Paper List
-
Formation of Artificial Neural Assemblies by Biologically Plausible Inhibition Mechanisms
This work addresses the core limitation of the Assembly Calculus model—its fixed-size, biologically implausible k-WTA selection process—by introducing...
-
How to make the most of your masked language model for protein engineering
This paper addresses the critical bottleneck of efficiently sampling high-quality, diverse protein sequences from Masked Language Models (MLMs) for pr...
-
Module control in youth symptom networks across COVID-19
This paper addresses the core challenge of distinguishing whether a prolonged societal stressor (COVID-19) fundamentally reorganizes the architecture ...
-
JEDI: Jointly Embedded Inference of Neural Dynamics
This paper addresses the core challenge of inferring context-dependent neural dynamics from noisy, high-dimensional recordings using a single unified ...
-
ATP Level and Phosphorylation Free Energy Regulate Trigger-Wave Speed and Critical Nucleus Size in Cellular Biochemical Systems
This work addresses the core challenge of quantitatively predicting how the cellular energy state (ATP level and phosphorylation free energy) governs ...
-
Packaging Jupyter notebooks as installable desktop apps using LabConstrictor
This paper addresses the core pain point of ensuring Jupyter notebook reproducibility and accessibility across different computing environments, parti...
-
SNPgen: Phenotype-Supervised Genotype Representation and Synthetic Data Generation via Latent Diffusion
This paper addresses the core challenge of generating privacy-preserving synthetic genotype data that maintains both statistical fidelity and downstre...
-
Continuous Diffusion Transformers for Designing Synthetic Regulatory Elements
This paper addresses the challenge of efficiently generating novel, cell-type-specific regulatory DNA sequences with high predicted activity while min...
SpikGPT: A High-Accuracy and Interpretable Spiking Attention Framework for Single-Cell Annotation
Department of Biomedical Informatics, Emory University | Department of Surgery, Duke University
30秒速读
IN SHORT: This paper addresses the core challenge of robust single-cell annotation across heterogeneous datasets with batch effects and the critical need to identify previously unseen cell populations.
核心创新
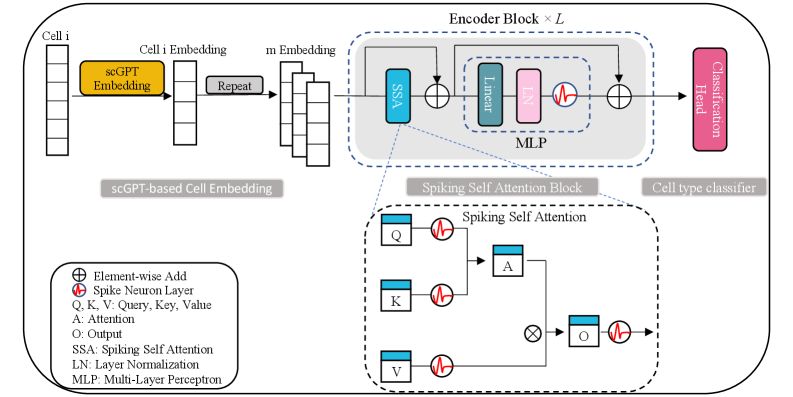
- Methodology First integration of spiking neural networks with transformer architecture for single-cell analysis, using Leaky Integrate-and-Fire (LIF) neurons in a multi-head Spiking Self-Attention mechanism for energy-efficient computation.
- Methodology Novel two-step embedding expansion strategy: repeating cell embeddings along feature channels (default m=300) and temporal dimensions (default T=4) to enhance representation richness and training stability.
