Paper List
-
An AI Implementation Science Study to Improve Trustworthy Data in a Large Healthcare System
This paper addresses the critical gap between theoretical AI research and real-world clinical implementation by providing a practical framework for as...
-
The BEAT-CF Causal Model: A model for guiding the design of trials and observational analyses of cystic fibrosis exacerbations
This paper addresses the critical gap in cystic fibrosis exacerbation management by providing a formal causal framework that integrates expert knowled...
-
Hierarchical Molecular Language Models (HMLMs)
This paper addresses the core challenge of accurately modeling context-dependent signaling, pathway cross-talk, and temporal dynamics across multiple ...
-
Stability analysis of action potential generation using Markov models of voltage‑gated sodium channel isoforms
This work addresses the challenge of systematically characterizing how the high-dimensional parameter space of Markov models for different sodium chan...
-
Approximate Bayesian Inference on Mechanisms of Network Growth and Evolution
This paper addresses the core challenge of inferring the relative contributions of multiple, simultaneous generative mechanisms in network formation w...
-
EnzyCLIP: A Cross-Attention Dual Encoder Framework with Contrastive Learning for Predicting Enzyme Kinetic Constants
This paper addresses the core challenge of jointly predicting enzyme kinetic parameters (Kcat and Km) by modeling dynamic enzyme-substrate interaction...
-
Tissue stress measurements with Bayesian Inversion Stress Microscopy
This paper addresses the core challenge of measuring absolute, tissue-scale mechanical stress without making assumptions about tissue rheology, which ...
-
DeepFRI Demystified: Interpretability vs. Accuracy in AI Protein Function Prediction
This study addresses the critical gap between high predictive accuracy and biological interpretability in DeepFRI, revealing that the model often prio...
Incorporating indel channels into average-case analysis of seed-chain-extend
Carnegie Mellon University, Pittsburgh, PA, USA
30秒速读
IN SHORT: This paper addresses the core pain point of bridging the theoretical gap for the widely used seed-chain-extend heuristic by providing the first rigorous average-case analysis that accounts for insertions and deletions (indels), not just substitutions.
核心创新
- Methodology Introduces a generalized definition of 'recoverability' and a 'homologous path' to mathematically model the correct alignment under indel mutation channels, moving beyond the simpler 'homologous diagonal' used for substitutions only.
