Paper List
-
A Unified Variational Principle for Branching Transport Networks: Wave Impedance, Viscous Flow, and Tissue Metabolism
This paper solves the core problem of predicting the empirically observed branching exponent (α≈2.7) in mammalian arterial trees, which neither Murray...
-
Household Bubbling Strategies for Epidemic Control and Social Connectivity
This paper addresses the core challenge of designing household merging (social bubble) strategies that effectively control epidemic risk while maximiz...
-
Empowering Chemical Structures with Biological Insights for Scalable Phenotypic Virtual Screening
This paper addresses the core challenge of bridging the gap between scalable chemical structure screening and biologically informative but resource-in...
-
A mechanical bifurcation constrains the evolution of cell sheet folding in the family Volvocaceae
This paper addresses the core problem of why there is an evolutionary gap in species with intermediate cell numbers (e.g., 256 cells) in Volvocaceae, ...
-
Bayesian Inference in Epidemic Modelling: A Beginner’s Guide Illustrated with the SIR Model
This guide addresses the core challenge of estimating uncertain epidemiological parameters (like transmission and recovery rates) from noisy, real-wor...
-
Geometric framework for biological evolution
This paper addresses the fundamental challenge of developing a coordinate-independent, geometric description of evolutionary dynamics that bridges gen...
-
A multiscale discrete-to-continuum framework for structured population models
This paper addresses the core challenge of systematically deriving uniformly valid continuum approximations from discrete structured population models...
-
Whole slide and microscopy image analysis with QuPath and OMERO
使QuPath能够直接分析存储在OMERO服务器中的图像而无需下载整个数据集,克服了大规模研究的本地存储限制。
Learning From Limited Data and Feedback for Cell Culture Process Monitoring: A Comparative Study
Complex Adaptive Systems Laboratory, The Data Science Institute, University of Technology Sydney, NSW 2007, Australia | CSL Innovation, Melbourne, VIC 3000, Australia
30秒速读
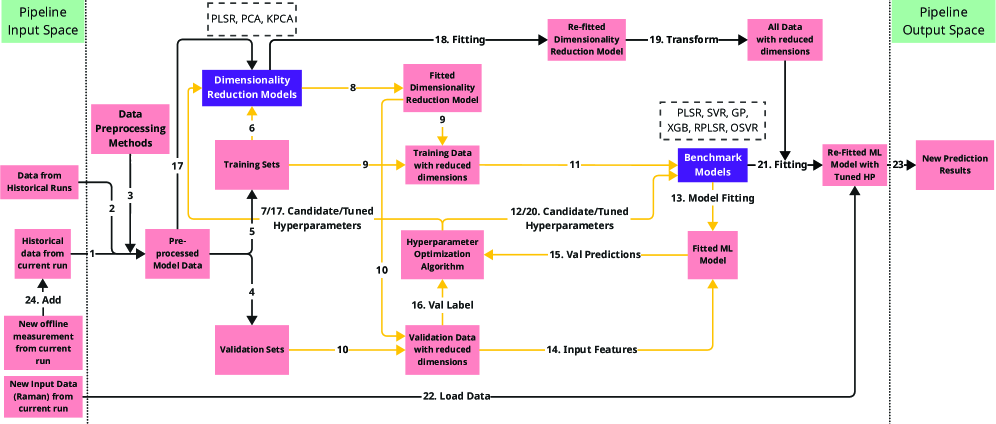
IN SHORT: This paper addresses the core challenge of developing accurate real-time bioprocess monitoring soft sensors under severe data constraints: limited historical batches, infrequent offline measurements (once/twice daily), heterogeneous process conditions, and high-dimensional Raman spectral inputs (3,325 wavenumbers).
核心创新
- Methodology Systematic benchmarking of three ML strategies (Dimensionality Reduction, Just-In-Time Learning, Online Learning) specifically tailored for cold-start bioprocess monitoring across simulated and real industrial datasets.
- Methodology Identification of key meta-features (feed media composition, process control strategies) that significantly impact model transferability between heterogeneous bioreactor runs.
- Methodology Demonstration that integrating Raman-based real-time predictions with lagged offline measurements enhances monitoring accuracy, providing a hybrid approach to overcome infrequent feedback.
