Paper List
-
Autonomous Agents Coordinating Distributed Discovery Through Emergent Artifact Exchange
This paper addresses the fundamental limitation of current AI-assisted scientific research by enabling truly autonomous, decentralized investigation w...
-
D-MEM: Dopamine-Gated Agentic Memory via Reward Prediction Error Routing
This paper addresses the fundamental scalability bottleneck in LLM agentic memory systems: the O(N²) computational complexity and unbounded API token ...
-
Countershading coloration in blue shark skin emerges from hierarchically organized and spatially tuned photonic architectures inside skin denticles
This paper solves the core problem of how blue sharks achieve their striking dorsoventral countershading camouflage, revealing that coloration origina...
-
Human-like Object Grouping in Self-supervised Vision Transformers
This paper addresses the core challenge of quantifying how well self-supervised vision models capture human-like object grouping in natural scenes, br...
-
Hierarchical pp-Adic Framework for Gene Regulatory Networks: Theory and Stability Analysis
This paper addresses the core challenge of mathematically capturing the inherent hierarchical organization and multi-scale stability of gene regulator...
-
Towards unified brain-to-text decoding across speech production and perception
This paper addresses the core challenge of developing a unified brain-to-text decoding framework that works across both speech production and percepti...
-
Dual-Laws Model for a theory of artificial consciousness
This paper addresses the core challenge of developing a comprehensive, testable theory of consciousness that bridges biological and artificial systems...
-
Pulse desynchronization of neural populations by targeting the centroid of the limit cycle in phase space
This work addresses the core challenge of determining optimal pulse timing and intensity for desynchronizing pathological neural oscillations when the...
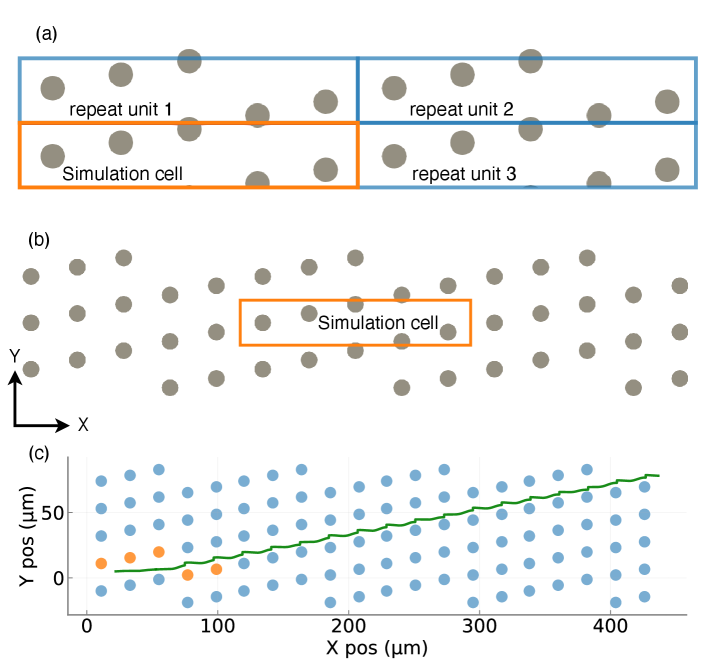
Physics-Guided Surrogate Modeling for Machine Learning–Driven DLD Design Optimization
Department of Mechanical Engineering, Lehigh University | Computational Engineering Department, Lawrence Livermore National Laboratory | Department of Industrial and Production Engineering, Bangladesh University of Engineering and Technology | Precision Medicine Translational Research Center, West China Hospital, Sichuan University
30秒速读
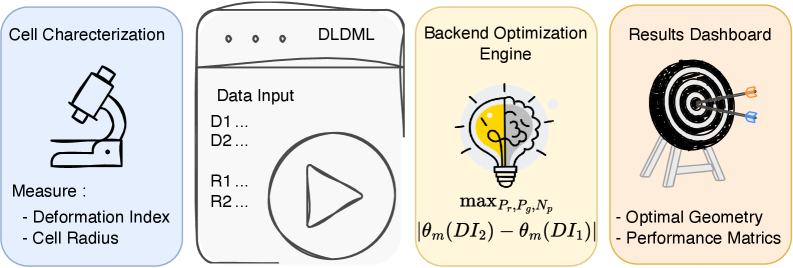
IN SHORT: This paper addresses the core bottleneck of translating microfluidic DLD devices from research prototypes to clinical applications by replacing weeks-long empirical design cycles with a physics-guided machine learning framework that delivers fabrication-ready specifications in under 60 seconds.
