Paper List
-
STAR-GO: Improving Protein Function Prediction by Learning to Hierarchically Integrate Ontology-Informed Semantic Embeddings
This paper addresses the core challenge of generalizing protein function prediction to unseen or newly introduced Gene Ontology (GO) terms by overcomi...
-
Incorporating indel channels into average-case analysis of seed-chain-extend
This paper addresses the core pain point of bridging the theoretical gap for the widely used seed-chain-extend heuristic by providing the first rigoro...
-
Competition, stability, and functionality in excitatory-inhibitory neural circuits
This paper addresses the core challenge of extending interpretable energy-based frameworks to biologically realistic asymmetric neural networks, where...
-
Enhancing Clinical Note Generation with ICD-10, Clinical Ontology Knowledge Graphs, and Chain-of-Thought Prompting Using GPT-4
This paper addresses the core challenge of generating accurate and clinically relevant patient notes from sparse inputs (ICD codes and basic demograph...
-
Learning From Limited Data and Feedback for Cell Culture Process Monitoring: A Comparative Study
This paper addresses the core challenge of developing accurate real-time bioprocess monitoring soft sensors under severe data constraints: limited his...
-
Cell-cell communication inference and analysis: biological mechanisms, computational approaches, and future opportunities
This review addresses the critical need for a systematic framework to navigate the rapidly expanding landscape of computational methods for inferring ...
-
Generating a Contact Matrix for Aged Care Settings in Australia: an agent-based model study
This study addresses the critical gap in understanding heterogeneous contact patterns within aged care facilities, where existing population-level con...
-
Emergent Spatiotemporal Dynamics in Large-Scale Brain Networks with Next Generation Neural Mass Models
This work addresses the core challenge of understanding how complex, brain-wide spatiotemporal patterns emerge from the interaction of biophysically d...
The Effective Reproduction Number in the Kermack-McKendrick model with age of infection and reinfection
School of Mathematical Sciences, Beijing Normal University, Beijing 100875, People’s Republic of China.
30秒速读
IN SHORT: This paper addresses the challenge of accurately estimating the time-varying effective reproduction number ℛ(t) in epidemics by incorporating two critical real-world complexities: the age of infection (time since infection) and the possibility of reinfection.
核心创新
- Methodology Introduces a novel extension of the classical Kermack-McKendrick SIRS model by formally incorporating both infection-age structure (a) and a reinfection term (δ), moving beyond constant transmission rate assumptions.
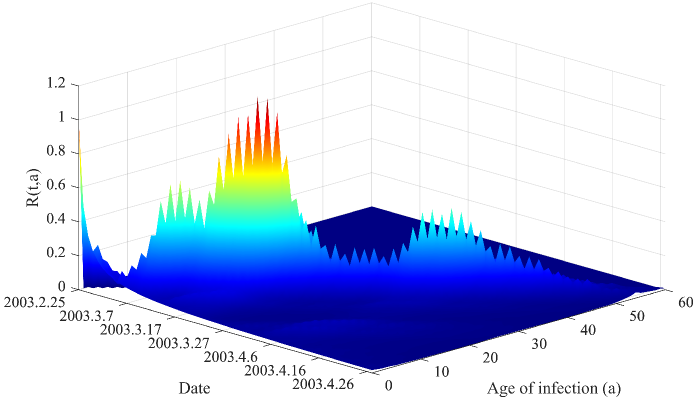
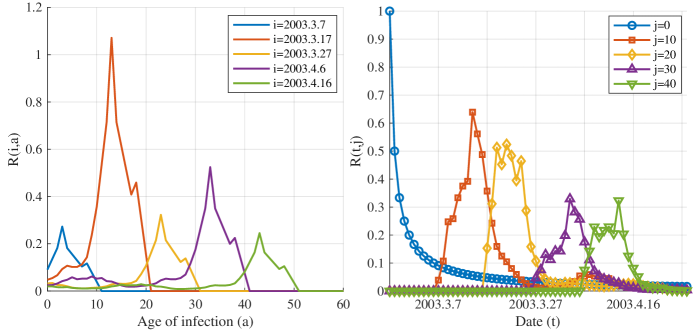
- Methodology Derives a rigorous mathematical framework using Volterra integral equations, the contraction mapping principle, and measure-valued solutions (e.g., Dirac mass for initial cohorts) to connect the flow of new infections N(t) to the reproductive power ℛ(t,a) and ultimately ℛ(t).