主要结论
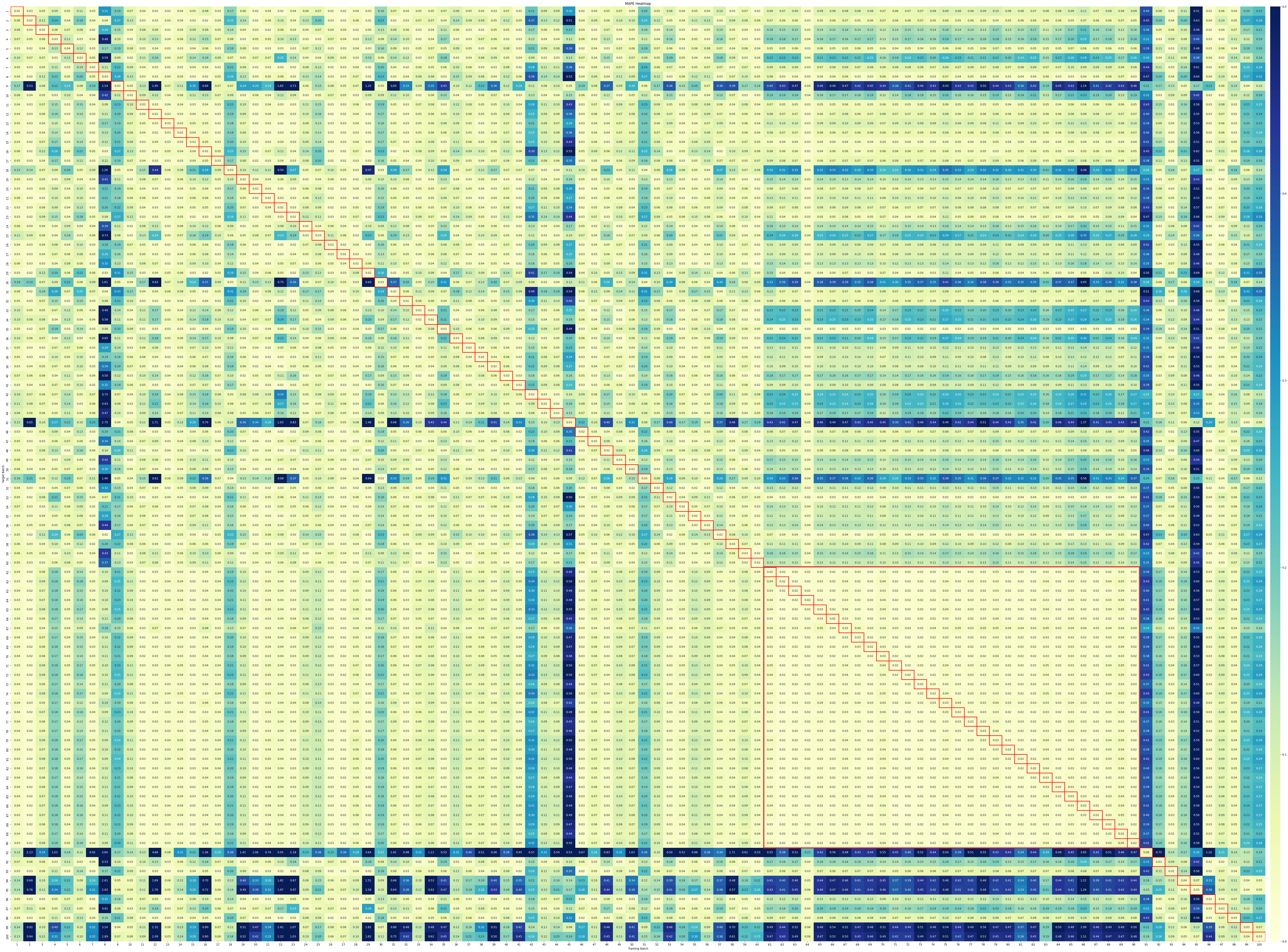
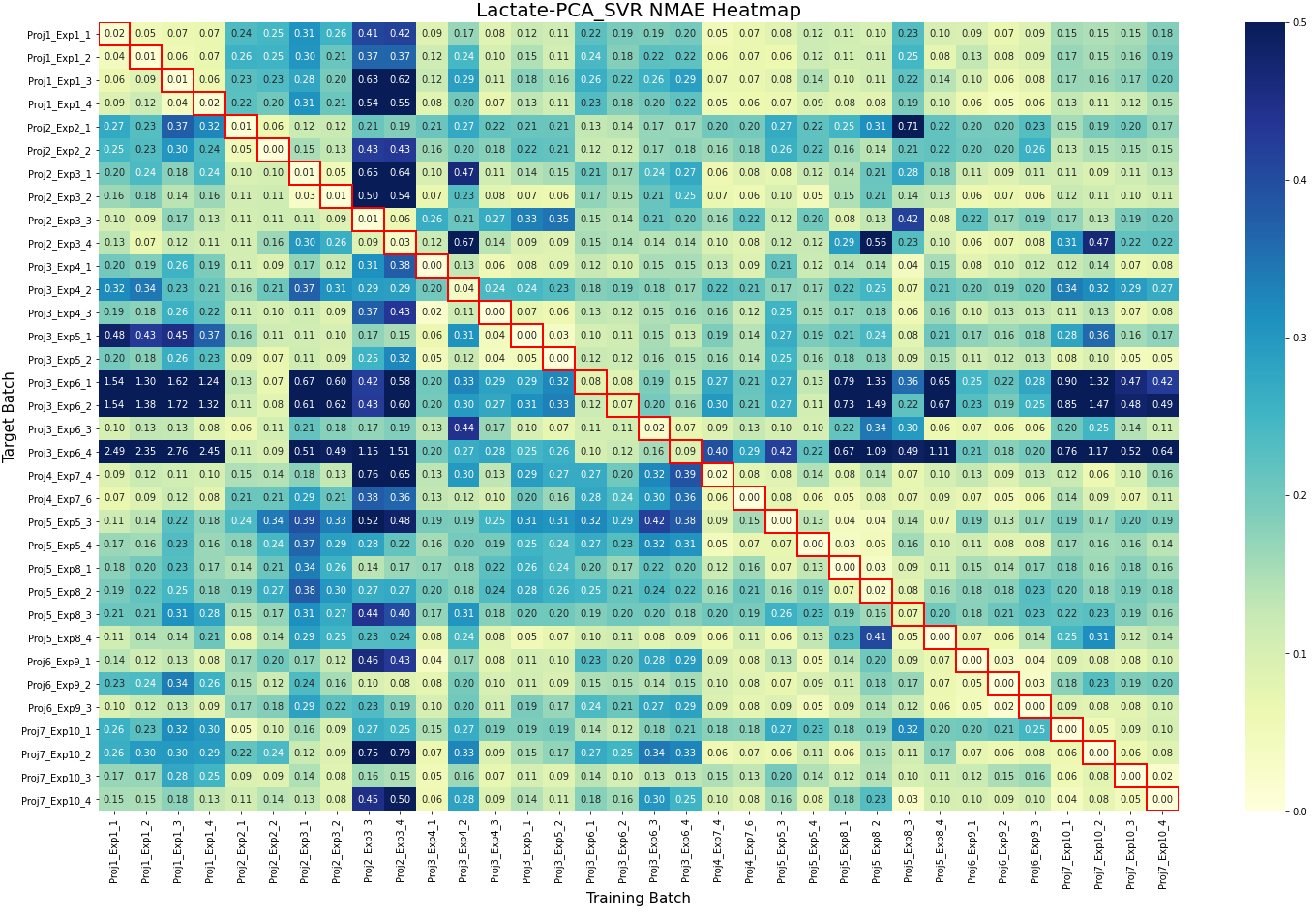
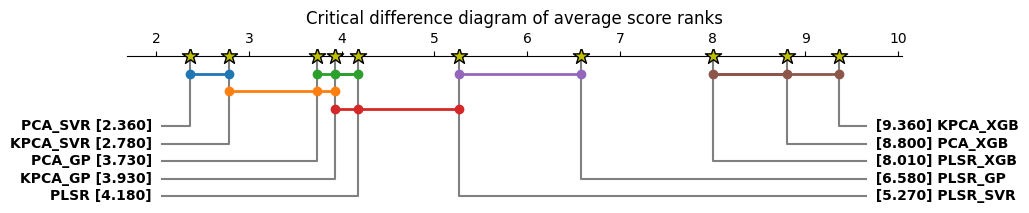
- Batch learning methods (e.g., PLSR, SVR) perform well in homogeneous settings but struggle in cold-start scenarios, where Just-In-Time Learning (JITL) and Online Learning (OL) show superior adaptability with statistically significant improvements (p<0.05 in Friedman tests).
- Dimensionality Reduction is critical for handling high-dimensional Raman data (3,325 features vs. <30 samples), with supervised methods like PLSR outperforming unsupervised PCA when offline measurements are available.
- Model transferability depends heavily on process meta-features; feed media composition explains up to 40% of performance variance across runs, highlighting the need for context-aware training strategies.
摘要: In cell culture bioprocessing, real-time batch process monitoring (BPM) refers to the continuous tracking and analysis of key process variables—such as viable cell density, nutrient levels, metabolite concentrations, and product titer—throughout the duration of a batch run. This enables early detection of deviations and supports timely control actions to ensure optimal cell growth and product quality. BPM plays a critical role in ensuring the quality and regulatory compliance of biopharmaceutical manufacturing processes. However, the development of accurate soft sensors for BPM is hindered by key challenges, including limited historical data, infrequent feedback, heterogeneous process conditions, and high-dimensional sensory inputs. This study presents a comprehensive benchmarking analysis of machine learning (ML) methods designed to address these challenges, with a focus on learning from historical data with limited volume and relevance in the context of bioprocess monitoring. We evaluate multiple ML approaches—including feature dimensionality reduction, online learning, and just-in-time learning—across three datasets, one in silico dataset and two real-world experimental datasets. Our findings highlight the importance of training strategies in handling limited data and feedback, with batch learning proving effective in homogeneous settings, while just-in-time learning and online learning demonstrate superior adaptability in cold-start scenarios. Additionally, we identify key meta-features, such as feed media composition and process control strategies, that significantly impact model transferability. The results also suggest that integrating Raman-based predictions with lagged offline measurements enhances monitoring accuracy, offering a promising direction for future bioprocess soft sensor development.