Paper List
-
The Effective Reproduction Number in the Kermack-McKendrick model with age of infection and reinfection
This paper addresses the challenge of accurately estimating the time-varying effective reproduction number ℛ(t) in epidemics by incorporating two crit...
-
Covering Relations in the Poset of Combinatorial Neural Codes
This work addresses the core challenge of navigating the complex poset structure of neural codes to systematically test the conjecture linking convex ...
-
Collective adsorption of pheromones at the water-air interface
This paper addresses the core challenge of understanding how amphiphilic pheromones, previously assumed to be transported in the gas phase, can be sta...
-
pHapCompass: Probabilistic Assembly and Uncertainty Quantification of Polyploid Haplotype Phase
This paper addresses the core challenge of accurately assembling polyploid haplotypes from sequencing data, where read assignment ambiguity and an exp...
-
Setting up for failure: automatic discovery of the neural mechanisms of cognitive errors
This paper addresses the core challenge of automating the discovery of biologically plausible recurrent neural network (RNN) dynamics that can replica...
-
Influence of Object Affordance on Action Language Understanding: Evidence from Dynamic Causal Modeling Analysis
This study addresses the core challenge of moving beyond correlational evidence to establish the *causal direction* and *temporal dynamics* of how obj...
-
Revealing stimulus-dependent dynamics through statistical complexity
This paper addresses the core challenge of detecting stimulus-specific patterns in neural population dynamics that remain hidden to traditional variab...
-
Exactly Solvable Population Model with Square-Root Growth Noise and Cell-Size Regulation
This paper addresses the fundamental gap in understanding how microscopic growth fluctuations, specifically those with size-dependent (square-root) no...
On the Approximation of Phylogenetic Distance Functions by Artificial Neural Networks
Indiana University, Bloomington, IN 47405, USA
30秒速读
IN SHORT: This paper addresses the core challenge of developing computationally efficient and scalable neural network architectures that can learn accurate phylogenetic distance functions from simulated data, bridging the gap between simple distance methods and complex model-based inference.
核心创新
- Methodology Introduces minimal, permutation-invariant neural architectures (Sequence networks S and Pair networks P) specifically designed to approximate phylogenetic distance functions, ensuring invariance to taxa ordering without costly data augmentation.
