Paper List
-
Developing the PsyCogMetrics™ AI Lab to Evaluate Large Language Models and Advance Cognitive Science
This paper addresses the critical gap between sophisticated LLM evaluation needs and the lack of accessible, scientifically rigorous platforms that in...
-
Equivalence of approximation by networks of single- and multi-spike neurons
This paper resolves the fundamental question of whether single-spike spiking neural networks (SNNs) are inherently less expressive than multi-spike SN...
-
The neuroscience of transformers
提出了Transformer架构与皮层柱微环路之间的新颖计算映射,连接了现代AI与神经科学。
-
Framing local structural identifiability and observability in terms of parameter-state symmetries
This paper addresses the core challenge of systematically determining which parameters and states in a mechanistic ODE model can be uniquely inferred ...
-
Leveraging Phytolith Research using Artificial Intelligence
This paper addresses the critical bottleneck in phytolith research by automating the labor-intensive manual microscopy process through a multimodal AI...
-
Neural network-based encoding in free-viewing fMRI with gaze-aware models
This paper addresses the core challenge of building computationally efficient and ecologically valid brain encoding models for naturalistic vision by ...
-
Scalable DNA Ternary Full Adder Enabled by a Competitive Blocking Circuit
This paper addresses the core bottleneck of carry information attenuation and limited computational scale in DNA binary adders by introducing a scalab...
-
ELISA: An Interpretable Hybrid Generative AI Agent for Expression-Grounded Discovery in Single-Cell Genomics
This paper addresses the critical bottleneck of translating high-dimensional single-cell transcriptomic data into interpretable biological hypotheses ...
Enhancing Clinical Note Generation with ICD-10, Clinical Ontology Knowledge Graphs, and Chain-of-Thought Prompting Using GPT-4
Computer Science, Old Dominion University | Biomedical Informatics, University of Arkansas for Medical Sciences
30秒速读
IN SHORT: This paper addresses the core challenge of generating accurate and clinically relevant patient notes from sparse inputs (ICD codes and basic demographics) by augmenting Chain-of-Thought prompting with semantic search and structured medical knowledge graphs.
核心创新
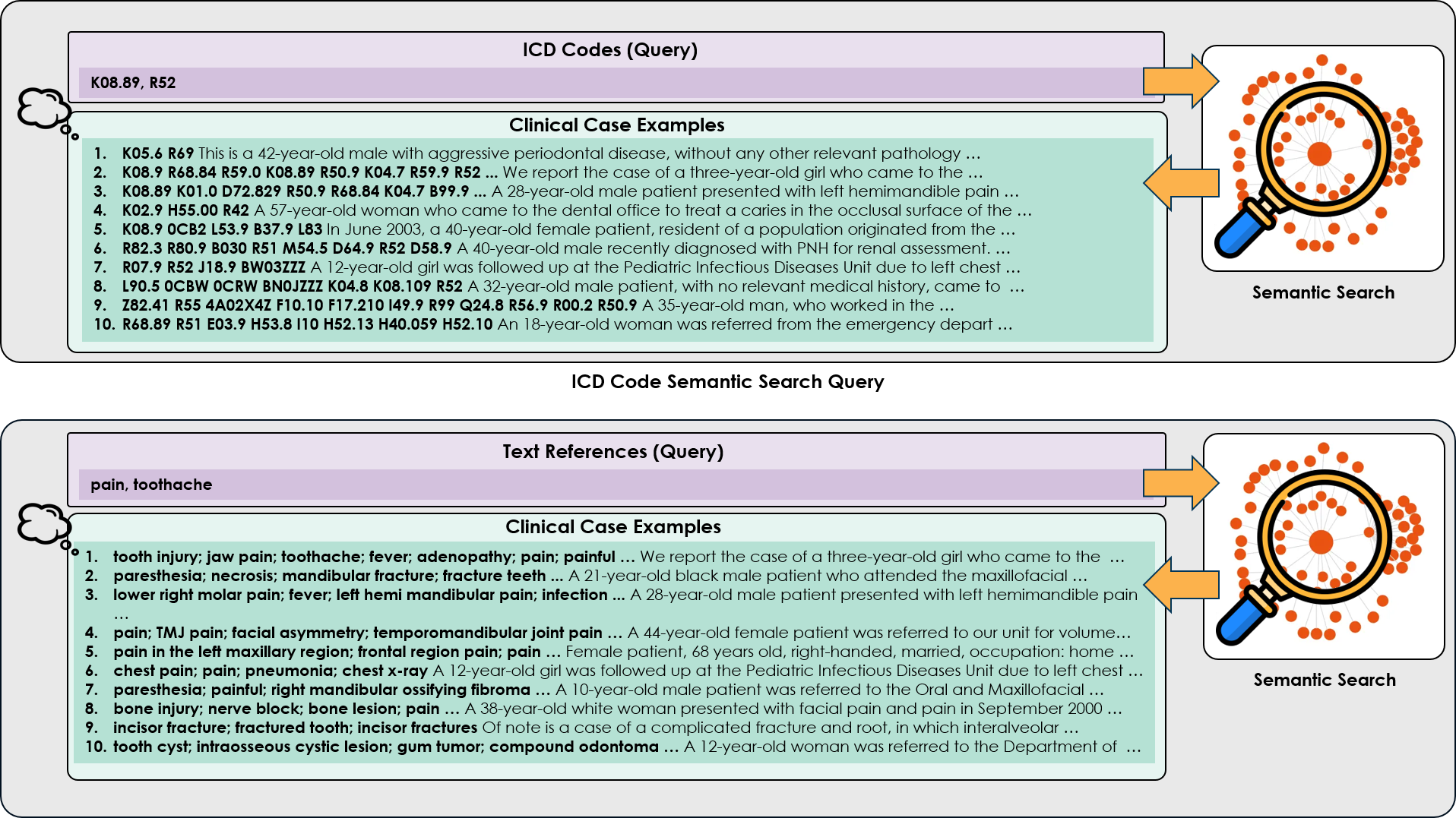
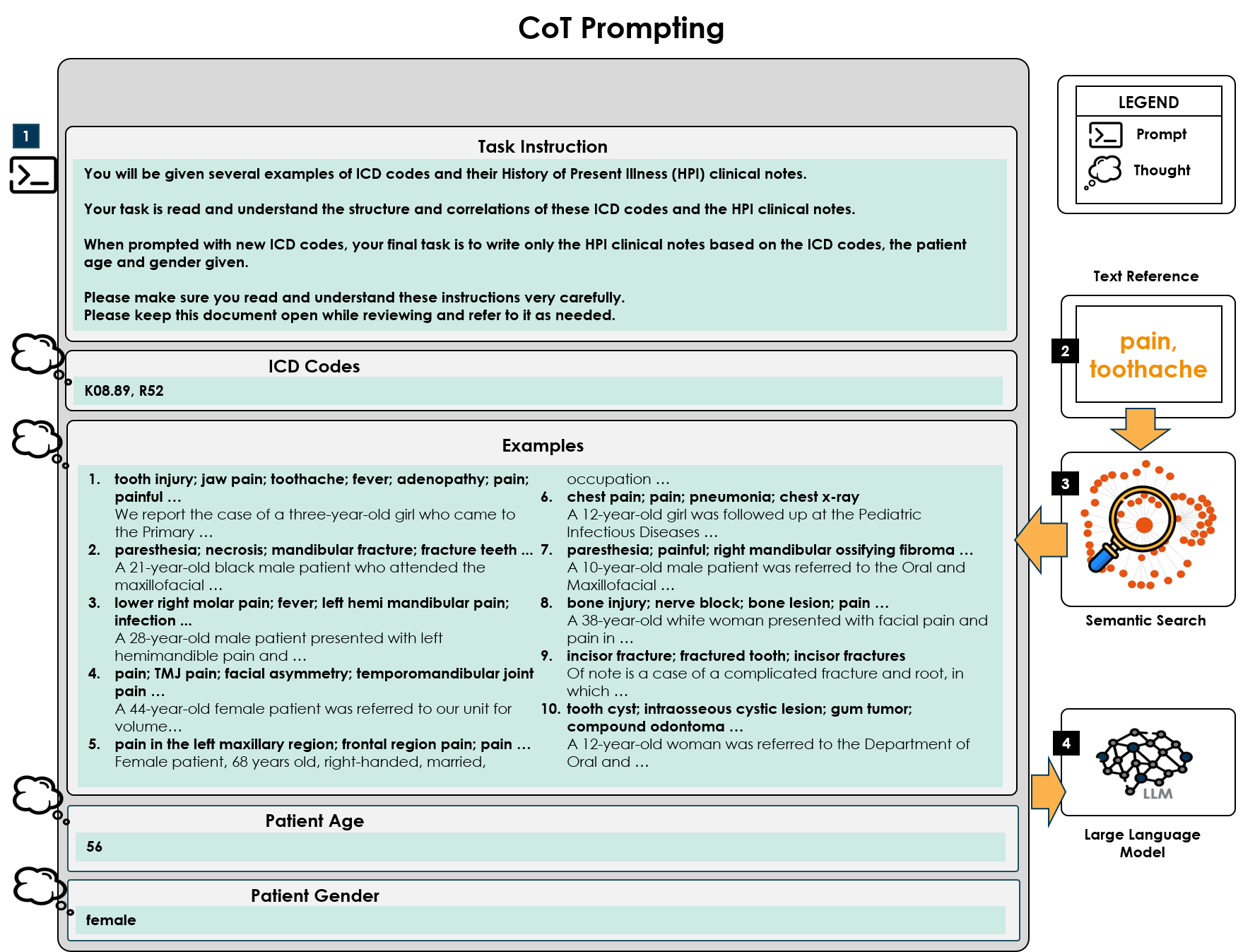
- Methodology Proposes a novel hybrid prompting framework that integrates traditional Chain-of-Thought reasoning with semantic search results from a clinical corpus (CodiEsp dataset) to provide contextual examples.
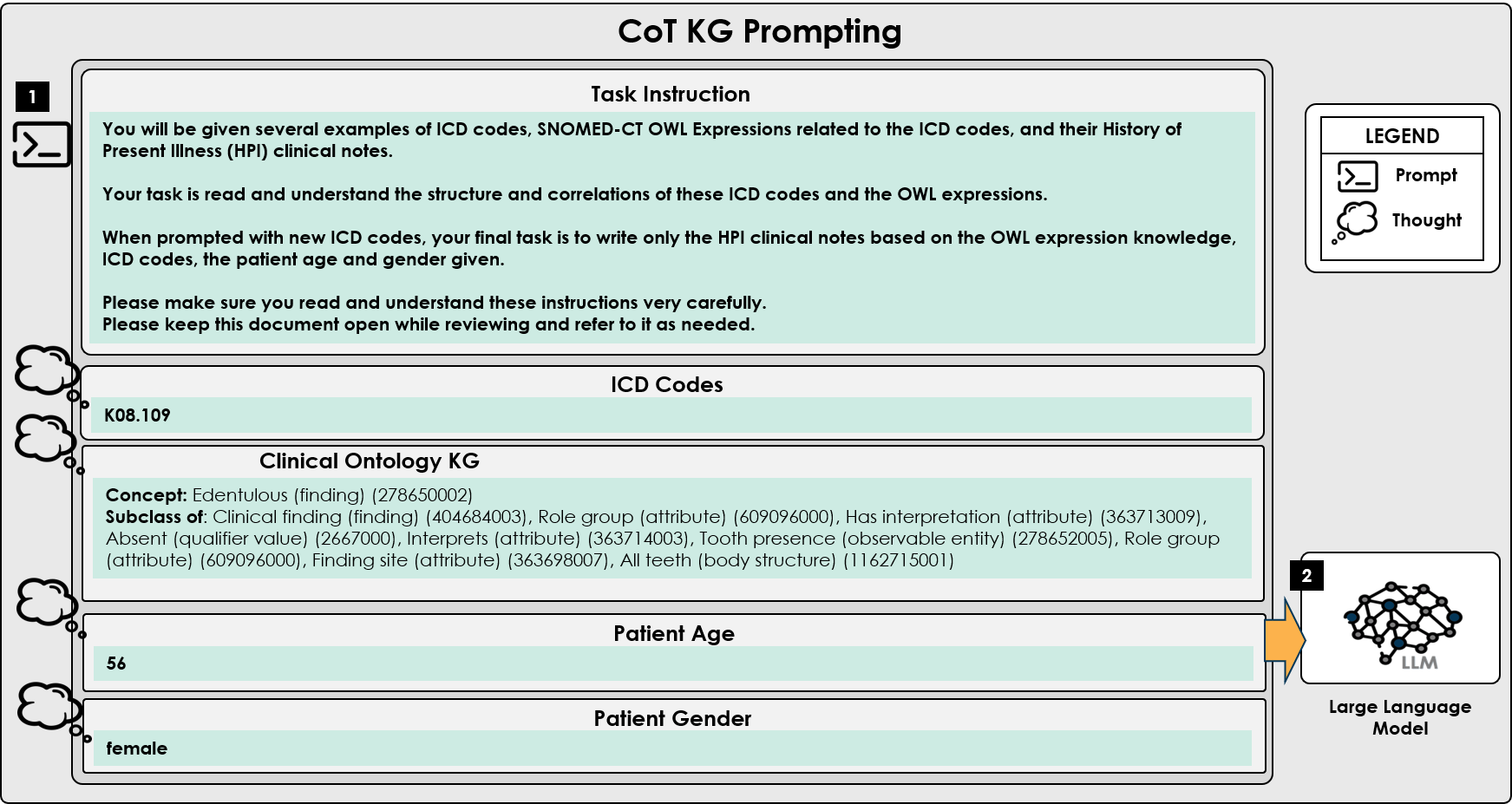
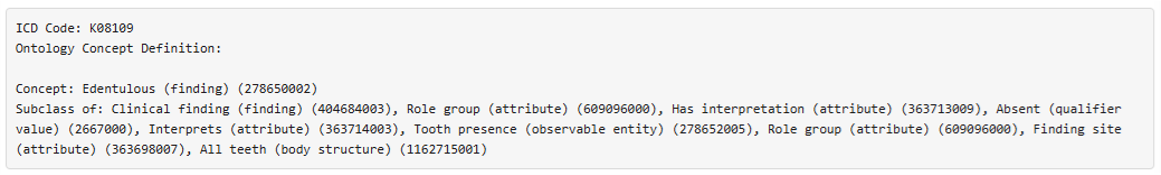
