Paper List
-
A Theoretical Framework for the Formation of Large Animal Groups: Topological Coordination, Subgroup Merging, and Velocity Inheritance
This paper addresses the core problem of how large, coordinated animal groups form in nature, challenging the classical view of gradual aggregation by...
-
CONFIDE: Hallucination Assessment for Reliable Biomolecular Structure Prediction and Design
This paper addresses the critical limitation of current protein structure prediction models (like AlphaFold3) where high-confidence scores (pLDDT) can...
-
Generative design and validation of therapeutic peptides for glioblastoma based on a potential target ATP5A
This paper addresses the critical bottleneck in therapeutic peptide design: how to efficiently optimize lead peptides with geometric constraints while...
-
Pharmacophore-based design by learning on voxel grids
This paper addresses the computational bottleneck and limited novelty in conventional pharmacophore-based virtual screening by introducing a voxel cap...
-
Human-Centred Evaluation of Text-to-Image Generation Models for Self-expression of Mental Distress: A Dataset Based on GPT-4o
This paper addresses the critical gap in evaluating how AI-generated images can effectively support cross-cultural mental distress communication, part...
-
ANNE Apnea Paper
This paper addresses the core challenge of achieving accurate, event-level sleep apnea detection and characterization using a non-intrusive, multimoda...
-
DeeDeeExperiment: Building an infrastructure for integrating and managing omics data analysis results in R/Bioconductor
This paper addresses the critical bottleneck of managing and organizing the growing volume of differential expression and functional enrichment analys...
-
Cross-Species Antimicrobial Resistance Prediction from Genomic Foundation Models
This paper addresses the core challenge of predicting antimicrobial resistance across phylogenetically distinct bacterial species, where traditional m...
DeepFRI Demystified: Interpretability vs. Accuracy in AI Protein Function Prediction
Yale University | Microsoft
30秒速读
IN SHORT: This study addresses the critical gap between high predictive accuracy and biological interpretability in DeepFRI, revealing that the model often prioritizes structural motifs over functional residues, complicating reliable identification of drug targets.
核心创新
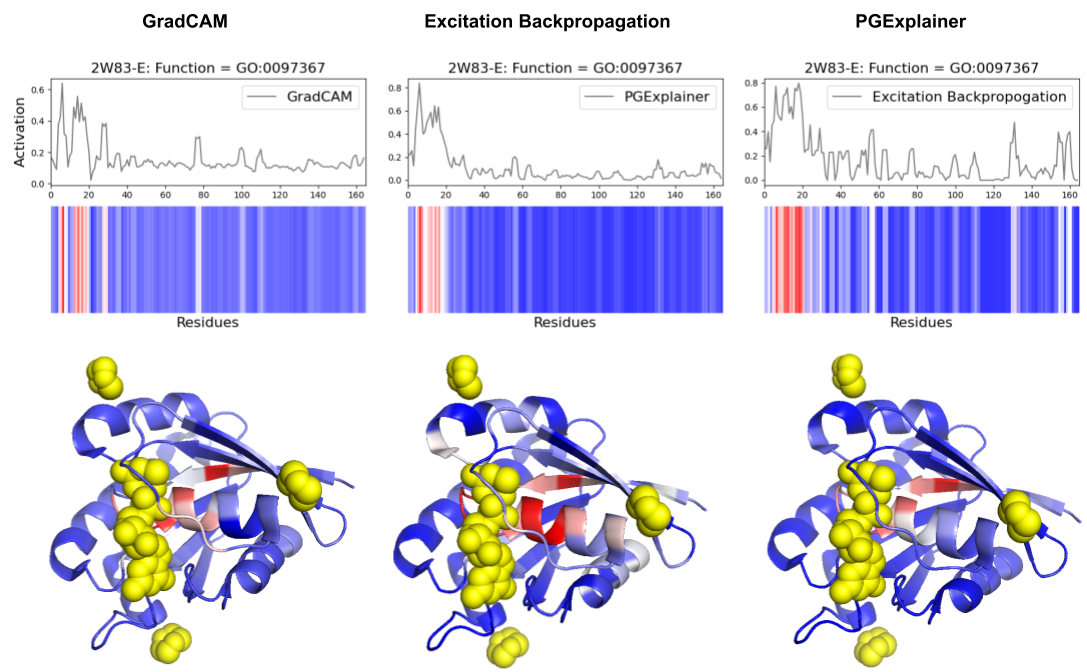
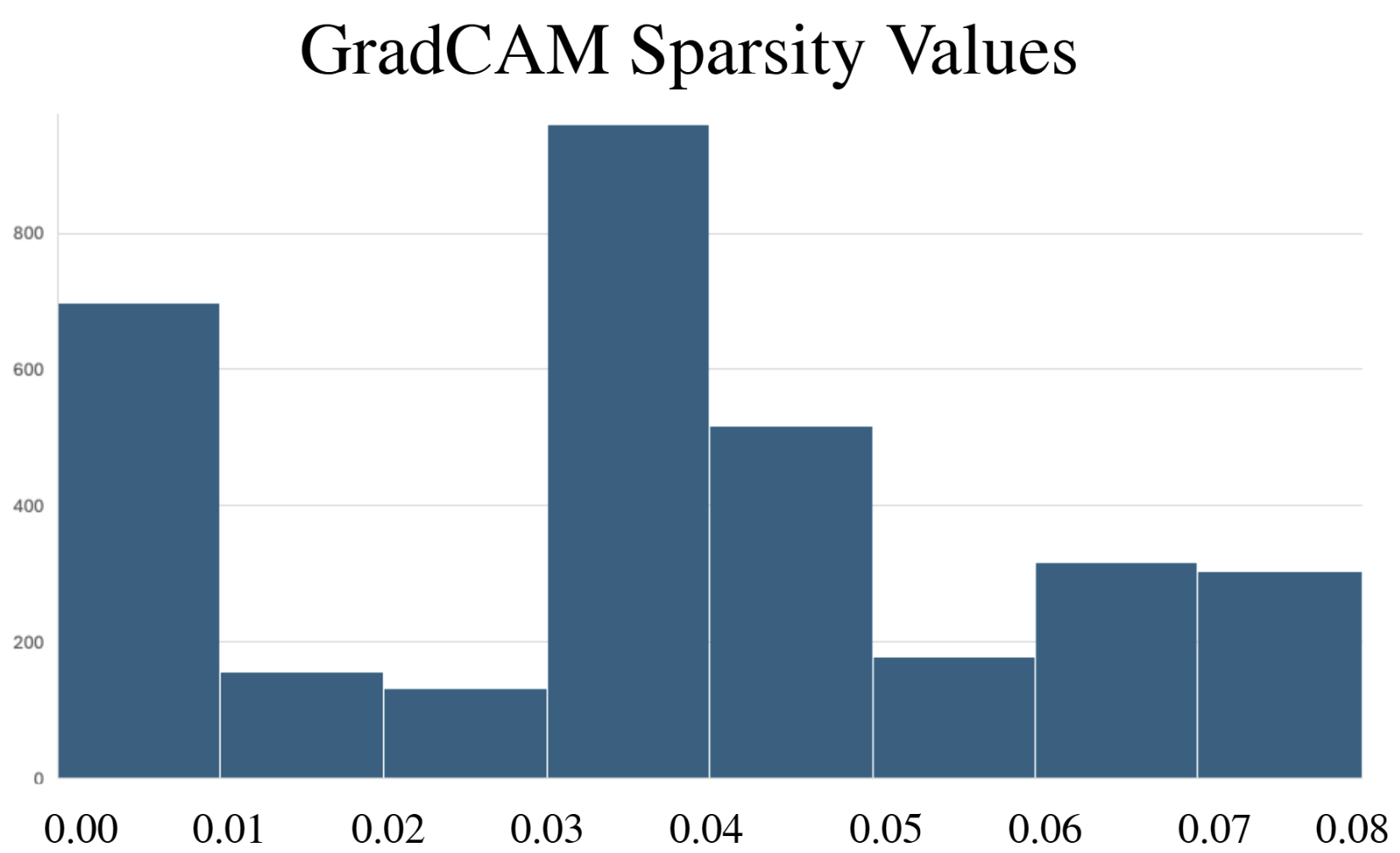
- Methodology Comprehensive benchmarking of three post-hoc explainability methods (GradCAM, Excitation Backpropagation, PGExplainer) on DeepFRI with quantitative sparsity analysis.
