Paper List
-
Macroscopic Dominance from Microscopic Extremes: Symmetry Breaking in Spatial Competition
This paper addresses the fundamental question of how microscopic stochastic advantages in spatial exploration translate into macroscopic resource domi...
-
Linear Readout of Neural Manifolds with Continuous Variables
This paper addresses the core challenge of quantifying how the geometric structure of high-dimensional neural population activity (neural manifolds) d...
-
Theory of Cell Body Lensing and Phototaxis Sign Reversal in “Eyeless” Mutants of Chlamydomonas
This paper solves the core puzzle of how eyeless mutants of Chlamydomonas exhibit reversed phototaxis by quantitatively modeling the competition betwe...
-
Cross-Species Transfer Learning for Electrophysiology-to-Transcriptomics Mapping in Cortical GABAergic Interneurons
This paper addresses the challenge of predicting transcriptomic identity from electrophysiological recordings in human cortical interneurons, where li...
-
Uncovering statistical structure in large-scale neural activity with Restricted Boltzmann Machines
This paper addresses the core challenge of modeling large-scale neural population activity (1500-2000 neurons) with interpretable higher-order interac...
-
Realizing Common Random Numbers: Event-Keyed Hashing for Causally Valid Stochastic Models
This paper addresses the critical problem that standard stateful PRNG implementations in agent-based models violate causal validity by making random d...
-
A Standardized Framework for Evaluating Gene Expression Generative Models
This paper addresses the critical lack of standardized evaluation protocols for single-cell gene expression generative models, where inconsistent metr...
-
Single Molecule Localization Microscopy Challenge: A Biologically Inspired Benchmark for Long-Sequence Modeling
This paper addresses the core challenge of evaluating state-space models on biologically realistic, sparse, and stochastic temporal processes, which a...
