Paper List
-
Evolutionarily Stable Stackelberg Equilibrium
通过要求追随者策略对突变入侵具有鲁棒性,弥合了斯塔克尔伯格领导力模型与演化稳定性之间的鸿沟。
-
Recovering Sparse Neural Connectivity from Partial Measurements: A Covariance-Based Approach with Granger-Causality Refinement
通过跨多个实验会话累积协方差统计,实现从部分记录到完整神经连接性的重建。
-
Atomic Trajectory Modeling with State Space Models for Biomolecular Dynamics
ATMOS通过提供一个基于SSM的高效框架,用于生物分子的原子级轨迹生成,弥合了计算昂贵的MD模拟与时间受限的深度生成模型之间的差距。
-
Slow evolution towards generalism in a model of variable dietary range
通过证明是种群统计噪声(而非确定性动力学)驱动了模式形成和泛化食性的演化,解决了间接竞争下物种形成的悖论。
-
Grounded Multimodal Retrieval-Augmented Drafting of Radiology Impressions Using Case-Based Similarity Search
通过将印象草稿基于检索到的历史病例,并采用明确引用和基于置信度的拒绝机制,解决放射学报告生成中的幻觉问题。
-
Unified Policy–Value Decomposition for Rapid Adaptation
通过双线性分解在策略和价值函数之间共享低维目标嵌入,实现对新颖任务的零样本适应。
-
Mathematical Modeling of Cancer–Bacterial Therapy: Analysis and Numerical Simulation via Physics-Informed Neural Networks
提供了一个严格的、无网格的PINN框架,用于模拟和分析细菌癌症疗法中复杂的、空间异质的相互作用。
-
Sample-Efficient Adaptation of Drug-Response Models to Patient Tumors under Strong Biological Domain Shift
通过从无标记分子谱中学习可迁移表征,利用最少的临床数据实现患者药物反应的有效预测。
Learning From Limited Data and Feedback for Cell Culture Process Monitoring: A Comparative Study
Complex Adaptive Systems Laboratory, The Data Science Institute, University of Technology Sydney, NSW 2007, Australia | CSL Innovation, Melbourne, VIC 3000, Australia
30秒速读
IN SHORT: This paper addresses the core challenge of developing accurate real-time bioprocess monitoring soft sensors under severe data constraints: limited historical batches, infrequent offline measurements (once/twice daily), heterogeneous process conditions, and high-dimensional Raman spectral inputs (3,325 wavenumbers).
核心创新
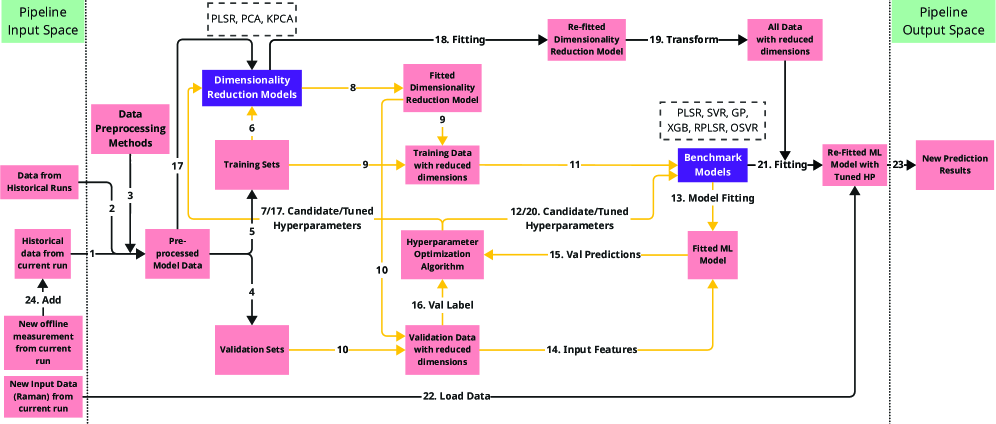
- Methodology Systematic benchmarking of three ML strategies (Dimensionality Reduction, Just-In-Time Learning, Online Learning) specifically tailored for cold-start bioprocess monitoring across simulated and real industrial datasets.
- Methodology Identification of key meta-features (feed media composition, process control strategies) that significantly impact model transferability between heterogeneous bioreactor runs.
- Methodology Demonstration that integrating Raman-based real-time predictions with lagged offline measurements enhances monitoring accuracy, providing a hybrid approach to overcome infrequent feedback.
