Paper List
-
MCP-AI: Protocol-Driven Intelligence Framework for Autonomous Reasoning in Healthcare
This paper addresses the critical gap in healthcare AI systems that lack contextual reasoning, long-term state management, and verifiable workflows by...
-
Model Gateway: Model Management Platform for Model-Driven Drug Discovery
This paper addresses the critical bottleneck of fragmented, ad-hoc model management in pharmaceutical research by providing a centralized, scalable ML...
-
Tree Thinking in the Genomic Era: Unifying Models Across Cells, Populations, and Species
This paper addresses the fragmentation of tree-based inference methods across biological scales by identifying shared algorithmic principles and stati...
-
SSDLabeler: Realistic semi-synthetic data generation for multi-label artifact classification in EEG
This paper addresses the core challenge of training robust multi-label EEG artifact classifiers by overcoming the scarcity and limited diversity of ma...
-
Decoding Selective Auditory Attention to Musical Elements in Ecologically Valid Music Listening
This paper addresses the core challenge of objectively quantifying listeners' selective attention to specific musical components (e.g., vocals, drums,...
-
Physics-Guided Surrogate Modeling for Machine Learning–Driven DLD Design Optimization
This paper addresses the core bottleneck of translating microfluidic DLD devices from research prototypes to clinical applications by replacing weeks-...
-
Mechanistic Interpretability of Antibody Language Models Using SAEs
This work addresses the core challenge of achieving both interpretability and controllable generation in domain-specific protein language models, spec...
-
Fluctuating Environments Favor Extreme Dormancy Strategies and Penalize Intermediate Ones
This paper addresses the core challenge of determining how organisms should tune dormancy duration to match the temporal autocorrelation of their envi...
DeepFRI Demystified: Interpretability vs. Accuracy in AI Protein Function Prediction
Yale University | Microsoft
30秒速读
IN SHORT: This study addresses the critical gap between high predictive accuracy and biological interpretability in DeepFRI, revealing that the model often prioritizes structural motifs over functional residues, complicating reliable identification of drug targets.