核心创新
- Methodology First complete inverse design framework for DLD that transforms measured cellular deformability into optimized device geometry through physics-guided machine learning.
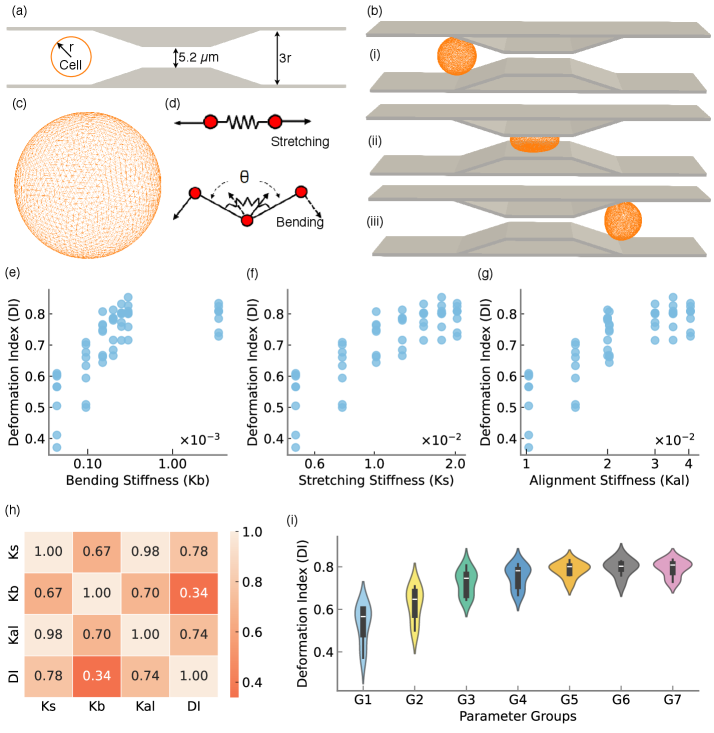
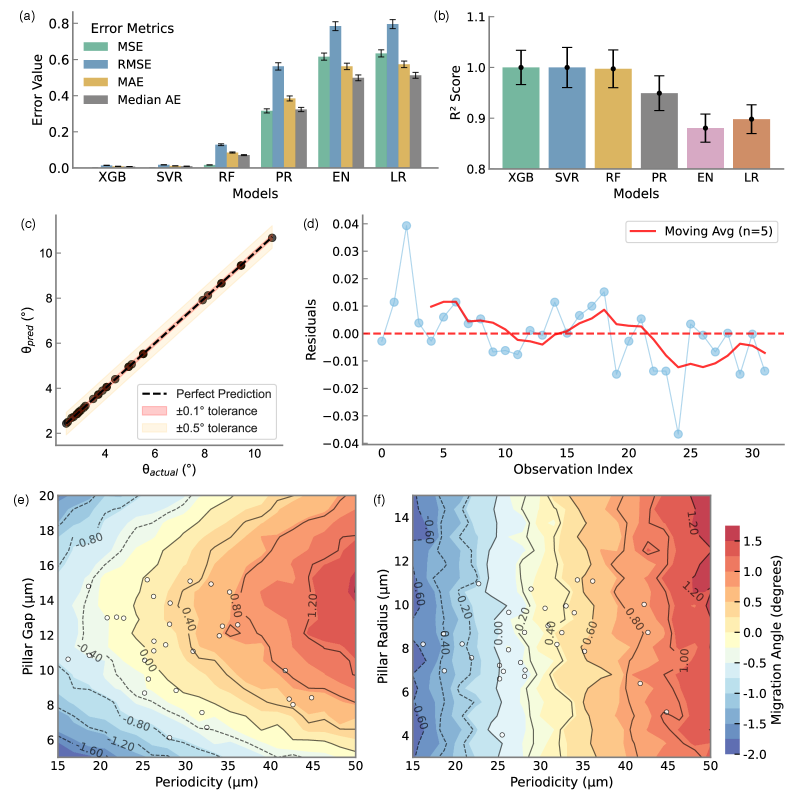
- Methodology Integration of high-fidelity Lattice-Boltzmann/Immersed-Boundary simulations with XGBoost surrogate models achieving sub-degree predictive accuracy (R²=0.9999, MSE=2×10⁻⁴).
