Paper List
-
MCP-AI: Protocol-Driven Intelligence Framework for Autonomous Reasoning in Healthcare
This paper addresses the critical gap in healthcare AI systems that lack contextual reasoning, long-term state management, and verifiable workflows by...
-
Model Gateway: Model Management Platform for Model-Driven Drug Discovery
This paper addresses the critical bottleneck of fragmented, ad-hoc model management in pharmaceutical research by providing a centralized, scalable ML...
-
Tree Thinking in the Genomic Era: Unifying Models Across Cells, Populations, and Species
This paper addresses the fragmentation of tree-based inference methods across biological scales by identifying shared algorithmic principles and stati...
-
SSDLabeler: Realistic semi-synthetic data generation for multi-label artifact classification in EEG
This paper addresses the core challenge of training robust multi-label EEG artifact classifiers by overcoming the scarcity and limited diversity of ma...
-
Decoding Selective Auditory Attention to Musical Elements in Ecologically Valid Music Listening
This paper addresses the core challenge of objectively quantifying listeners' selective attention to specific musical components (e.g., vocals, drums,...
-
Physics-Guided Surrogate Modeling for Machine Learning–Driven DLD Design Optimization
This paper addresses the core bottleneck of translating microfluidic DLD devices from research prototypes to clinical applications by replacing weeks-...
-
Mechanistic Interpretability of Antibody Language Models Using SAEs
This work addresses the core challenge of achieving both interpretability and controllable generation in domain-specific protein language models, spec...
-
Fluctuating Environments Favor Extreme Dormancy Strategies and Penalize Intermediate Ones
This paper addresses the core challenge of determining how organisms should tune dormancy duration to match the temporal autocorrelation of their envi...
Few-shot Protein Fitness Prediction via In-context Learning and Test-time Training
Department of Systems Biology, Harvard Medical School | Department of Biology, University of Copenhagen | Machine Intelligence, Novo Nordisk A/S | Microsoft Research, Cambridge, MA, USA | Dept. of Applied Mathematics and Computer Science, Technical University of Denmark
30秒速读
IN SHORT: This paper addresses the core challenge of accurately predicting protein fitness with only a handful of experimental observations, where data collection is prohibitively expensive and label availability is severely limited.
核心创新
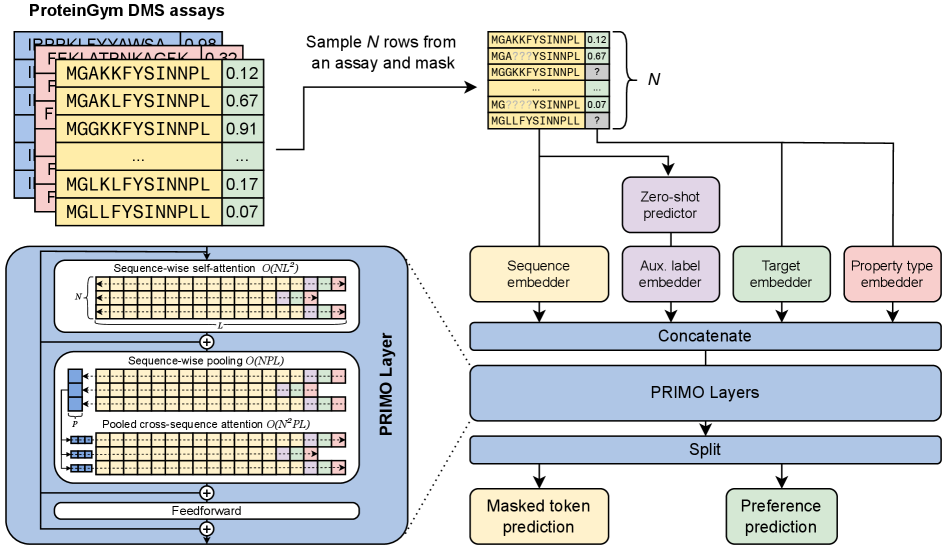
- Methodology Introduces PRIMO, a novel transformer-based framework that uniquely combines in-context learning with test-time training for few-shot protein fitness prediction.
- Methodology Proposes a hybrid masked token reconstruction objective with a preference-based loss function, enabling effective learning from sparse experimental labels across diverse assays.
- Methodology Develops a lightweight pooling attention mechanism that handles both substitution and indel mutations while maintaining computational efficiency, overcoming limitations of previous methods.
主要结论
- PRIMO with test-time training (TTT) achieves state-of-the-art few-shot performance, improving from a zero-shot Spearman correlation of 0.51 to 0.67 with 128 shots, outperforming Gaussian Process (0.56) and Ridge Regression (0.63) baselines.
- The framework demonstrates broad applicability across protein properties including stability (0.77 correlation with TTT), enzymatic activity (0.61), fluorescence (0.30), and binding (0.69), handling both substitution and indel mutations.
- PRIMO's performance highlights the critical importance of proper data splitting to avoid inflated results, as demonstrated by the 0.4 correlation inflation on RL40A_YEAST when using Metalic's overlapping train-test split.
摘要: Accurately predicting protein fitness with minimal experimental data is a persistent challenge in protein engineering. We introduce PRIMO (PRotein In-context Mutation Oracle), a transformer-based framework that leverages in-context learning and test-time training to adapt rapidly to new proteins and assays without large task-specific datasets. By encoding sequence information, auxiliary zero-shot predictions, and sparse experimental labels from many assays as a unified token set in a pre-training masked-language modeling paradigm, PRIMO learns to prioritize promising variants through a preference-based loss function. Across diverse protein families and properties—including both substitution and indel mutations—PRIMO outperforms zero-shot and fully supervised baselines. This work underscores the power of combining large-scale pre-training with efficient test-time adaptation to tackle challenging protein design tasks where data collection is expensive and label availability is limited.