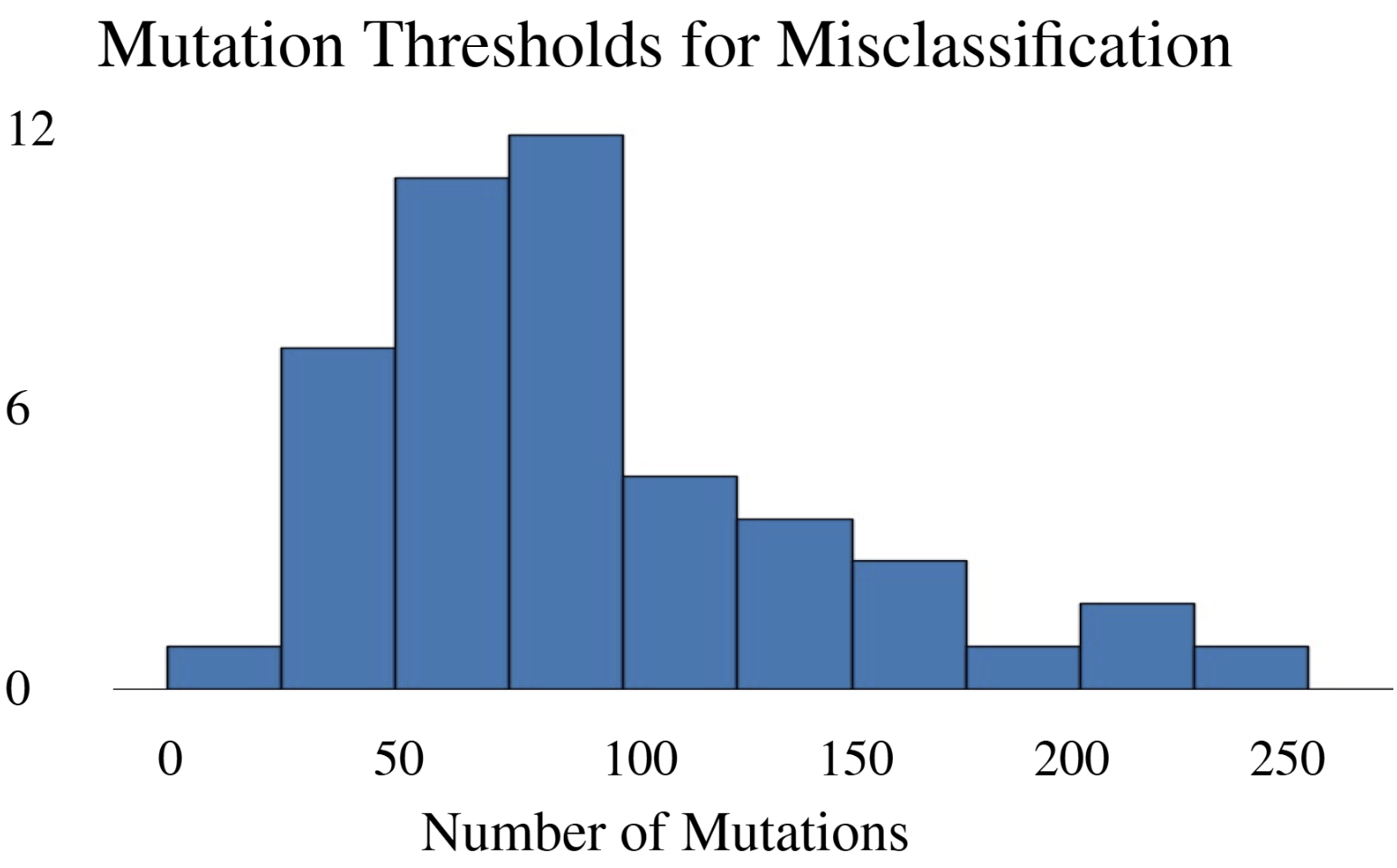
- Methodology Development of a modified DeepFool adversarial testing framework for protein sequences, measuring mutation thresholds required for misclassification.
- Biology Revealed that DeepFRI prioritizes amino acids controlling protein structure over function in >50% of tested proteins, highlighting a fundamental accuracy-interpretability trade-off.