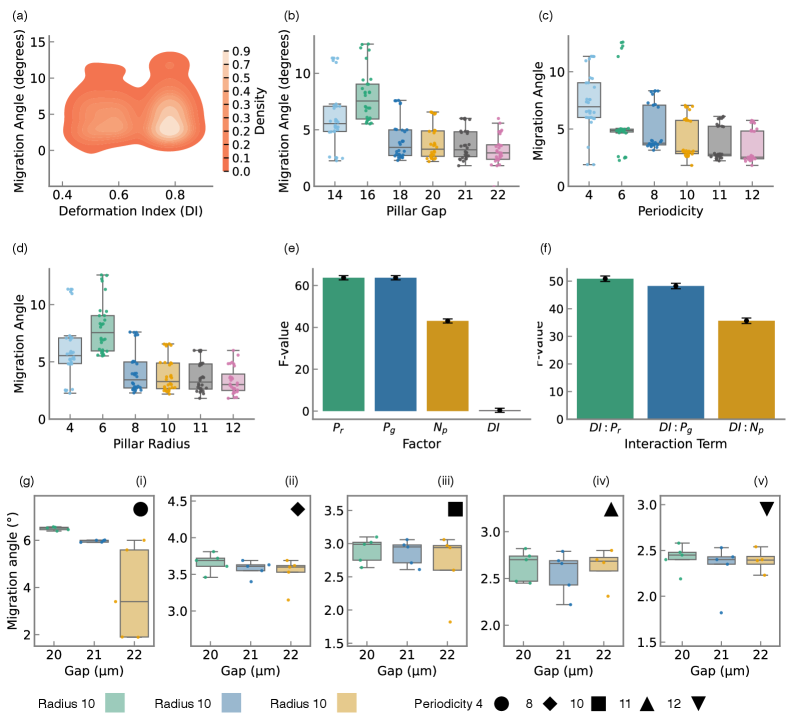
- Methodology Statistical quantification of deformability-geometry interactions via Type II ANOVA revealing significant interaction effects (F=48.23, p<10⁻³⁴) despite geometric dominance of main effects.
主要结论
- Geometric parameters dominate migration angle variance (F=63.72, p<10⁻³⁷), but cellular deformability exerts statistically significant effects through interactions with device geometry (F=48.23, p<10⁻³⁴).
- The XGBoost surrogate model achieves exceptional predictive accuracy (R²=0.9999, MSE=2×10⁻⁴), enabling sub-degree migration angle prediction across the design space.
- Bayesian optimization via tree-structured Parzen estimation identifies optimal DLD architectures in under 60 seconds, reducing design iteration from weeks of experimental prototyping to minutes of automated computation.
摘要: Microfluidic separation technologies have transformed label-free cell sorting by exploiting intrinsic biophysical properties, yet the translation of these platforms from laboratory prototypes to clinical applications remains constrained by the empirical, trial-and-error nature of device design. Deterministic Lateral Displacement (DLD) represents a paradigmatic example: while demonstrating robust discrimination of cells by size, shape, and deformability across diverse applications including circulating tumor cell isolation and malaria diagnostics, DLD performance exhibits extreme sensitivity to the coupled interplay between cellular mechanical phenotype and micron-scale geometric parameters, necessitating iterative fabrication-testing cycles that span weeks to months. We present the first complete inverse design framework that transforms measured cellular deformability into fabrication-ready DLD specifications through physics-guided machine learning. Our approach integrates high-fidelity lattice-Boltzmann and immersed-boundary simulations with gradient-boosted surrogate models to systematically map cellular mechanical properties to migration behavior across manufacturing-feasible geometric configurations (pillar radius, gap, periodicity). Type II ANOVA quantifies the relative influence of these parameters, revealing that while geometric factors dominate migration angle variance (F=63.72, p<10−37), cellular deformability exerts statistically significant effects through interactions with device geometry (F=48.23, p<10−34). The resulting XGBoost surrogate achieves sub-degree predictive accuracy (R2=0.9999, MSE =2×10−4), enabling Bayesian optimization via tree-structured Parzen estimation to identify optimal array architectures in under 60 seconds—reducing design iteration from weeks of experimental prototyping to minutes of automated computation. By deploying this validated pipeline as an accessible web application that accepts experimentally measured deformation indices and returns optimized device specifications with tolerance analysis, we democratize DLD design for researchers without specialized computational expertise, thereby accelerating the translation of microfluidic technologies from research-grade prototypes to application-specific, clinically deployable devices.