- Methodology Leverages theoretical results from metric embedding (Bourgain's theorem, Johnson-Lindenstrauss Lemma) to inform network design, explicitly linking embedding dimension to the number of taxa for efficient representation.
- Methodology Demonstrates how equivariant layers and attention mechanisms can be structured to handle both i.i.d. and spatially correlated sequence data (e.g., models with indels or rate variation), adapting to the complexity of the generative evolutionary model.
主要结论
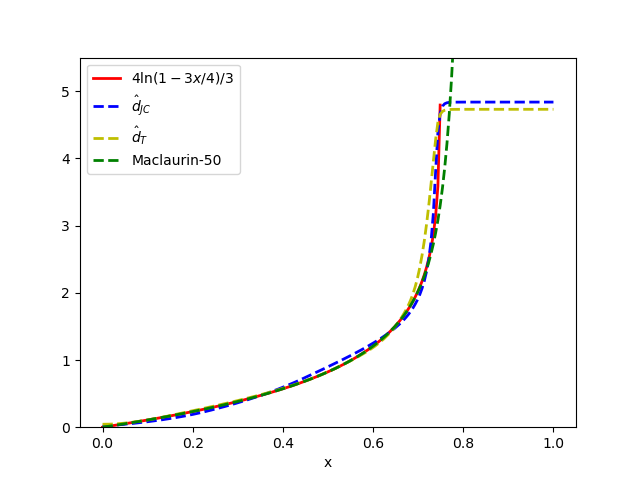
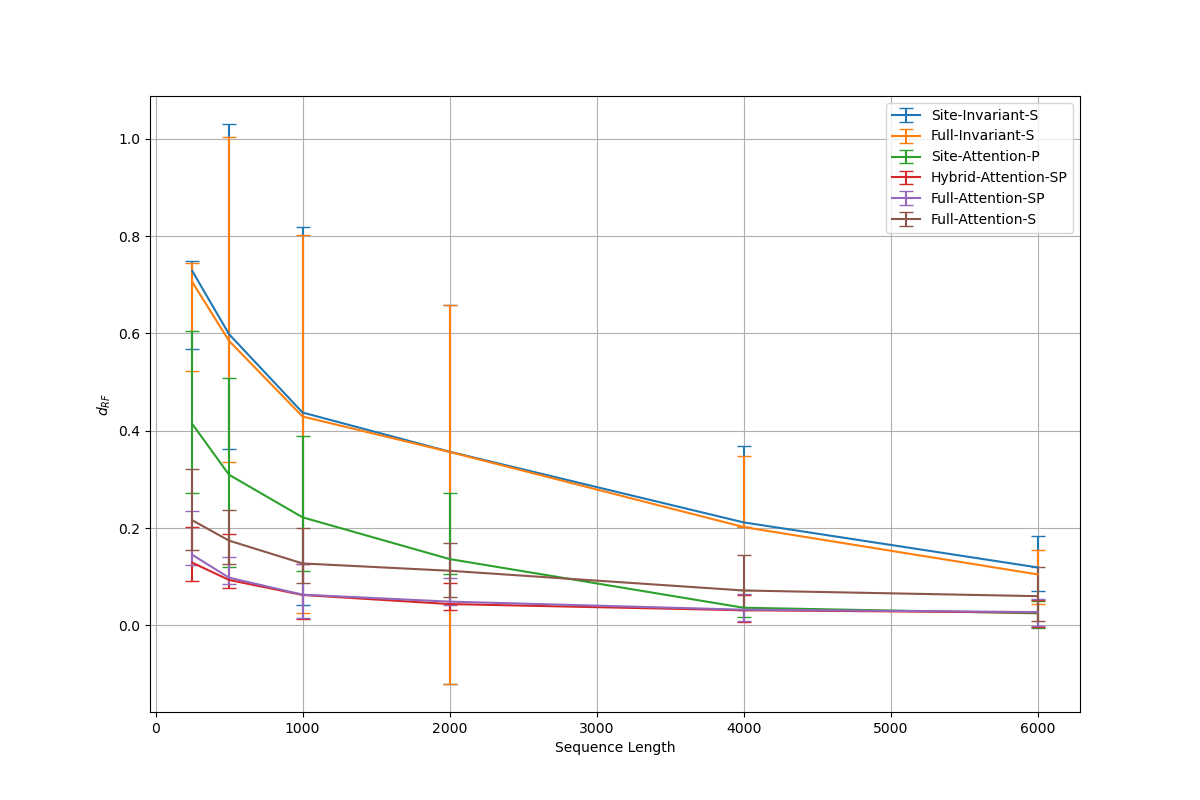
- The proposed minimal architectures (e.g., Sites-Invariant-S with ~7.6K parameters) achieve results comparable to state-of-the-art inference methods like IQ-TREE on simulated data under various models (JC, K2P, HKY, LG+indels), outperforming classic pairwise distance methods (d_H, d_JC, d_K2P) in most conditions.
- Architectures incorporating taxa-wise attention, while more memory-intensive, are necessary for complex evolutionary models with spatial dependencies; however, simpler networks suffice for simpler i.i.d. models, indicating an architecture-evolutionary model correspondence.
- Performance is highly sensitive to hyperparameters: validation error increases sharply with fewer than 4 attention heads or with hidden channel counts outside an optimal range (e.g., 32-128), aligning with theoretical requirements for learning graph-structured data.
摘要: Inferring the phylogenetic relationships among a sample of organisms is a fundamental problem in modern biology. While distance-based hierarchical clustering algorithms achieved early success on this task, these have been supplanted by Bayesian and maximum likelihood search procedures based on complex models of molecular evolution. In this work we describe minimal neural network architectures that can approximate classic phylogenetic distance functions and the properties required to learn distances under a variety of molecular evolutionary models. In contrast to model-based inference (and recently proposed model-free convolutional and transformer networks), these architectures have a small computational footprint and are scalable to large numbers of taxa and molecular characters. The learned distance functions generalize well and, given an appropriate training dataset, achieve results comparable to state-of-the art inference methods.