核心创新
- Methodology Comprehensive benchmarking of three post-hoc explainability methods (GradCAM, Excitation Backpropagation, PGExplainer) on DeepFRI with quantitative sparsity analysis.
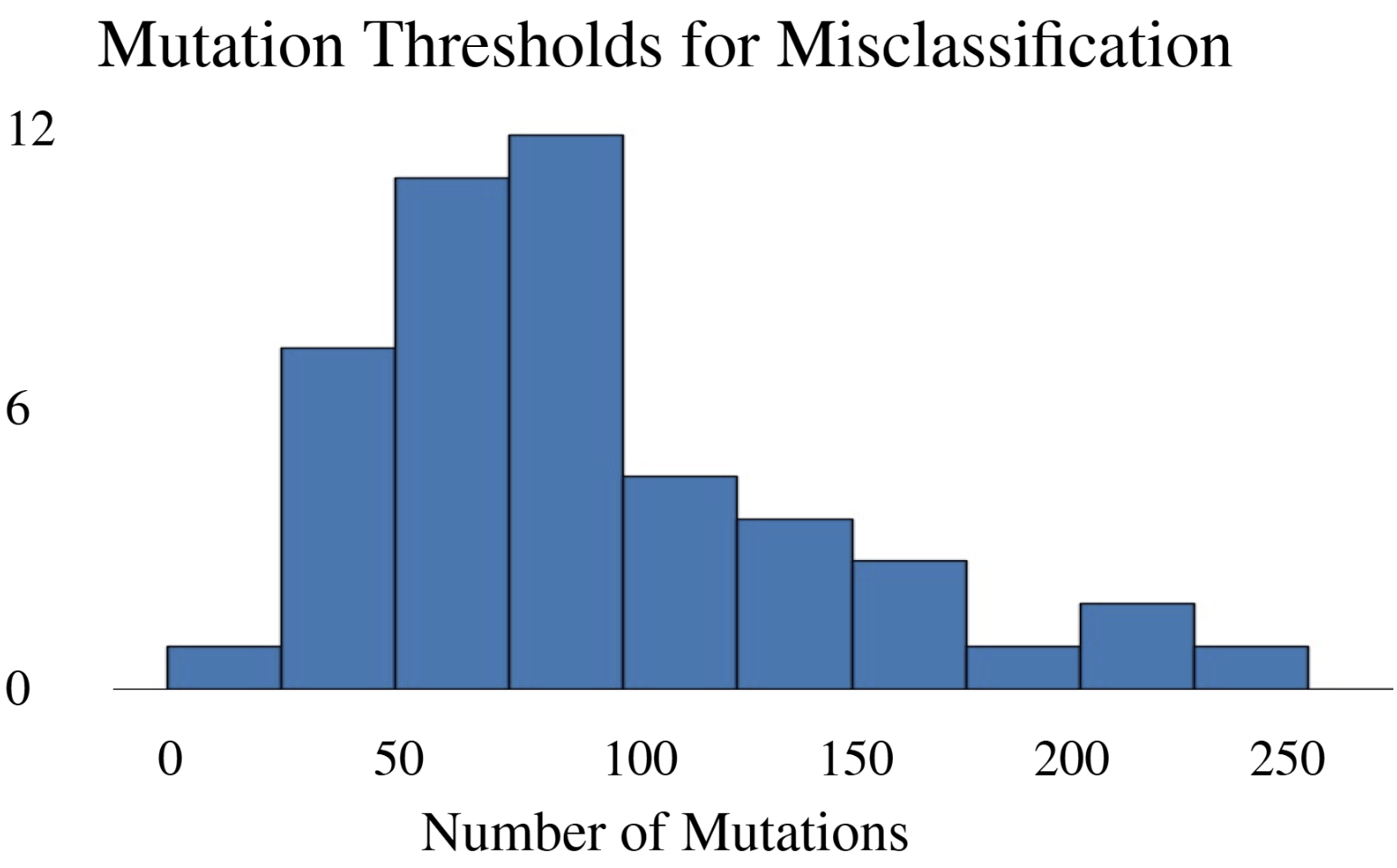
- Methodology Development of a modified DeepFool adversarial testing framework for protein sequences, measuring mutation thresholds required for misclassification.
- Biology Revealed that DeepFRI prioritizes amino acids controlling protein structure over function in >50% of tested proteins, highlighting a fundamental accuracy-interpretability trade-off.
主要结论
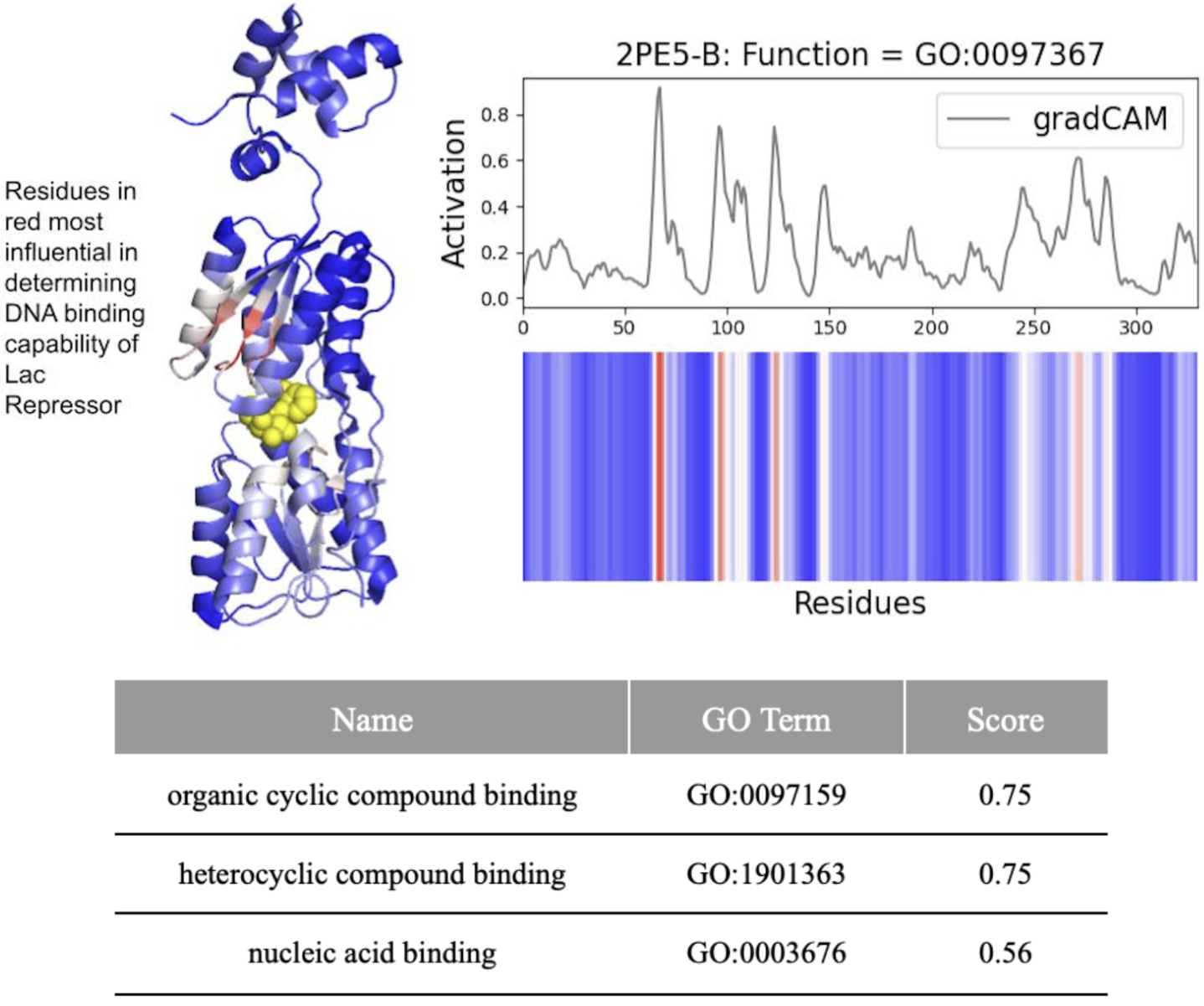
- DeepFRI required 206 mutations (62.4% of 330 residues) in the lac repressor for misclassification, demonstrating extreme robustness but potentially missing subtle functional alterations.
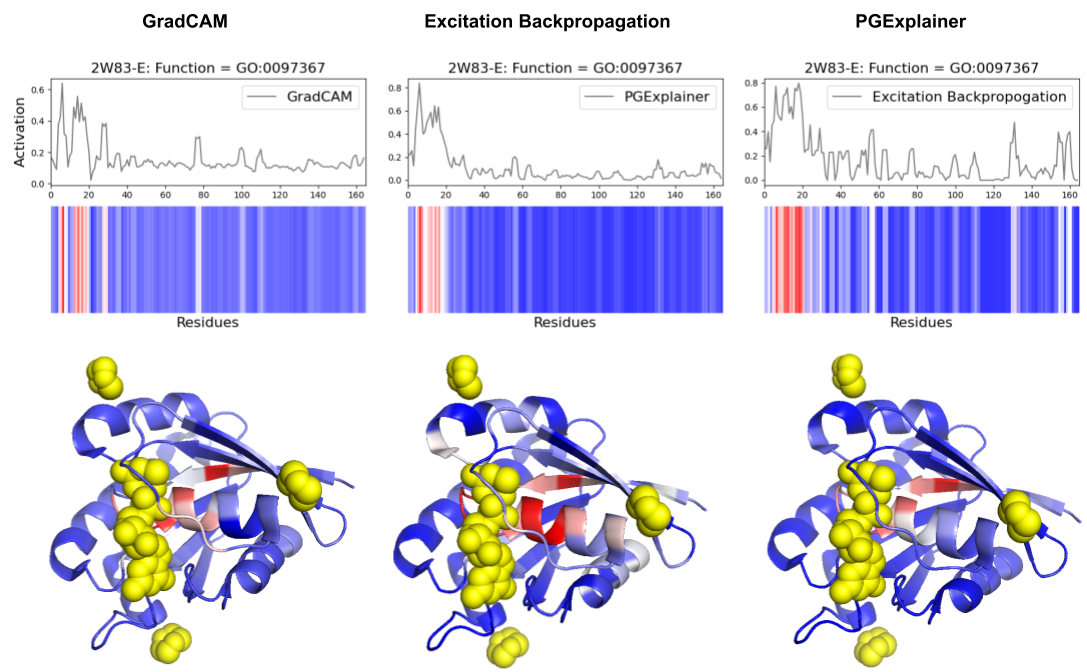
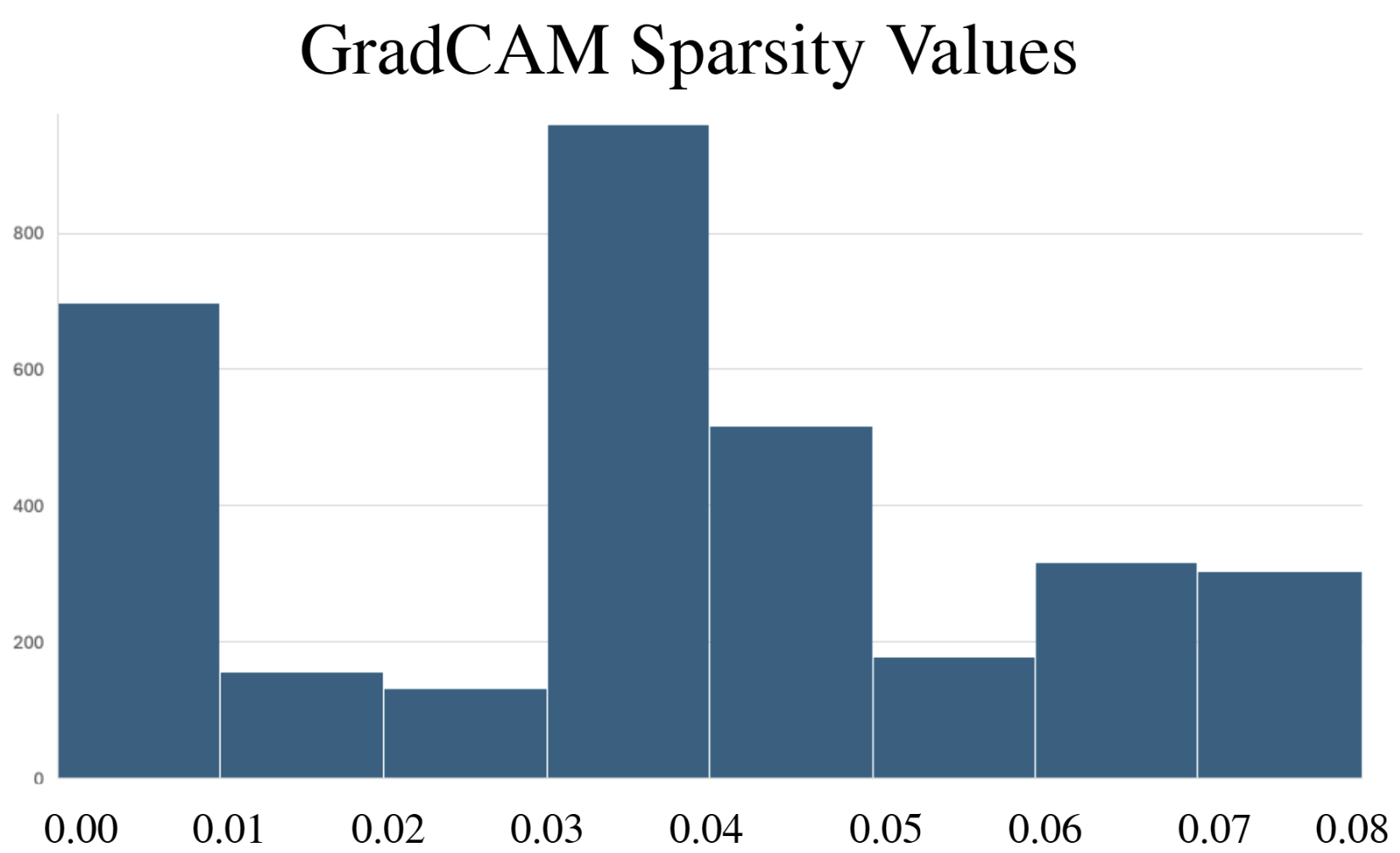
- Explainability methods showed significant granularity differences: PGExplainer was 3× sparser than GradCAM and 17× sparser than Excitation Backpropagation across 124 binding proteins.
- All three methods converged on biochemically critical P-loop residues (0-20) in ARF6 GTPase, validating DeepFRI's focus on conserved functional motifs in straightforward domains.
摘要: Machine learning technologies for protein function prediction are black box models. Despite their potential to identify key drug targets with high accuracy and accelerate therapy development, the adoption of these methods depends on verifying their findings. This study evaluates DeepFRI, a leading Graph Convolutional Network (GCN)-based tool, using advanced explainability techniques—GradCAM, Excitation Backpropagation, and PGExplainer—and adversarial robustness tests. Our findings reveal that the model’s predictions often prioritize conserved motifs over truly deterministic residues, complicating the identification of functional sites. Quantitative analyses show that explainability methods differ significantly in granularity, with GradCAM providing broad relevance and PGExplainer pinpointing specific active sites. These results highlight trade-offs between accuracy and interpretability, suggesting areas for improvement in DeepFRI’s architecture to enhance its trustworthiness in drug discovery and regulatory settings.