Paper List
-
Evolutionarily Stable Stackelberg Equilibrium
通过要求追随者策略对突变入侵具有鲁棒性,弥合了斯塔克尔伯格领导力模型与演化稳定性之间的鸿沟。
-
Recovering Sparse Neural Connectivity from Partial Measurements: A Covariance-Based Approach with Granger-Causality Refinement
通过跨多个实验会话累积协方差统计,实现从部分记录到完整神经连接性的重建。
-
Atomic Trajectory Modeling with State Space Models for Biomolecular Dynamics
ATMOS通过提供一个基于SSM的高效框架,用于生物分子的原子级轨迹生成,弥合了计算昂贵的MD模拟与时间受限的深度生成模型之间的差距。
-
Slow evolution towards generalism in a model of variable dietary range
通过证明是种群统计噪声(而非确定性动力学)驱动了模式形成和泛化食性的演化,解决了间接竞争下物种形成的悖论。
-
Grounded Multimodal Retrieval-Augmented Drafting of Radiology Impressions Using Case-Based Similarity Search
通过将印象草稿基于检索到的历史病例,并采用明确引用和基于置信度的拒绝机制,解决放射学报告生成中的幻觉问题。
-
Unified Policy–Value Decomposition for Rapid Adaptation
通过双线性分解在策略和价值函数之间共享低维目标嵌入,实现对新颖任务的零样本适应。
-
Mathematical Modeling of Cancer–Bacterial Therapy: Analysis and Numerical Simulation via Physics-Informed Neural Networks
提供了一个严格的、无网格的PINN框架,用于模拟和分析细菌癌症疗法中复杂的、空间异质的相互作用。
-
Sample-Efficient Adaptation of Drug-Response Models to Patient Tumors under Strong Biological Domain Shift
通过从无标记分子谱中学习可迁移表征,利用最少的临床数据实现患者药物反应的有效预测。
DeeDeeExperiment: Building an infrastructure for integrating and managing omics data analysis results in R/Bioconductor
Institute of Medical Biostatistics, Epidemiology and Informatics (IMBEI), University Medical Center Mainz | Research Center for Immunotherapy (FZI) Mainz | Department of Nephrology, Rheumatology and Kidney Transplantation, University Medical Center Mainz
30秒速读
IN SHORT: This paper addresses the critical bottleneck of managing and organizing the growing volume of differential expression and functional enrichment analysis results from complex omics experiments, which currently lack standardized data structures for storage and contextualization.
核心创新
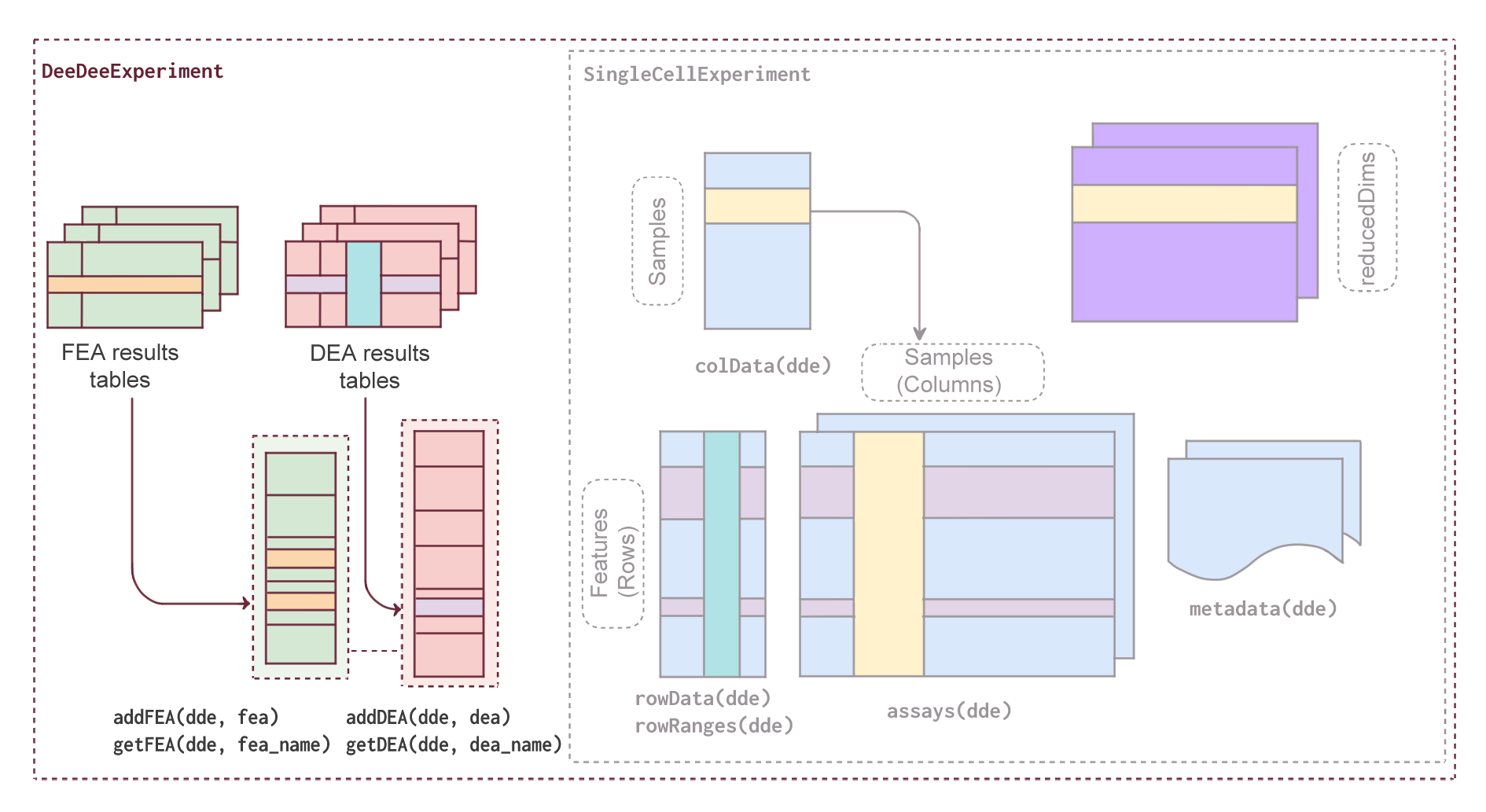
- Methodology Introduces the first standardized S4 class specifically designed to co-store DEA and FEA results with their metadata in a single, structured container within the Bioconductor ecosystem.
- Methodology Extends the widely adopted SingleCellExperiment class by adding dedicated slots for DEA and FEA results while maintaining full backward compatibility with existing Bioconductor tools.