CONFIDE: Hallucination Assessment for Reliable Biomolecular Structure Prediction and Design
The Chinese University of Hong Kong | Zhejiang University | Macao Polytechnic University | University of Electronic Science and Technology of China
30秒速读
IN SHORT: This paper addresses the critical limitation of current protein structure prediction models (like AlphaFold3) where high-confidence scores (pLDDT) can be misleading, failing to detect subtle structural errors like atomic clashes and topological traps, which undermines reliability in downstream applications like drug discovery.
核心创新
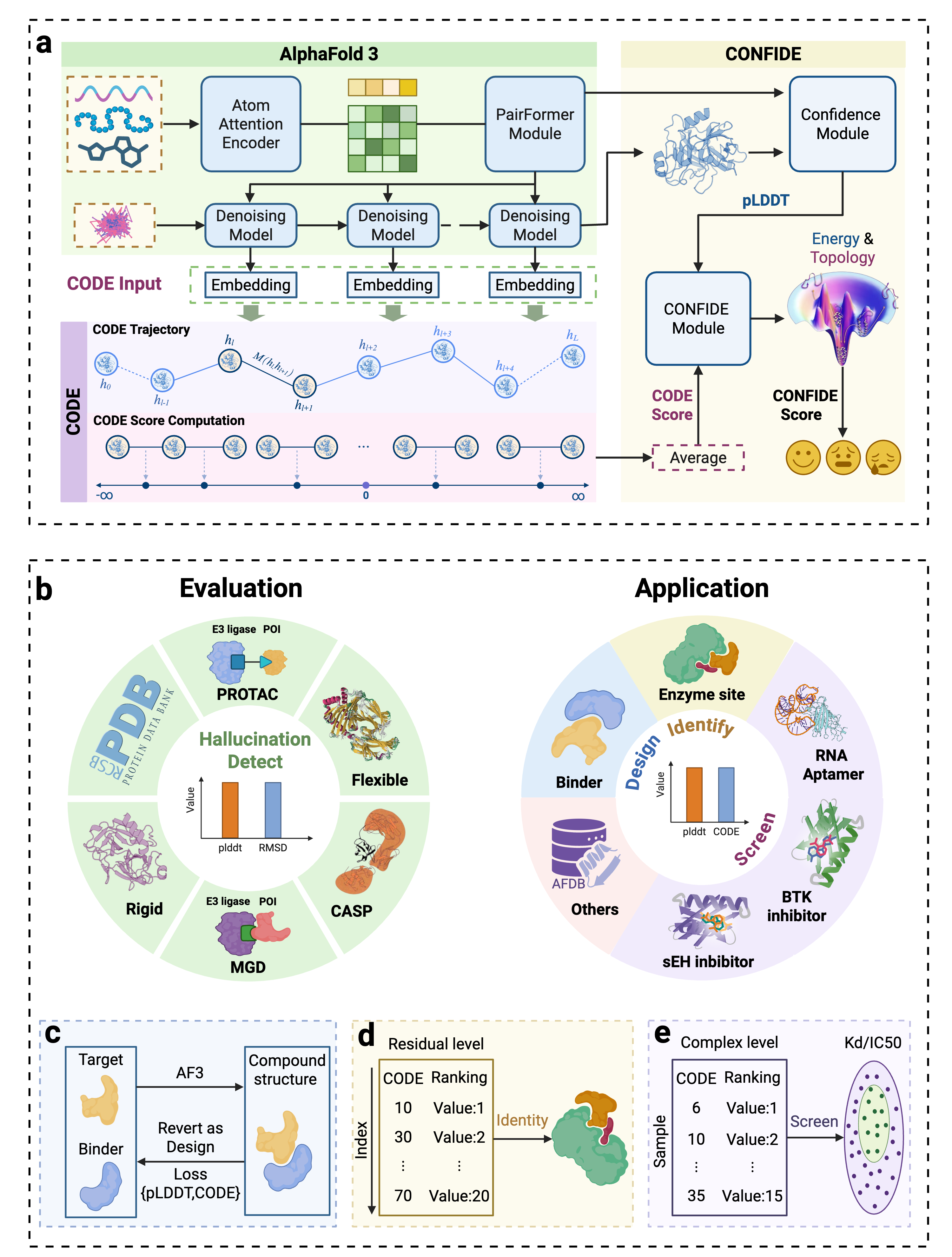
- Methodology Introduces CODE (Chain of Diffusion Embeddings), a novel, unsupervised metric derived from AlphaFold3's latent diffusion embeddings that directly quantifies topological frustration, a key factor in protein folding kinetics previously overlooked by confidence scores.