- Methodology Introduces the infusion of a structured clinical ontology knowledge graph (built from SNOMED CT OWL expressions) directly into the LLM prompt to ground generation in formal medical relationships and constraints.
- Methodology/Biology Demonstrates the first systematic approach to reverse the common ICD code classification task, instead generating comprehensive clinical notes from ICD codes as primary input, evaluated on six distinct clinical cases.
主要结论
- The proposed CoT prompting with semantic search (using ICD codes as query) consistently outperformed the standard one-shot baseline across six clinical cases, as evidenced by lower cosine distance scores (e.g., Case C showed a clear leftward shift in KDE peak, indicating higher semantic similarity to ground truth).
- Incorporating a clinical knowledge graph (SNOMED CT OWL) into the prompt (CoT KG) provided structured medical relationships, enriching the generated notes with domain-specific terminology and logical constraints derived from formal ontologies.
- The hybrid approach (CoT Semantic Search + KG) leverages both in-context examples from similar cases and formal medical knowledge, offering a robust framework for improving the factual accuracy and clinical relevance of LLM-generated notes from coded inputs.
摘要: In the past decade a surge in the amount of electronic health record (EHR) data in the United States, attributed to a favorable policy environment created by the Health Information Technology for Economic and Clinical Health (HITECH) Act of 2009 and the 21st Century Cures Act of 2016. Clinical notes for patients’ assessments, diagnoses, and treatments are captured in these EHRs in free-form text by physicians, who spend a considerable amount of time entering and editing them. Manually writing clinical notes takes a considerable amount of a doctor’s valuable time, increasing the patient’s waiting time and possibly delaying diagnoses. Large language models (LLMs) possess the ability to generate news articles that closely resemble human-written ones. We investigate the usage of Chain-of-Thought (CoT) prompt engineering to improve the LLM’s response in clinical note generation. In our prompts, we use as input International Classification of Diseases (ICD) codes and basic patient information. We investigate a strategy that combines the traditional CoT with semantic search results to improve the quality of generated clinical notes. Additionally, we infuse a knowledge graph (KG) built from clinical ontology to further enrich the domain-specific knowledge of generated clinical notes. We test our prompting technique on six clinical cases from the CodiEsp test dataset using GPT-4 and our results show that it outperformed the clinical notes generated by standard one-shot prompts.