主要结论
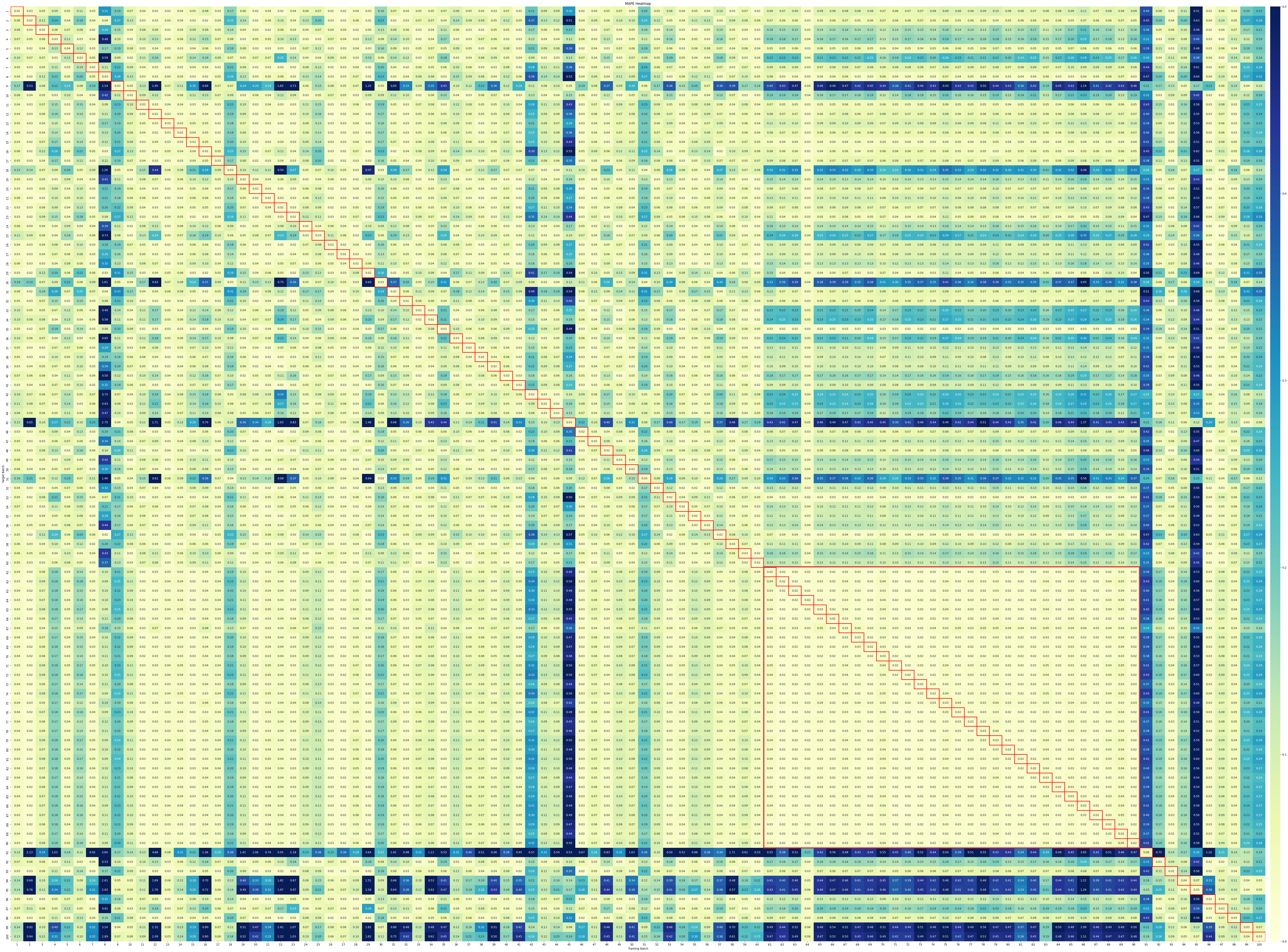
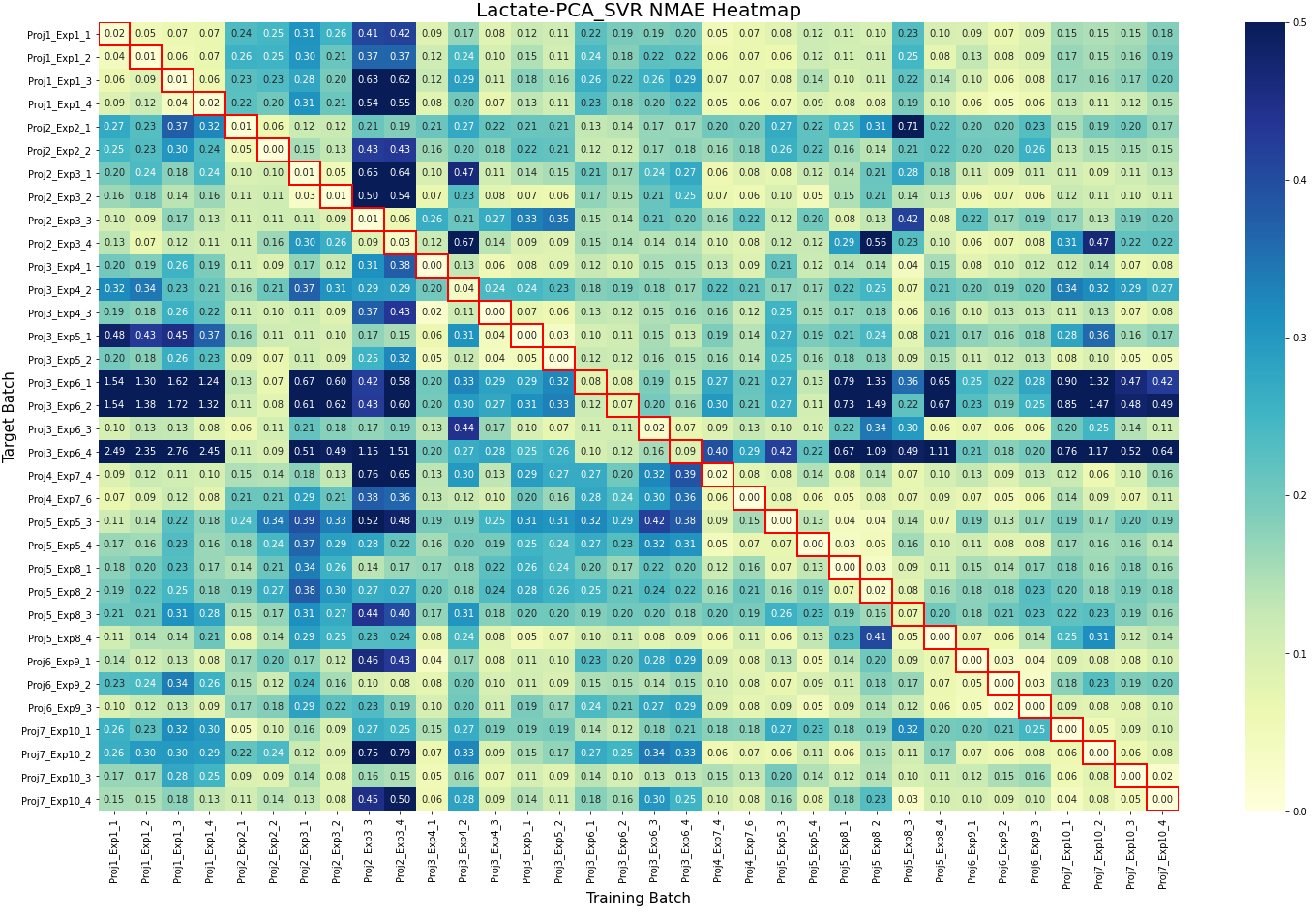
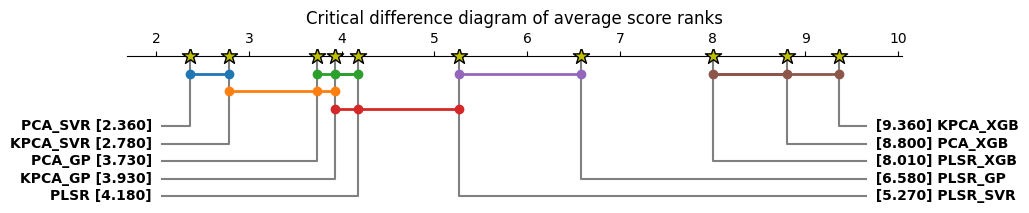
- Batch learning methods (e.g., PLSR, SVR) perform well in homogeneous settings but struggle in cold-start scenarios, where Just-In-Time Learning (JITL) and Online Learning (OL) show superior adaptability with statistically significant improvements (p<0.05 in Friedman tests).
- Dimensionality Reduction is critical for handling high-dimensional Raman data (3,325 features vs. <30 samples), with supervised methods like PLSR outperforming unsupervised PCA when offline measurements are available.
- Model transferability depends heavily on process meta-features; feed media composition explains up to 40% of performance variance across runs, highlighting the need for context-aware training strategies.
摘要: In cell culture bioprocessing, real-time batch process monitoring (BPM) refers to the continuous tracking and analysis of key process variables—such as viable cell density, nutrient levels, metabolite concentrations, and product titer—throughout the duration of a batch run. This enables early detection of deviations and supports timely control actions to ensure optimal cell growth and product quality. BPM plays a critical role in ensuring the quality and regulatory compliance of biopharmaceutical manufacturing processes. However, the development of accurate soft sensors for BPM is hindered by key challenges, including limited historical data, infrequent feedback, heterogeneous process conditions, and high-dimensional sensory inputs. This study presents a comprehensive benchmarking analysis of machine learning (ML) methods designed to address these challenges, with a focus on learning from historical data with limited volume and relevance in the context of bioprocess monitoring. We evaluate multiple ML approaches—including feature dimensionality reduction, online learning, and just-in-time learning—across three datasets, one in silico dataset and two real-world experimental datasets. Our findings highlight the importance of training strategies in handling limited data and feedback, with batch learning proving effective in homogeneous settings, while just-in-time learning and online learning demonstrate superior adaptability in cold-start scenarios. Additionally, we identify key meta-features, such as feed media composition and process control strategies, that significantly impact model transferability. The results also suggest that integrating Raman-based predictions with lagged offline measurements enhances monitoring accuracy, offering a promising direction for future bioprocess soft sensor development.