- Methodology Implements a contrast-centric architecture that organizes results from multiple comparisons (including limma multi-contrast objects and muscat pseudobulk analyses) with efficient storage through pointer-based referencing.
主要结论
- DeeDeeExperiment provides a robust, standardized framework that enables efficient organization and retrieval of DEA/FEA results across multiple contrasts within a single data object.
- The implementation maintains full compatibility with the Bioconductor ecosystem, supporting interoperability with downstream tools like scater for visualization and iSEE for interactive exploration.
- By consolidating analysis results and metadata, the framework supports more nuanced quantitative approaches beyond simple overlap strategies, enabling trustworthy summaries of complex experimental measurements.
摘要: Summary: Modern omics experiments now involve multiple conditions and complex designs, producing an increasingly large set of differential expression and functional enrichment analysis results. However, no standardized data structure exists to store and contextualize these results together with their metadata, leaving researchers with an unmanageable and potentially non-reproducible collection of results that are difficult to navigate and/or share. Here we introduce DeeDeeExperiment, a new S4 class for managing and storing omics data analysis results, implemented within the Bioconductor ecosystem, which promotes interoperability, reproducibility and good documentation. This class extends the widely used SingleCellExperiment object by introducing dedicated slots for Differential Expression (DEA) and Functional Enrichment Analysis (FEA) results, allowing users to organize, store, and retrieve information on multiple contrasts and associated metadata within a single data object, ultimately streamlining the management and interpretation of many omics datasets. Availability and implementation: DeeDeeExperiment is available on Bioconductor under the MIT license (https://bioconductor.org/packages/DeeDeeExperiment), with its development version also available on Github (https://github.com/imbeimainz/DeeDeeExperiment).