Paper List
-
Macroscopic Dominance from Microscopic Extremes: Symmetry Breaking in Spatial Competition
This paper addresses the fundamental question of how microscopic stochastic advantages in spatial exploration translate into macroscopic resource domi...
-
Linear Readout of Neural Manifolds with Continuous Variables
This paper addresses the core challenge of quantifying how the geometric structure of high-dimensional neural population activity (neural manifolds) d...
-
Theory of Cell Body Lensing and Phototaxis Sign Reversal in “Eyeless” Mutants of Chlamydomonas
This paper solves the core puzzle of how eyeless mutants of Chlamydomonas exhibit reversed phototaxis by quantitatively modeling the competition betwe...
-
Cross-Species Transfer Learning for Electrophysiology-to-Transcriptomics Mapping in Cortical GABAergic Interneurons
This paper addresses the challenge of predicting transcriptomic identity from electrophysiological recordings in human cortical interneurons, where li...
-
Uncovering statistical structure in large-scale neural activity with Restricted Boltzmann Machines
This paper addresses the core challenge of modeling large-scale neural population activity (1500-2000 neurons) with interpretable higher-order interac...
-
Realizing Common Random Numbers: Event-Keyed Hashing for Causally Valid Stochastic Models
This paper addresses the critical problem that standard stateful PRNG implementations in agent-based models violate causal validity by making random d...
-
A Standardized Framework for Evaluating Gene Expression Generative Models
This paper addresses the critical lack of standardized evaluation protocols for single-cell gene expression generative models, where inconsistent metr...
-
Single Molecule Localization Microscopy Challenge: A Biologically Inspired Benchmark for Long-Sequence Modeling
This paper addresses the core challenge of evaluating state-space models on biologically realistic, sparse, and stochastic temporal processes, which a...
STAR-GO: Improving Protein Function Prediction by Learning to Hierarchically Integrate Ontology-Informed Semantic Embeddings
Department of Computer Engineering, Bogazici University, Istanbul, Turkiye
30秒速读
IN SHORT: This paper addresses the core challenge of generalizing protein function prediction to unseen or newly introduced Gene Ontology (GO) terms by overcoming the limitations of existing models that either prioritize graph structure at the expense of semantic meaning or vice versa.
核心创新
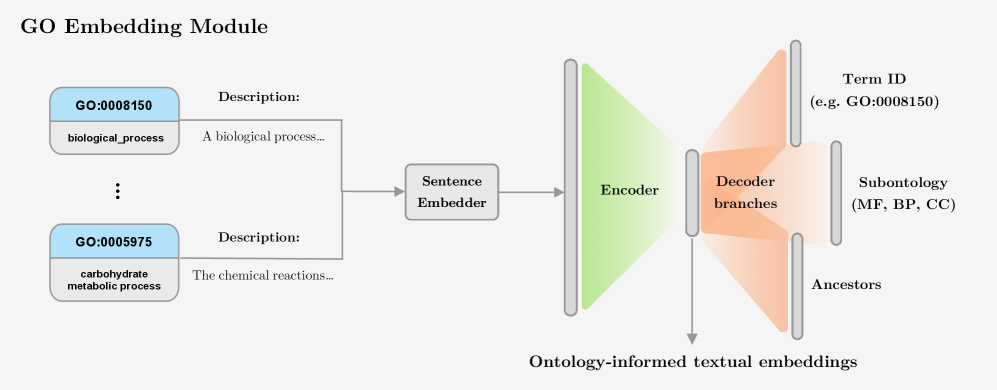
- Methodology Introduces a novel GO embedding module that integrates textual definitions (via SBERT-BioBERT) with ontology graph structure through a multi-task autoencoder, learning unified representations that preserve both semantic similarity and hierarchical dependencies.
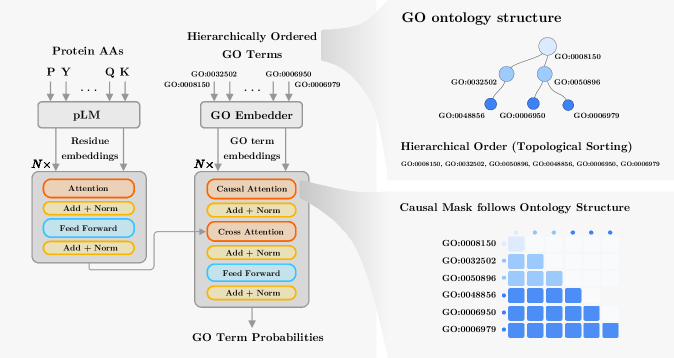
- Methodology Proposes a hierarchical Transformer decoder that processes GO terms in topological order (ancestors to descendants) using causal self-attention, enabling information propagation across ontology levels and capturing functional dependencies.
- Biology Demonstrates superior zero-shot generalization to unseen GO terms, particularly for Molecular Function and Biological Process terms, by effectively leveraging semantic information from textual definitions, which transfers better to novel ontology concepts than purely structural embeddings.
主要结论
- STAR-GO achieves state-of-the-art or competitive performance across all three GO subontologies (BP, CC, MF), with the highest AUC scores (e.g., 0.989 for BP, 0.988 for CC, 0.995 for MF), indicating strong term-level discriminability.
- In zero-shot evaluation on 16 held-out GO terms, STAR-GO variants achieve the highest AUCs in 13 cases, significantly outperforming baselines like DeepGOZero and DeepGO-SE, demonstrating superior generalization to unseen functions.
- Ablation studies reveal that semantic embeddings (STAR_T) achieve the best zero-shot results for most MF and BP terms (e.g., AUC of 0.949 for GO:0001228), while structural embeddings (STAR_S) perform best for a few terms but poorly for MF, highlighting the critical role of semantic information for generalization.
摘要: Motivation: Accurate prediction of protein function is essential for elucidating molecular mechanisms and advancing biological and therapeutic discovery. Yet experimental annotation lags far behind the rapid growth of protein sequence data. Computational approaches address this gap by associating proteins with Gene Ontology (GO) terms, which encode functional knowledge through hierarchical relations and textual definitions. However, existing models often emphasize one modality over the other, limiting their ability to generalize, particularly to unseen or newly introduced GO terms that frequently arise as the ontology evolves, and making the previously trained models outdated. Results: We present STAR-GO, a Transformer-based framework that jointly models the semantic and structural characteristics of GO terms to enhance zero-shot protein function prediction. STAR-GO integrates textual definitions with ontology graph structure to learn unified GO representations, which are processed in hierarchical order to propagate information from general to specific terms. These representations are then aligned with protein sequence embeddings to capture sequence–function relationships. STAR-GO achieves state-of-the-art performance and superior zero-shot generalization, demonstrating the utility of integrating semantics and structure for robust and adaptable protein function prediction. Availability: Code and pre-trained models are available at https://github.com/boun-tabi-lifelu/stargo.