Paper List
-
Ill-Conditioning in Dictionary-Based Dynamic-Equation Learning: A Systems Biology Case Study
This paper addresses the critical challenge of numerical ill-conditioning and multicollinearity in library-based sparse regression methods (e.g., SIND...
-
Hybrid eTFCE–GRF: Exact Cluster-Size Retrieval with Analytical pp-Values for Voxel-Based Morphometry
This paper addresses the computational bottleneck in voxel-based neuroimaging analysis by providing a method that delivers exact cluster-size retrieva...
-
abx_amr_simulator: A simulation environment for antibiotic prescribing policy optimization under antimicrobial resistance
This paper addresses the critical challenge of quantitatively evaluating antibiotic prescribing policies under realistic uncertainty and partial obser...
-
PesTwin: a biology-informed Digital Twin for enabling precision farming
This paper addresses the critical bottleneck in precision agriculture: the inability to accurately forecast pest outbreaks in real-time, leading to su...
-
Equivariant Asynchronous Diffusion: An Adaptive Denoising Schedule for Accelerated Molecular Conformation Generation
This paper addresses the core challenge of generating physically plausible 3D molecular structures by bridging the gap between autoregressive methods ...
-
Omics Data Discovery Agents
This paper addresses the core challenge of making published omics data computationally reusable by automating the extraction, quantification, and inte...
-
Single-cell directional sensing at ultra-low chemoattractant concentrations from extreme first-passage events
This work addresses the core challenge of how a cell can rapidly and accurately determine the direction of a chemoattractant source when the signal is...
-
SDSR: A Spectral Divide-and-Conquer Approach for Species Tree Reconstruction
This paper addresses the computational bottleneck in reconstructing species trees from thousands of species and multiple genes by introducing a scalab...
Incorporating indel channels into average-case analysis of seed-chain-extend
Carnegie Mellon University, Pittsburgh, PA, USA
30秒速读
IN SHORT: This paper addresses the core pain point of bridging the theoretical gap for the widely used seed-chain-extend heuristic by providing the first rigorous average-case analysis that accounts for insertions and deletions (indels), not just substitutions.
核心创新
- Methodology Introduces a generalized definition of 'recoverability' and a 'homologous path' to mathematically model the correct alignment under indel mutation channels, moving beyond the simpler 'homologous diagonal' used for substitutions only.
