Paper List
-
MCP-AI: Protocol-Driven Intelligence Framework for Autonomous Reasoning in Healthcare
This paper addresses the critical gap in healthcare AI systems that lack contextual reasoning, long-term state management, and verifiable workflows by...
-
Model Gateway: Model Management Platform for Model-Driven Drug Discovery
This paper addresses the critical bottleneck of fragmented, ad-hoc model management in pharmaceutical research by providing a centralized, scalable ML...
-
Tree Thinking in the Genomic Era: Unifying Models Across Cells, Populations, and Species
This paper addresses the fragmentation of tree-based inference methods across biological scales by identifying shared algorithmic principles and stati...
-
SSDLabeler: Realistic semi-synthetic data generation for multi-label artifact classification in EEG
This paper addresses the core challenge of training robust multi-label EEG artifact classifiers by overcoming the scarcity and limited diversity of ma...
-
Decoding Selective Auditory Attention to Musical Elements in Ecologically Valid Music Listening
This paper addresses the core challenge of objectively quantifying listeners' selective attention to specific musical components (e.g., vocals, drums,...
-
Physics-Guided Surrogate Modeling for Machine Learning–Driven DLD Design Optimization
This paper addresses the core bottleneck of translating microfluidic DLD devices from research prototypes to clinical applications by replacing weeks-...
-
Mechanistic Interpretability of Antibody Language Models Using SAEs
This work addresses the core challenge of achieving both interpretability and controllable generation in domain-specific protein language models, spec...
-
Fluctuating Environments Favor Extreme Dormancy Strategies and Penalize Intermediate Ones
This paper addresses the core challenge of determining how organisms should tune dormancy duration to match the temporal autocorrelation of their envi...
Incorporating indel channels into average-case analysis of seed-chain-extend
Carnegie Mellon University, Pittsburgh, PA, USA
30秒速读
IN SHORT: This paper addresses the core pain point of bridging the theoretical gap for the widely used seed-chain-extend heuristic by providing the first rigorous average-case analysis that accounts for insertions and deletions (indels), not just substitutions.
核心创新
- Methodology Introduces a generalized definition of 'recoverability' and a 'homologous path' to mathematically model the correct alignment under indel mutation channels, moving beyond the simpler 'homologous diagonal' used for substitutions only.