- Theory Develops new mathematical machinery to handle the dependence structure of neighboring anchors and the existence of 'clipping anchors' (partially correct anchors), which are unique challenges introduced by indels.
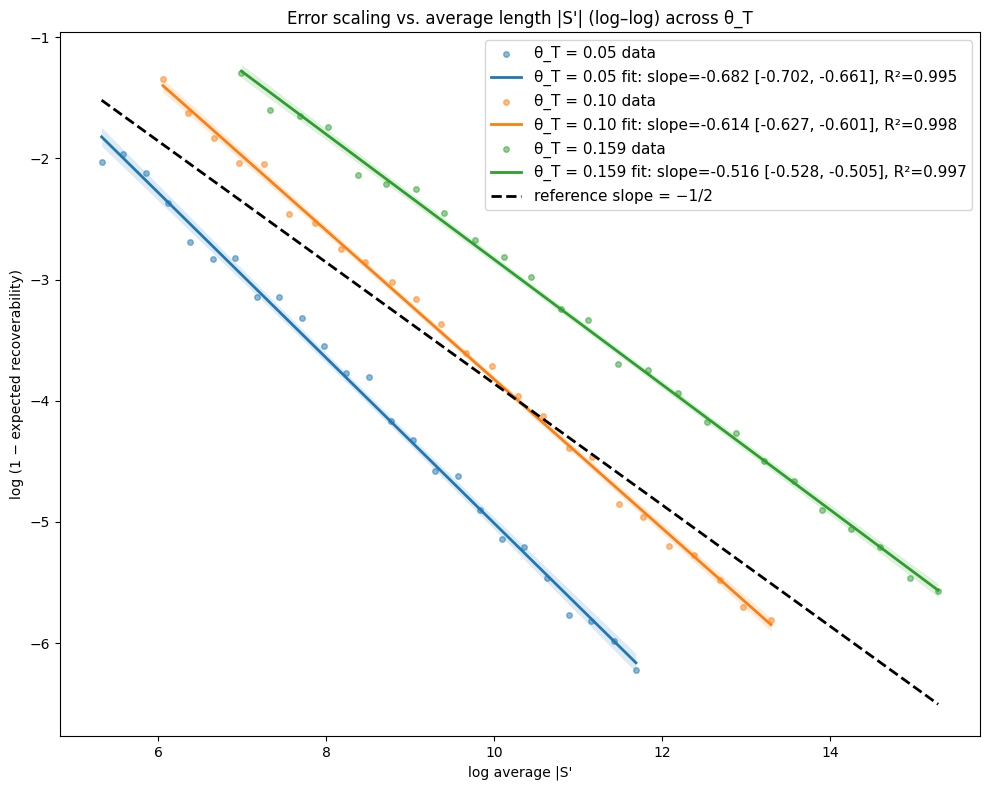
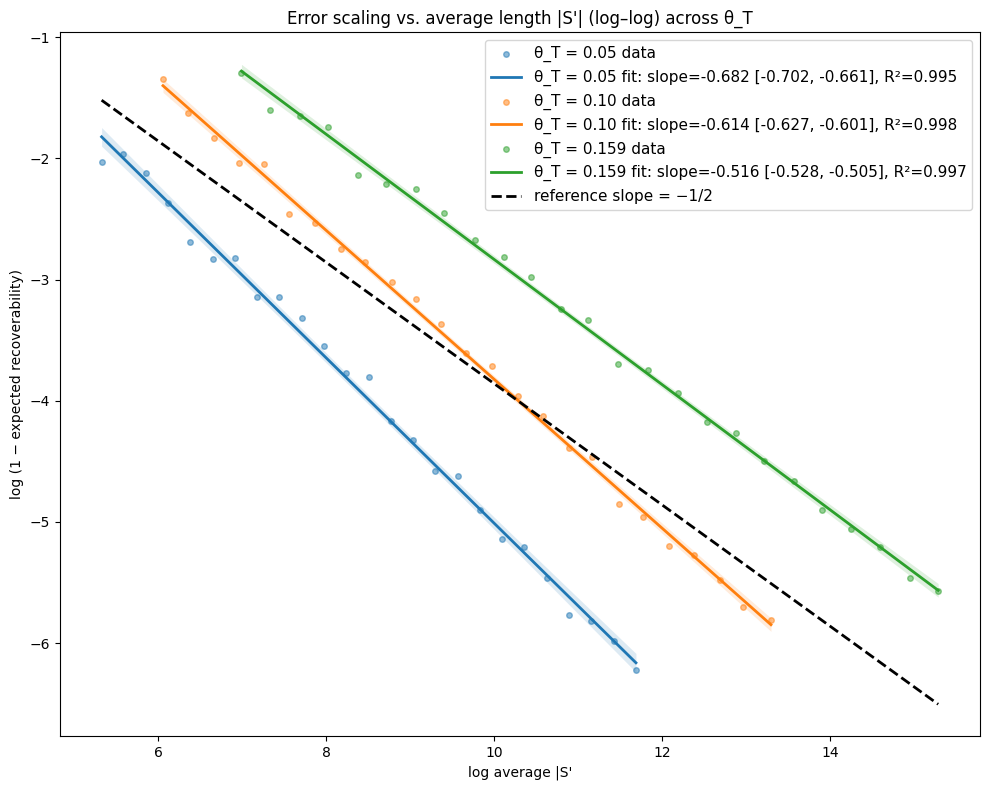
- Theory Proves that under a total mutation rate θ_T < 0.159, optimal linear-gap cost chaining achieves an expected recoverability of ≥ 1 - O(1/√m), generalizing the prior substitution-only result to a biologically realistic model.
主要结论
- The expected recoverability of an optimal chain under linear-gap cost chaining is ≥ 1 - O(1/√m) when the total mutation rate θ_T (sum of substitution, insertion, deletion rates) is less than 0.159.
- The expected runtime of the algorithm is O(m n^(3.15·θ_T) log n). For example, at a θ_T of 0.05 (similar to human-chimp divergence), the exponent is ~1.12, leading to near-linear scaling.
- The analysis successfully bridges theory and practice by extending the proof framework to handle indels, justifying the heuristic's empirical effectiveness on real genomic data which contains indels.
摘要: Given a sequence s1 of n letters drawn i.i.d. from an alphabet of size σ and a mutated substring s2 of length m<n, we often want to recover the mutation history that generated s2 from s1. Modern sequence aligners are widely used for this task, and many employ the seed-chain-extend heuristic with k-mer seeds. Previously, Shaw and Yu showed that optimal linear-gap cost chaining can produce a chain with 1−O(1/m) recoverability, the proportion of the mutation history that is recovered, in O(mn^(2.43θ) log n) expected time, where θ<0.206 is the mutation rate under a substitution-only channel and s1 is assumed to be uniformly random. However, a gap remains between theory and practice, since real genomic data includes insertions and deletions (indels), and yet seed-chain-extend remains effective. In this paper, we generalize those prior results by introducing mathematical machinery to deal with the two new obstacles introduced by indel channels: the dependence of neighboring anchors and the presence of anchors that are only partially correct. We are thus able to prove that the expected recoverability of an optimal chain is ≥1−O(1/√m) and the expected runtime is O(mn^(3.15·θ_T) log n), when the total mutation rate given by the sum of the substitution, insertion, and deletion mutation rates (θ_T = θ_i + θ_d + θ_s) is less than 0.159.