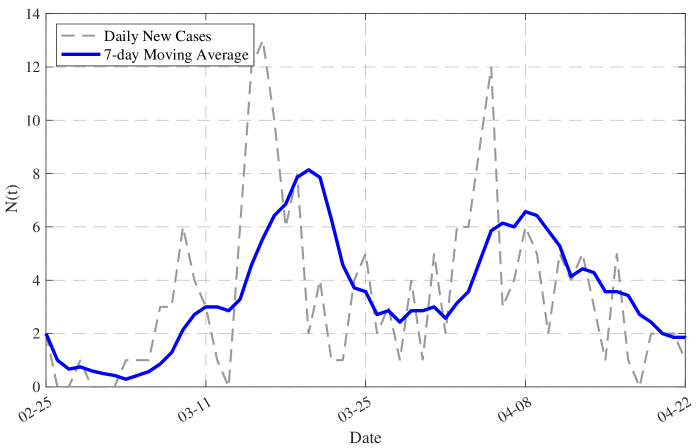
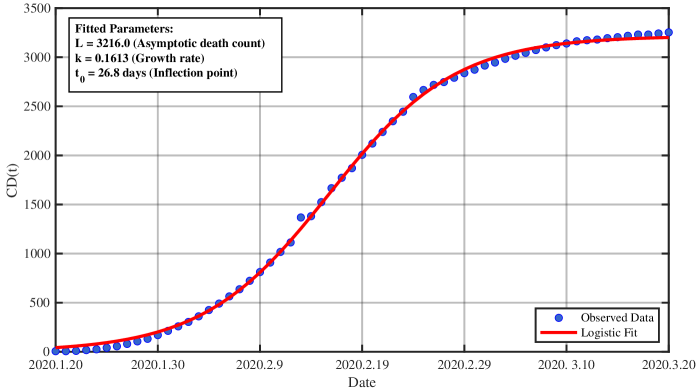
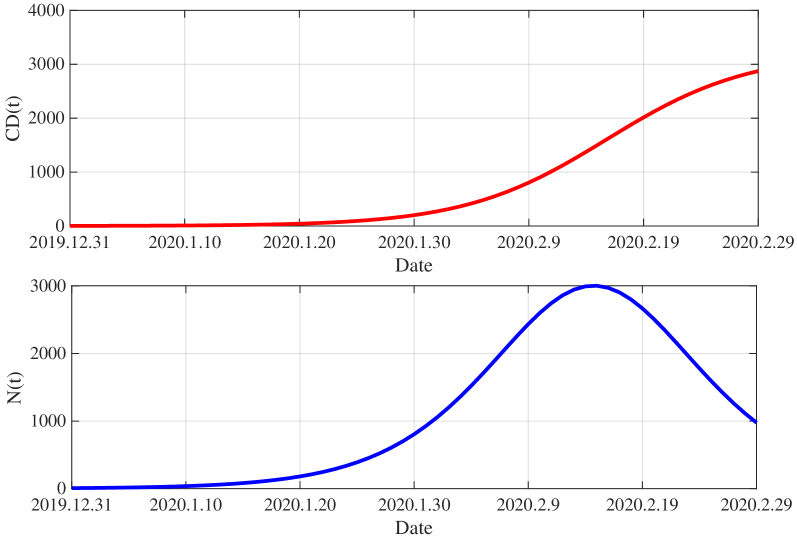
- Methodology/Biology Develops a practical parameter identification methodology that works with two common but challenging data types: 1) direct daily new case counts (applied to 2003 SARS in Singapore) and 2) cumulative death counts when new infection data is unreliable (applied to COVID-19 in China).
主要结论
- The model successfully formulates the infection dynamics as a nonlinear Volterra integral equation of the second kind for N(t) (Eq. 2.14), providing a solvable link between observable data and the underlying transmission parameters.
- Theoretical analysis justifies the use of a Dirac mass initial condition (representing a single cohort infected at time t0) via a limiting process of approximating functions i_κ(a), proving uniform convergence of the solution N_κ(t) to N(t) (Theorem 3.2).
- The derived framework enables the identification of the effective reproduction number ℛ(t) from epidemic curves, demonstrated through application to real-world SARS and COVID-19 datasets, bridging theoretical constructs with practical public health analytics.
摘要: This study introduces a novel epidemiological model that expands upon the Kermack-McKendrick model by incorporating the age of infection and reinfection. By including infection age, we can classify participants, which enables a more targeted analysis within the modeling framework. The reinfection term addresses the real-world occurrences of secondary or recurrent viral infections. In the theoretical part, we apply the contraction mapping principle, the dominated convergence theorem, and the properties of Volterra integral equations to derive analytical expressions for the number of newly infected individuals denoted by N(t). Then, we establish a Volterra integral equation for N(t) and study its initial conditions for both a single cohort and multiple cohorts. From this equation, we derive a method for identifying the effective reproduction number, denoted as ℛ(t). In the practical aspect, we present two distinct methods and separately apply them to analyze the daily new infection cases from the 2003 SARS outbreak in Singapore and the cumulative number of deaths from the COVID-19 epidemic in China. This work effectively bridges theoretical epidemiology and computational modeling, providing a robust framework for analyzing infection dynamics influenced by infection-age-structured transmission and reinfection mechanisms.