Paper List
-
Macroscopic Dominance from Microscopic Extremes: Symmetry Breaking in Spatial Competition
This paper addresses the fundamental question of how microscopic stochastic advantages in spatial exploration translate into macroscopic resource domi...
-
Linear Readout of Neural Manifolds with Continuous Variables
This paper addresses the core challenge of quantifying how the geometric structure of high-dimensional neural population activity (neural manifolds) d...
-
Theory of Cell Body Lensing and Phototaxis Sign Reversal in “Eyeless” Mutants of Chlamydomonas
This paper solves the core puzzle of how eyeless mutants of Chlamydomonas exhibit reversed phototaxis by quantitatively modeling the competition betwe...
-
Cross-Species Transfer Learning for Electrophysiology-to-Transcriptomics Mapping in Cortical GABAergic Interneurons
This paper addresses the challenge of predicting transcriptomic identity from electrophysiological recordings in human cortical interneurons, where li...
-
Uncovering statistical structure in large-scale neural activity with Restricted Boltzmann Machines
This paper addresses the core challenge of modeling large-scale neural population activity (1500-2000 neurons) with interpretable higher-order interac...
-
Realizing Common Random Numbers: Event-Keyed Hashing for Causally Valid Stochastic Models
This paper addresses the critical problem that standard stateful PRNG implementations in agent-based models violate causal validity by making random d...
-
A Standardized Framework for Evaluating Gene Expression Generative Models
This paper addresses the critical lack of standardized evaluation protocols for single-cell gene expression generative models, where inconsistent metr...
-
Single Molecule Localization Microscopy Challenge: A Biologically Inspired Benchmark for Long-Sequence Modeling
This paper addresses the core challenge of evaluating state-space models on biologically realistic, sparse, and stochastic temporal processes, which a...
Hierarchical Molecular Language Models (HMLMs)
Department of Chemical Engineering, University of Arkansas, Fayetteville, AR 72701, USA
30秒速读
IN SHORT: This paper addresses the core challenge of accurately modeling context-dependent signaling, pathway cross-talk, and temporal dynamics across multiple biological scales in cellular signaling networks.
核心创新
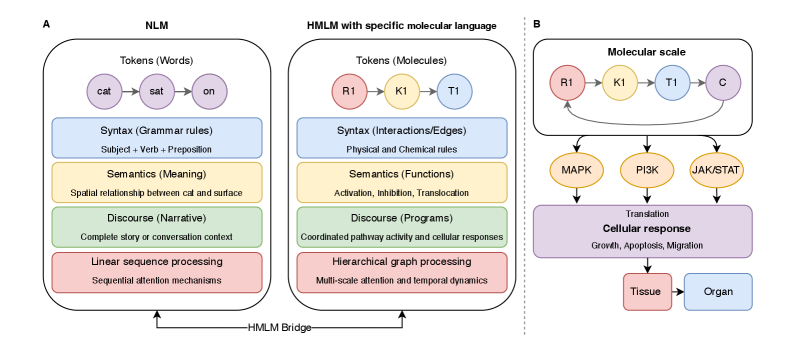
- Methodology Introduces cellular signaling as a molecular language with unique grammar and semantics, establishing a theoretical foundation for molecular artificial intelligence (MAI).
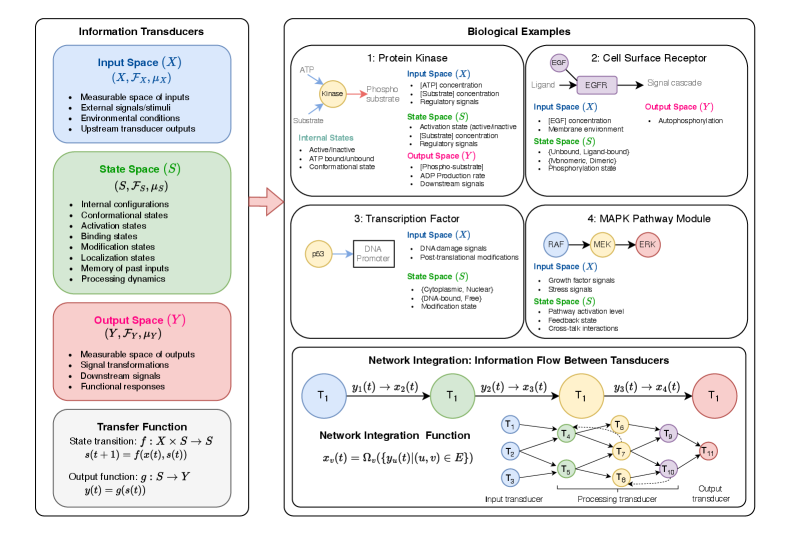
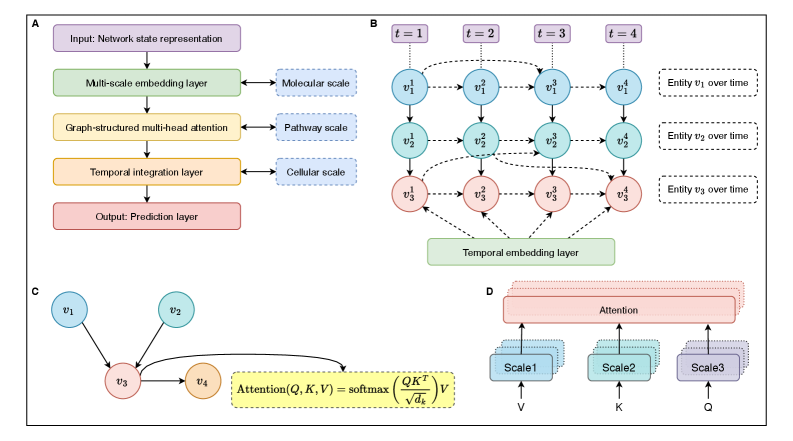
- Methodology Develops HMLMs as a novel computational architecture adapting transformer architecture to model signaling networks as information-processing systems across molecular, pathway, and cellular scales.