- Methodology Proposes CONFIDE, a unified evaluation framework that integrates the energetic perspective of pLDDT with the topological perspective of CODE, providing a more comprehensive and reliable assessment of predicted biomolecular structures.
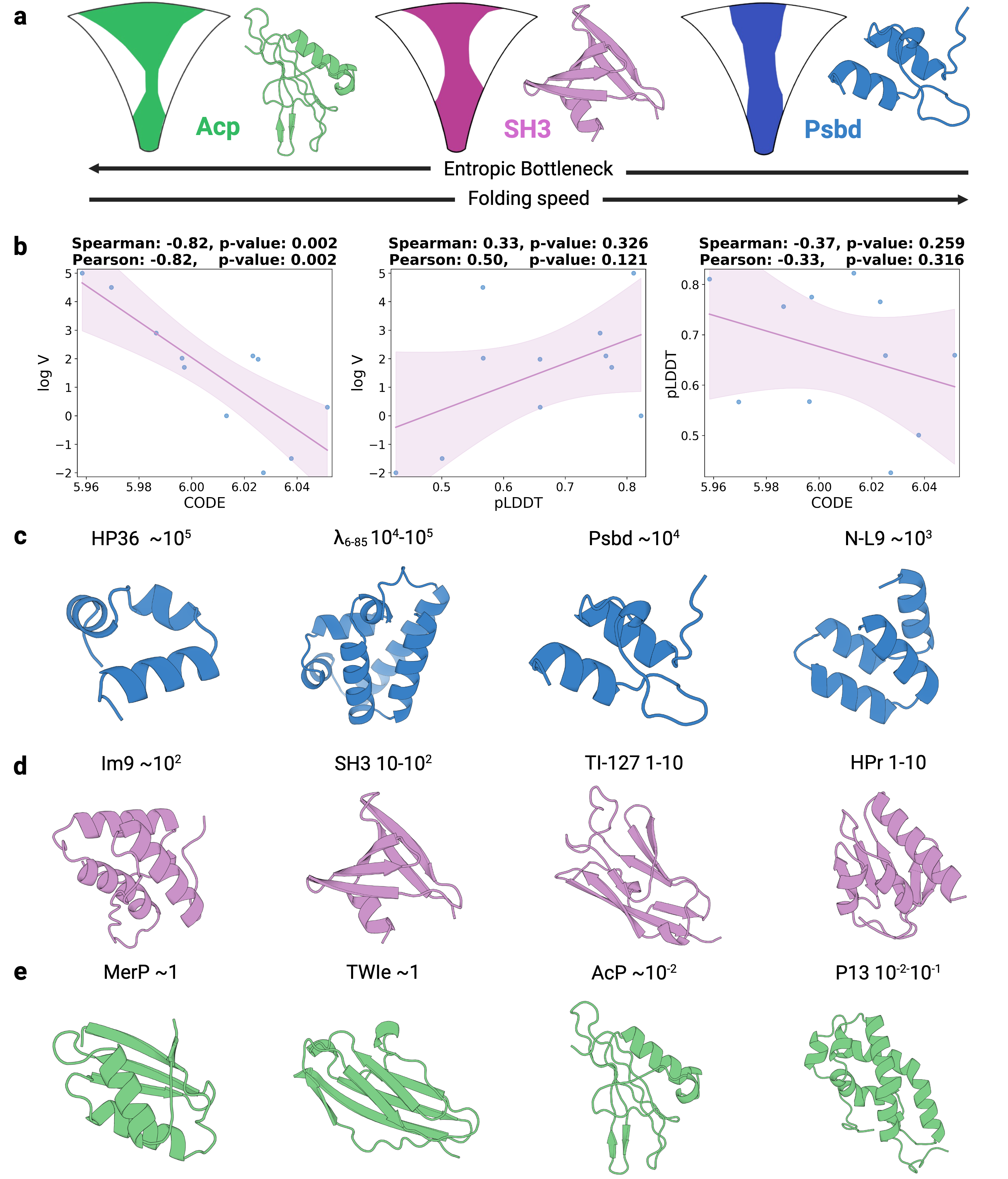
- Biology Establishes a strong empirical link between the CODE metric and protein folding rates driven by topological frustration (Spearman correlation of -0.82, p=0.002), offering a data-driven proxy for a complex biophysical phenomenon.
主要结论
- CODE demonstrates a strong, statistically significant correlation with protein folding rates mediated by topological frustration (Spearman ρ = -0.82, p=0.002), far outperforming pLDDT (ρ = 0.33, p=0.326).
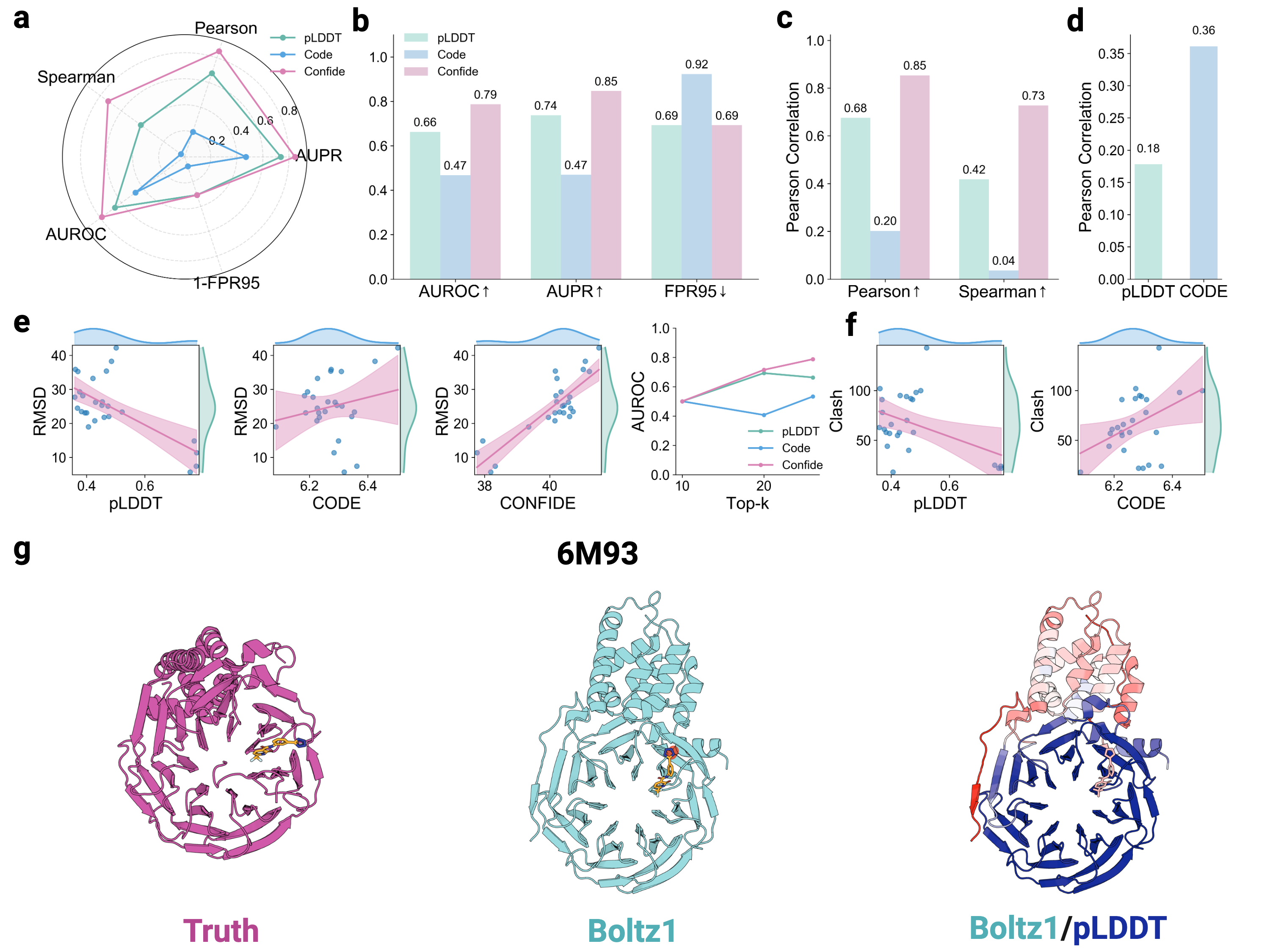
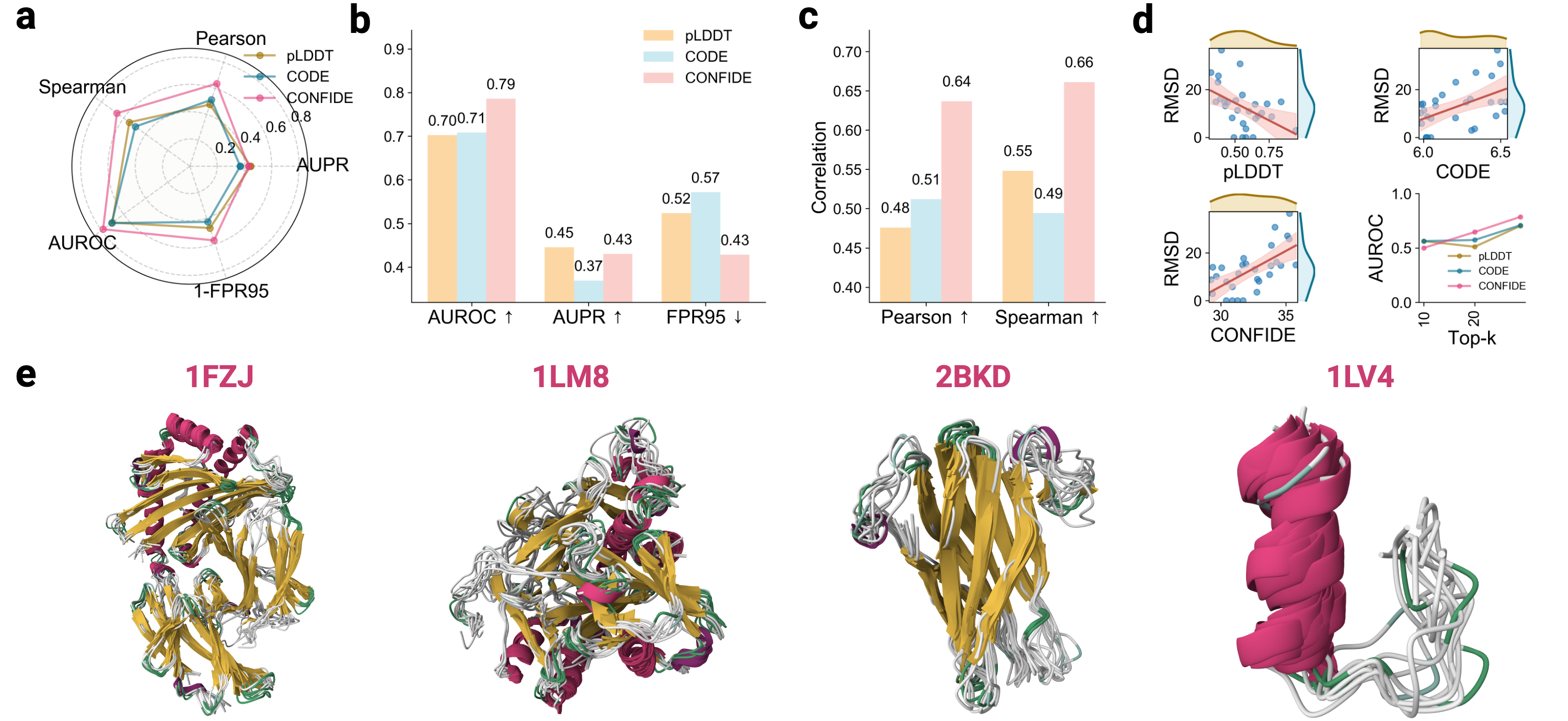
- The CONFIDE framework significantly improves hallucination detection, achieving a Spearman correlation of 0.73 with RMSD on molecular glue benchmarks, a 73.8% relative improvement over pLDDT's correlation of 0.42.