- Theory Develops new mathematical machinery to handle the dependence structure of neighboring anchors and the existence of 'clipping anchors' (partially correct anchors), which are unique challenges introduced by indels.
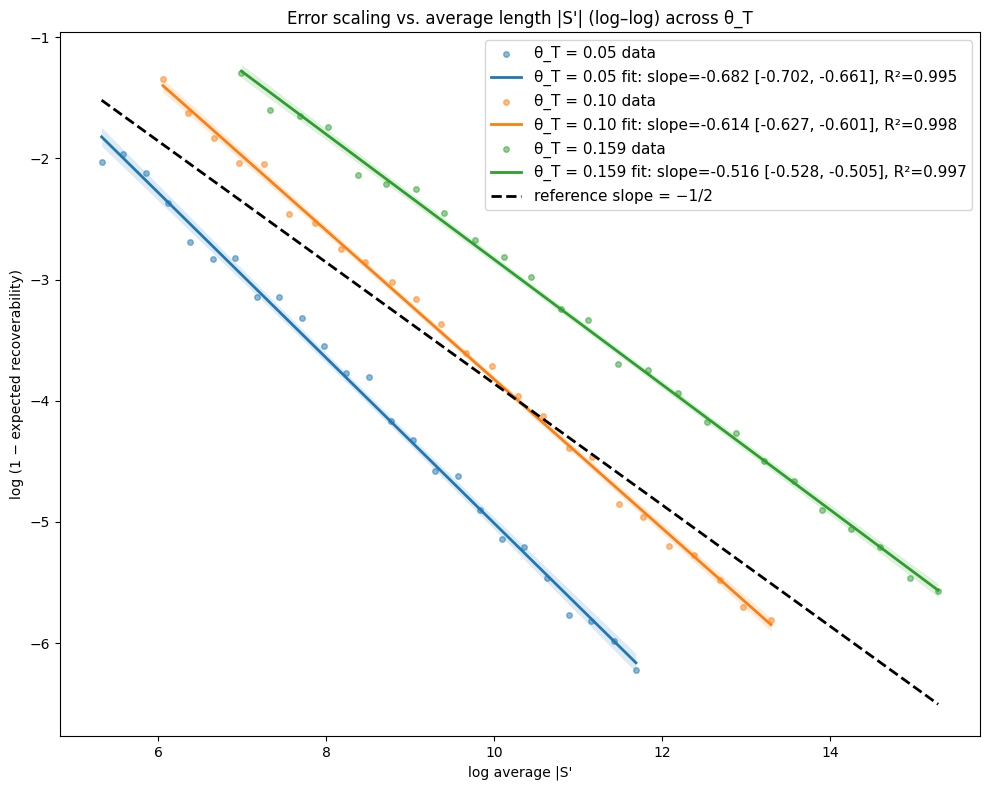
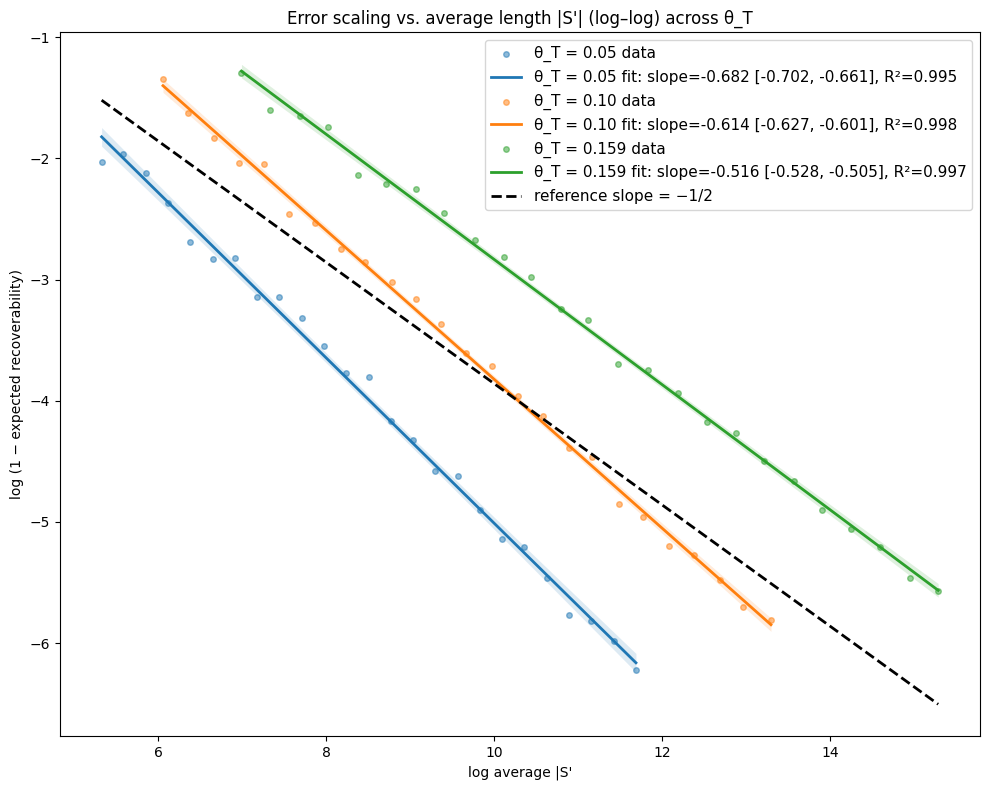
- Theory Proves that under a total mutation rate θ_T < 0.159, optimal linear-gap cost chaining achieves an expected recoverability of ≥ 1 - O(1/√m), generalizing the prior substitution-only result to a biologically realistic model.
主要结论
- The expected recoverability of an optimal chain under linear-gap cost chaining is ≥ 1 - O(1/√m) when the total mutation rate θ_T (sum of substitution, insertion, deletion rates) is less than 0.159.
- The expected runtime of the algorithm is O(m n^(3.15·θ_T) log n). For example, at a θ_T of 0.05 (similar to human-chimp divergence), the exponent is ~1.12, leading to near-linear scaling.
- The analysis successfully bridges theory and practice by extending the proof framework to handle indels, justifying the heuristic's empirical effectiveness on real genomic data which contains indels.
摘要: Given a sequence s1 of n letters drawn i.i.d. from an alphabet of size σ and a mutated substring s2 of length m<n, we often want to recover the mutation history that generated s2 from s1. Modern sequence aligners are widely used for this task, and many employ the seed-chain-extend heuristic with k-mer seeds. Previously, Shaw and Yu showed that optimal linear-gap cost chaining can produce a chain with 1−O(1/m) recoverability, the proportion of the mutation history that is recovered, in O(mn^(2.43θ) log n) expected time, where θ<0.206 is the mutation rate under a substitution-only channel and s1 is assumed to be uniformly random. However, a gap remains between theory and practice, since real genomic data includes insertions and deletions (indels), and yet seed-chain-extend remains effective. In this paper, we generalize those prior results by introducing mathematical machinery to deal with the two new obstacles introduced by indel channels: the dependence of neighboring anchors and the presence of anchors that are only partially correct. We are thus able to prove that the expected recoverability of an optimal chain is ≥1−O(1/√m) and the expected runtime is O(mn^(3.15·θ_T) log n), when the total mutation rate given by the sum of the substitution, insertion, and deletion mutation rates (θ_T = θ_i + θ_d + θ_s) is less than 0.159.