- Methodology Implements graph-structured attention mechanisms and hierarchical scale-bridging operators (aggregation, decomposition, translation) to accommodate signaling network topology and multi-scale organization.
主要结论
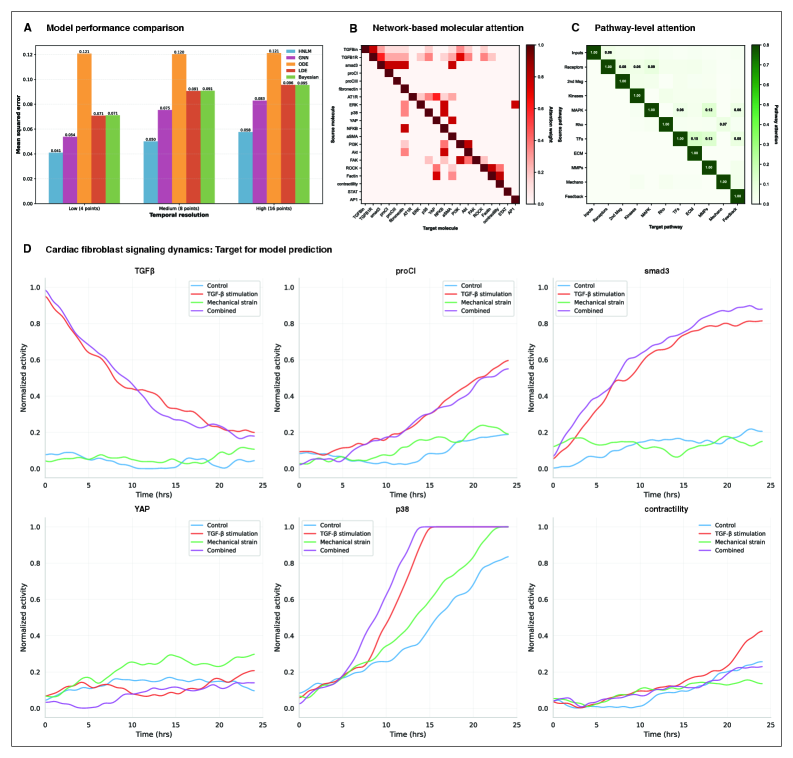
- HMLM achieved MSE of 0.058 for temporal signaling predictions, representing 30% improvement over GNNs (0.083) and 52% improvement over ODE models (0.121).
- Under sparse temporal sampling with only 4 timepoints, HMLM maintained superior performance with MSE = 0.041, demonstrating robustness to limited temporal data.
- Attention mechanisms identified biologically plausible pathway interactions including mechanotransduction-MAPK coupling and TGFβ to ERK signaling, validating the model's ability to capture meaningful biological relationships.
摘要: Cellular signaling networks represent complex information processing systems that have been modeled via traditional mathematical or statistical approaches. However, these methods often struggle to capture context-dependent signaling, pathway cross-talk, and temporal dynamics across multiple biological scales. Here, we introduce hierarchical molecular language models (HMLMs), a novel architecture that proposes a molecular network-specifiac large language model (LLM) to use in intracellular communication as a specialized molecular language, which includes molecules as tokens, protein interactions, post-translational modifications, and regulatory events modeled as semantic relationships within an adapted transformer architecture. HMLMs employ graph-structured attention mechanisms to accommodate signaling network topology while integrating information across the molecular, pathway, and cellular scales through hierarchical attention patterns. We demonstrate HMLM superiority using a cardiac fibroblast signaling network comprising over 100 molecular species across functional modules connected by regulatory edges. HMLM achieved a mean squared error (MSE) of 0.058 for temporal signaling predictions, representing 30% improvement over graph neural networks (GNNs: 0.083) and 52% improvement over ordinary differential equation models (ODEs: 0.121), with particular advantages under sparse temporal sampling conditions where HMLM maintained MSE = 0.041 with only 4 timepoints. The attention-based computational analysis identified key inter-pathway cross-talk patterns through learned attention mechanisms, including mechanotransduction-MAPK coupling and TGFβ to ERK signaling, demonstrating the model's capability to capture biologically plausible pathway interactions from network topology and temporal dynamics and convergent regulatory mechanisms controlling fibrosis markers in simulated cardiac fibroblast networks. The HMLMs offer a foundation for AI-driven biology and medicine with predictable scaling characteristics suitable for interactive applications. By bridging molecular mechanisms with cellular phenotypes through AI-driven molecular language representation, HMLMs provide a powerful paradigm for systems biology that advances precision medicine applications and therapeutic discovery in the era of AI.