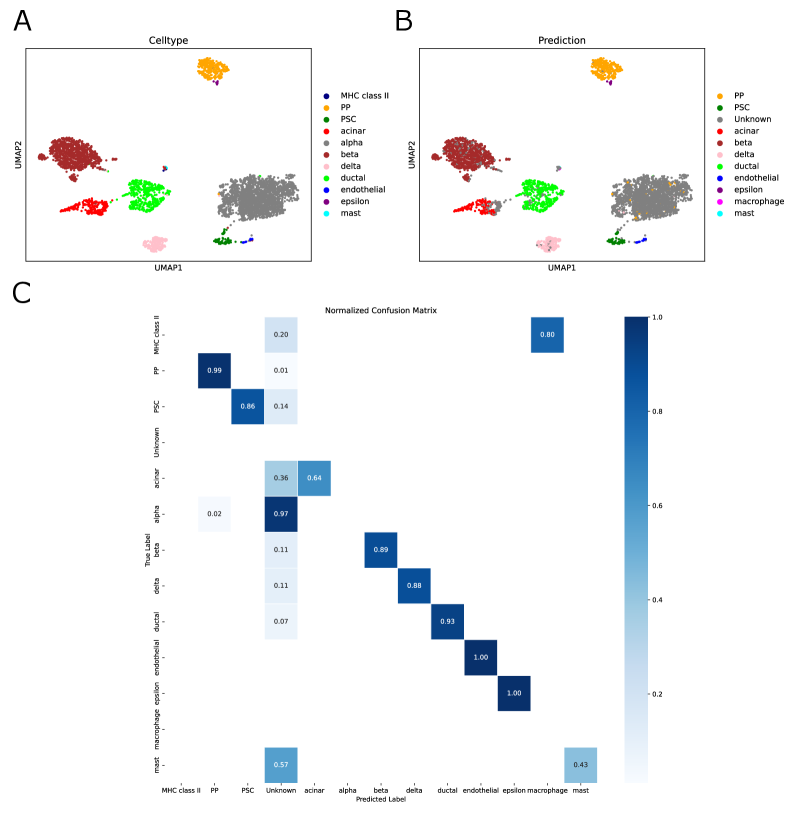
- Biology Confidence-based rejection mechanism that successfully identifies 97% of unseen 'alpha cells' as 'Unknown' in pancreas datasets, enabling robust detection of novel cell types absent from training data.
主要结论
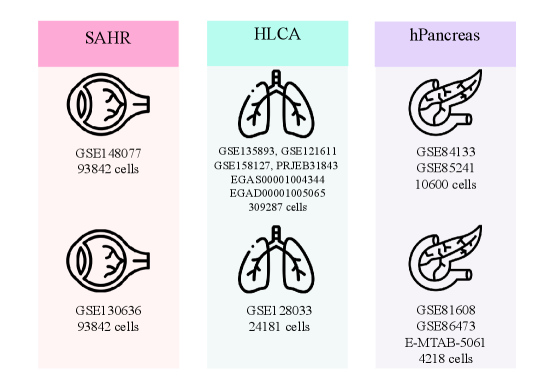
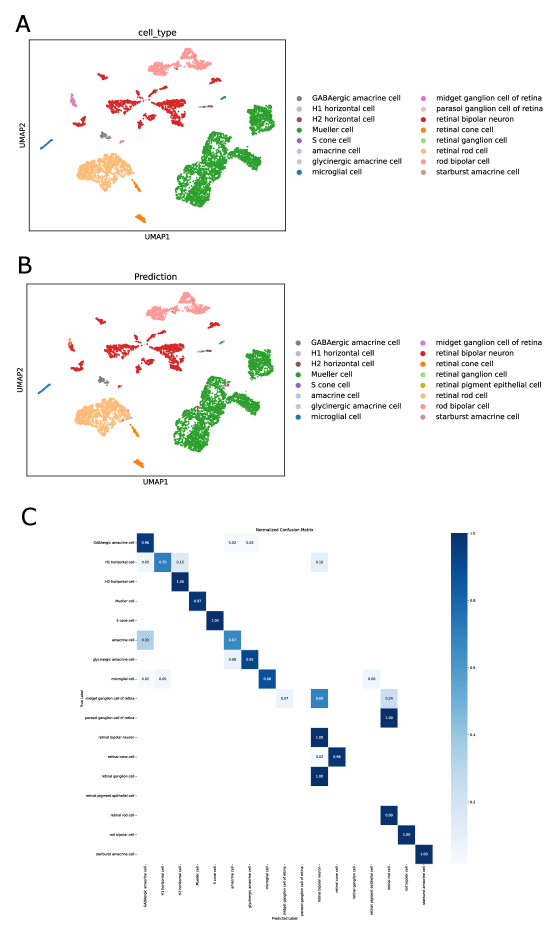
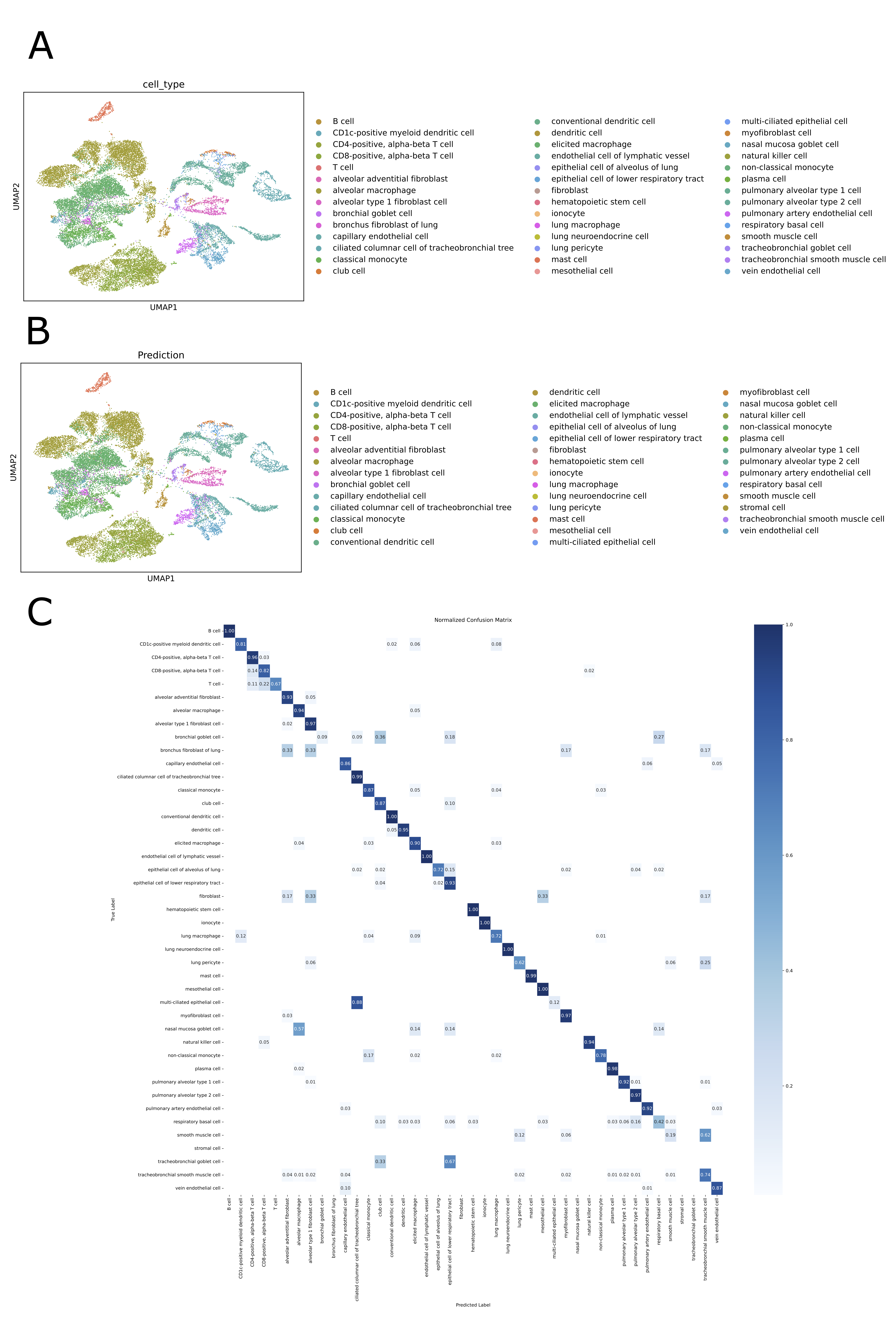
- SpikGPT achieves accuracy of 0.991 on SAHR dataset and 0.920 on HLCA dataset, outperforming or matching 8 benchmark methods including scGPT, CCA, and scPred.
- The model demonstrates superior robustness to batch effects, maintaining macro F1-score of 0.711 on heterogeneous HLCA data where traditional methods like SingleR drop to 0.207 F1-score.
- SpikGPT successfully identifies 97% of unseen 'alpha cells' as 'Unknown' using confidence thresholding (p<0.05), enabling reliable detection of novel cell populations.
摘要: Accurate and scalable cell type annotation remains a challenge in single-cell transcriptomics, especially when datasets exhibit strong batch effects or contain previously unseen cell populations. Here we introduce SpikGPT, a hybrid deep learning framework that integrates scGPT-derived cell embeddings with a spiking Transformer architecture to achieve efficient and robust annotation. scGPT provides biologically informed dense representations of each cell, which are further processed by a multi-head Spiking Self-Attention mechanism, energy-efficient feature extraction. Across multiple benchmark datasets, SpikGPT consistently matches or exceeds the performance of leading annotation tools. Notably, SpikGPT uniquely identifies unseen cell types by assigning low-confidence predictions to an 'Unknown' category, allowing accurate rejection of cell states absent from the training reference. Together, these results demonstrate that SpikGPT is a versatile and reliable annotation tool capable of generalizing across datasets, resolving complex cellular heterogeneity, and facilitating discovery of novel or disease-associated cell populations.