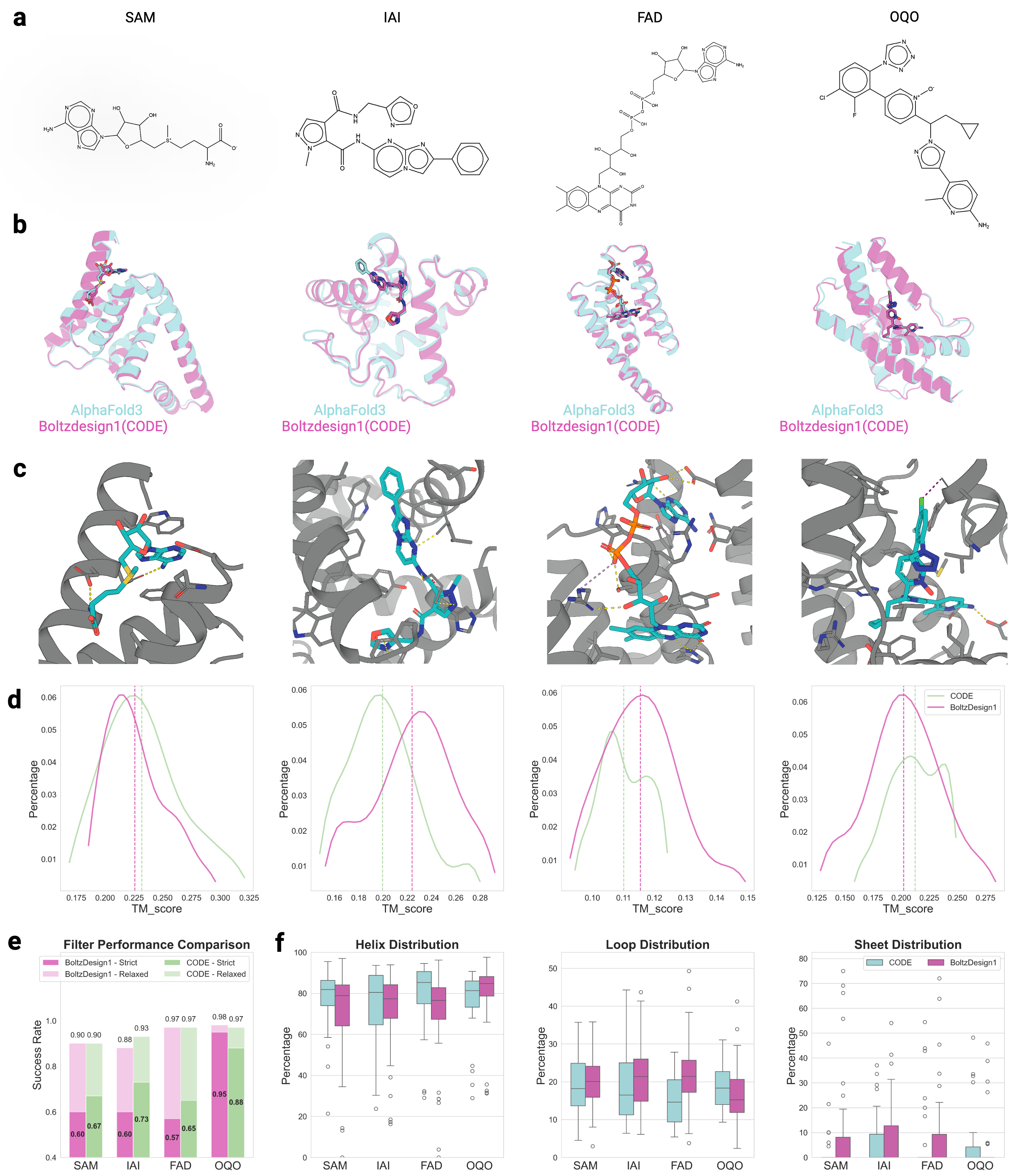
- CONFIDE enables practical downstream applications, improving binder design success rates (e.g., +13% for IAI) and accurately predicting mutation-induced binding affinity changes (Spearman ρ = 0.83 for BTK vs. Fenebrutinib, compared to pLDDT's ρ = 0.03).
摘要: Reliable evaluation of protein structure predictions remains challenging, as metrics like pLDDT capture energetic stability but often miss subtle errors such as atomic clashes or conformational traps reflecting topological frustration within the protein-folding energy landscape. We present CODE (Chain of Diffusion Embeddings), a self-evaluating metric empirically found to quantify topological frustration directly from the latent diffusion embeddings of the AlphaFold3 series of structure predictors in a fully unsupervised manner. Integrating this with pLDDT, we propose CONFIDE, a unified evaluation framework that combines energetic and topological perspectives to improve the reliability of AlphaFold3 and related models. CODE strongly correlates with protein folding rates driven by topological frustration, achieving a correlation of 0.82 compared to pLDDT’s 0.33 (a relative improvement of 148%). CONFIDE significantly enhances the reliability of quality evaluation in molecular glue structure prediction benchmarks, achieving a Spearman correlation of 0.73 with RMSD, compared to pLDDT’s correlation of 0.42, a relative improvement of 73.8%. Beyond quality assessment, our approach applies to diverse drug-design tasks, including all-atom binder design, enzymatic active-site mapping, mutation-induced binding-affinity prediction, nucleic acid aptamer screening, and flexible protein modeling. By combining data-driven embeddings with theoretical insight, CODE and CONFIDE outperform existing metrics across a wide range of biomolecular systems, offering robust and versatile tools to refine structure predictions, advance structural biology, and accelerate drug discovery.