian COVID-19 cohort, including scenarios with irregular sampling and missing data.
- Theory Shows that parameter space reparameterization (e.g., using R0, e0, s0) can mitigate ill-conditioning in complex models, improving posterior alignment between PF and CNF methods.
主要结论
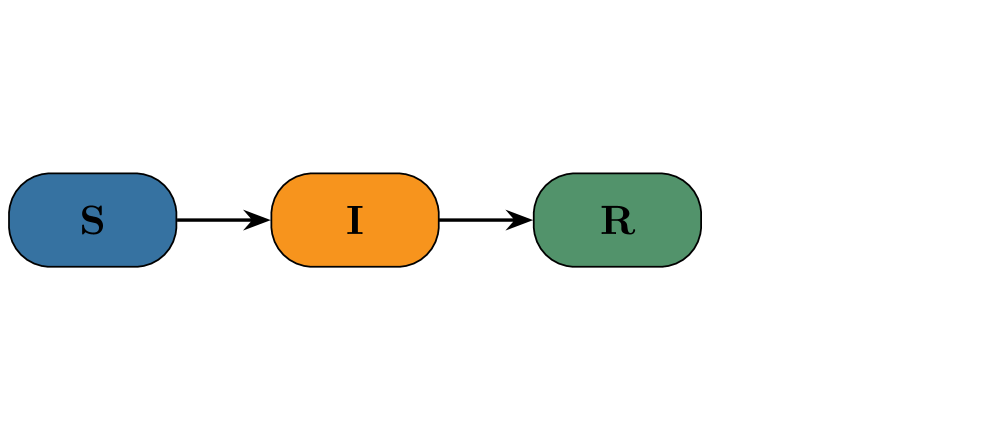
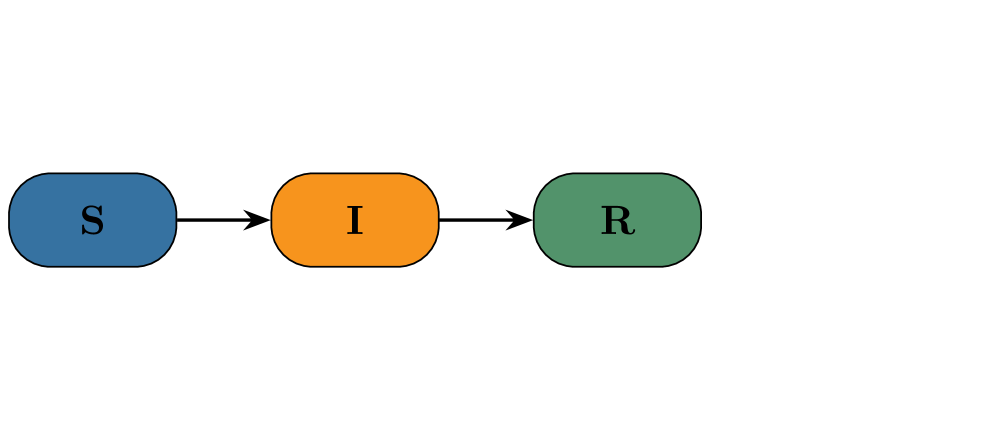
- Both PF and CNF provided robust and reliable inference on the stochastic SIR model with synthetic data, validating the implementation framework.
- For the complex two-variant SEIR model, both methods yielded good fits to synthetic data, but ill-conditioning led to differences in marginal posterior shapes; reparameterization with dimension reduction improved posterior alignment.
- Application to real Ethiopian cohort data demonstrated the operational robustness of both PF and CNF under conditions of real-world noise and irregular data sampling, proving their practical utility.
摘要: Global pandemics, such as the recent COVID-19 crisis, highlight the need for stochastic epidemic models that can capture the randomness inherent in the spread of disease. Such models must be accompanied by methods for estimating parameters in order to generate fast nowcasts and short-term forecasts that can inform public health decisions. This paper presents a comparison of two advanced Bayesian inference methods: 1) pseudo-marginal particle Markov chain Monte Carlo, short Particle Filters (PF), and 2) Conditional Normalizing Flows (CNF). We investigate their performance on two commonly used compartmental models: a classical Susceptible-Infected-Recovered (SIR) model and a two-variant Susceptible-Exposed-Infected-Recovered (SEIR) model, complemented by an observation model that maps latent trajectories to empirical data. Addressing the challenges of intractable likelihoods for parameter inference in stochastic settings, our analysis highlights how these likelihood-free methods provide accurate and robust inference capabilities. The results of our simulation study further underscore the effectiveness of these approaches in capturing the stochastic dynamics of epidemics, providing prediction capabilities for the control of epidemic outbreaks. Results on an Ethiopian cohort study demonstrate operational robustness under real‑world noise and irregular data sampling. To facilitate reuse and to enable building pipelines that ultimately contribute to better informed decision making in public health, we make code and synthetic datasets publicly available.