主要结论
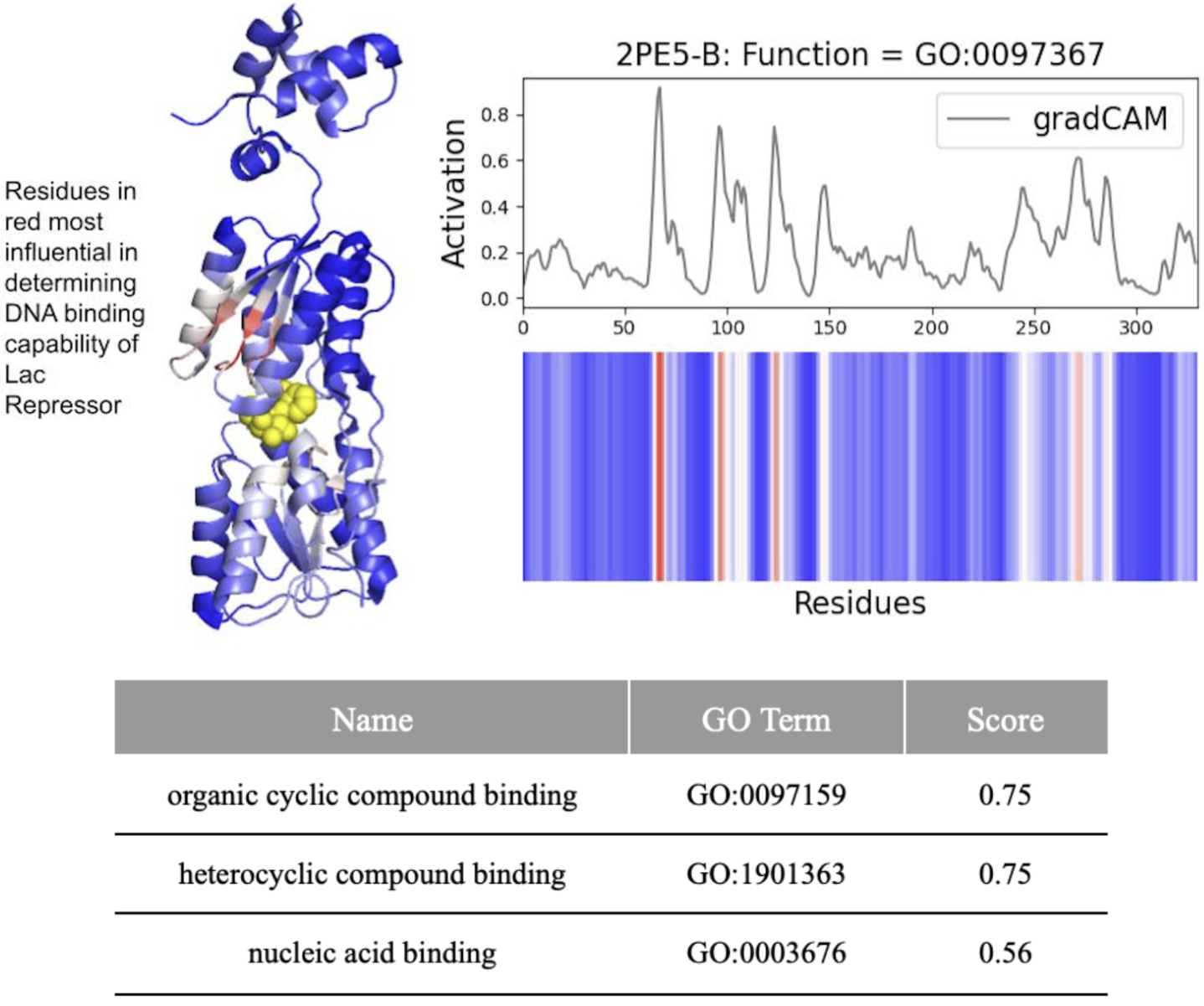
- DeepFRI required 206 mutations (62.4% of 330 residues) in the lac repressor for misclassification, demonstrating extreme robustness but potentially missing subtle functional alterations.
- Explainability methods showed significant granularity differences: PGExplainer was 3× sparser than GradCAM and 17× sparser than Excitation Backpropagation across 124 binding proteins.
- All three methods converged on biochemically critical P-loop residues (0-20) in ARF6 GTPase, validating DeepFRI's focus on conserved functional motifs in straightforward domains.
摘要: Machine learning technologies for protein function prediction are black box models. Despite their potential to identify key drug targets with high accuracy and accelerate therapy development, the adoption of these methods depends on verifying their findings. This study evaluates DeepFRI, a leading Graph Convolutional Network (GCN)-based tool, using advanced explainability techniques—GradCAM, Excitation Backpropagation, and PGExplainer—and adversarial robustness tests. Our findings reveal that the model’s predictions often prioritize conserved motifs over truly deterministic residues, complicating the identification of functional sites. Quantitative analyses show that explainability methods differ significantly in granularity, with GradCAM providing broad relevance and PGExplainer pinpointing specific active sites. These results highlight trade-offs between accuracy and interpretability, suggesting areas for improvement in DeepFRI’s architecture to enhance its trustworthiness in drug discovery and regulatory settings.