- Theory Develops new mathematical machinery to handle the dependence structure of neighboring anchors and the existence of 'clipping anchors' (partially correct anchors), which are unique challenges introduced by indels.
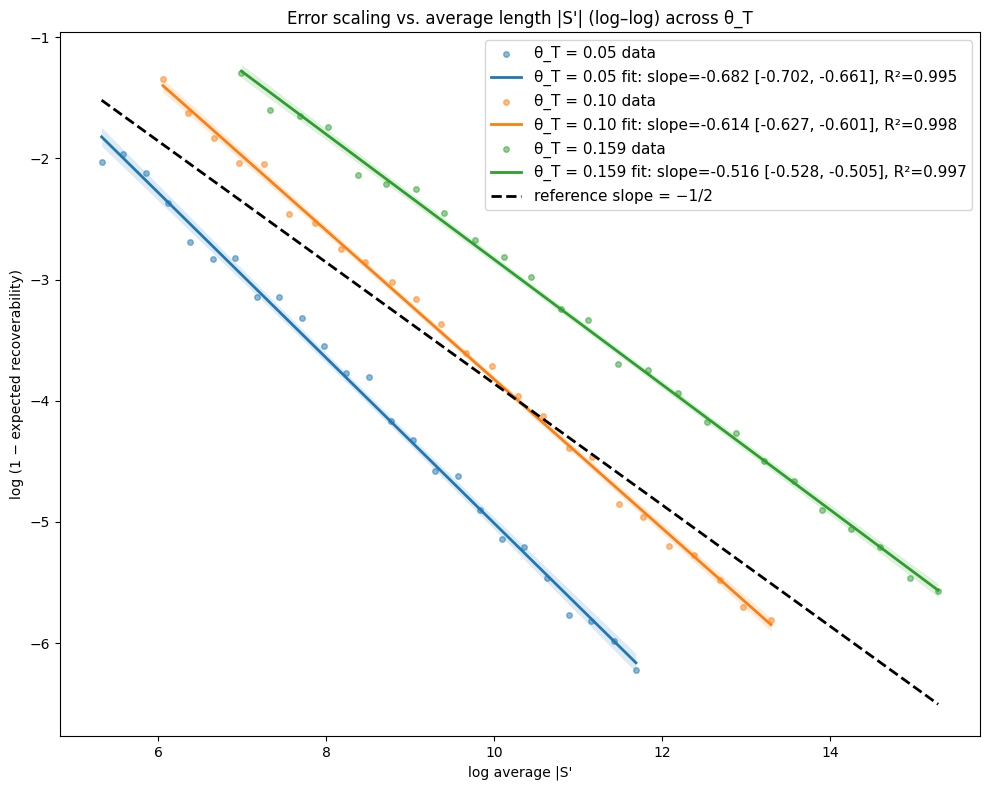
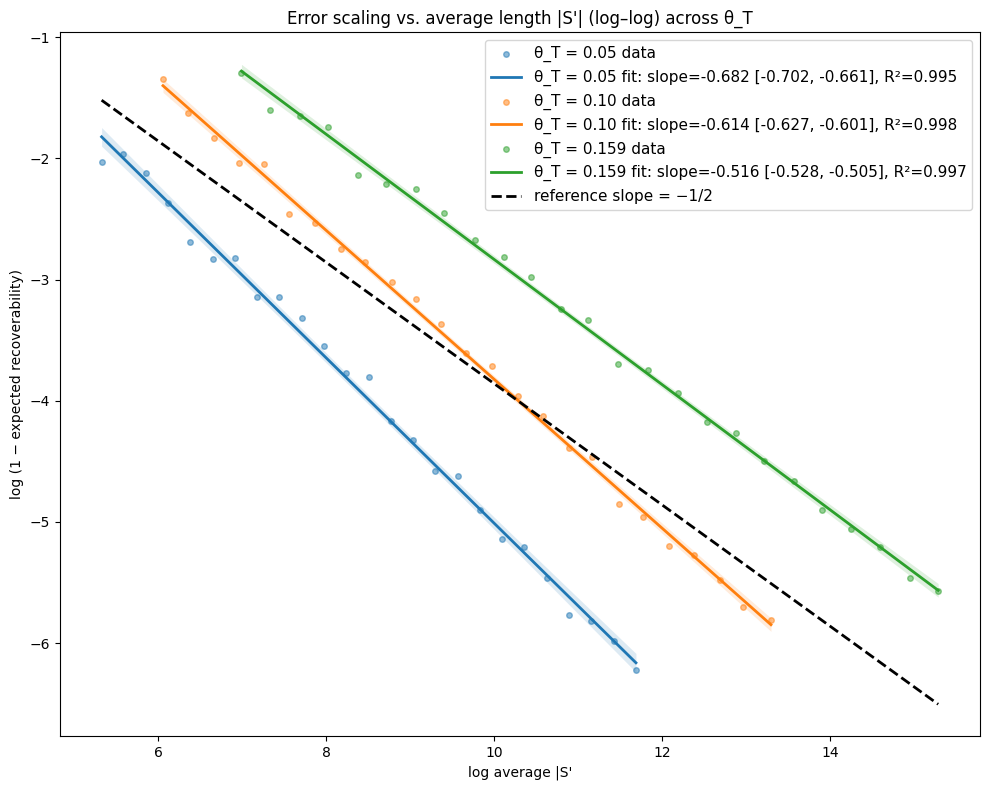
- Theory Proves that under a total mutation rate θ_T < 0.159, optimal linear-gap cost chaining achieves an expected recoverability of ≥ 1 - O(1/√m), generalizing the prior substitution-only result to a biologically realistic model.
主要结论
- The expected recoverability of an optimal chain under linear-gap cost chaining is ≥ 1 - O(1/√m) when the total mutation rate θ_T (sum of substitution, insertion, deletion rates) is less than 0.159.
- The expected runtime of the algorithm is O(m n^(3.15·θ_T) log n). For example, at a θ_T of 0.05 (similar to human-chimp divergence), the exponent is ~1.12, leading to near-linear scaling.
- The analysis successfully bridges theory and practice by extending the proof framework to handle indels, justifying the heuristic's empirical effectiveness on real genomic data which contains indels.
摘要: Given a sequence s1 of n letters drawn i.i.d. from an alphabet of size σ and a mutated substring s2 of length m<n, we often want to recover the mutation history that generated s2 from s1. Modern sequence aligners are widely used for this task, and many employ the seed-chain-extend heuristic with k-mer seeds. Previously, Shaw and Yu showed that optimal linear-gap cost chaining can produce a chain with 1−O(1/m) recoverability, the proportion of the mutation history that is recovered, in O(mn^(2.43θ) log n) expected time, where θ<0.206 is the mutation rate under a substitution-only channel and s1 is assumed to be uniformly random. However, a gap remains between theory and practice, since real genomic data includes insertions and deletions (indels), and yet seed-chain-extend remains effective. In this paper, we generalize those prior results by introducing mathematical machinery to deal with the two new obstacles introduced by indel channels: the dependence of neighboring anchors and the presence of anchors that are only partially correct. We are thus able to prove that the expected recoverability of an optimal chain is ≥1−O(1/√m) and the expected runtime is O(mn^(3.15·θ_T) log n), when the total mutation rate given by the sum of the substitution, insertion, and deletion mutation rates (θ_T = θ_i + θ_d + θ_s) is less than 0.159.